- 1 Division of Psychiatry, Royal Edinburgh Hospital, University of Edinburgh, Edinburgh, UK
- 2 Molecular Medicine Centre, Western General Hospital, University of Edinburgh, Edinburgh, UK
- 3 Department of Psychiatry, Yale Medical School, New Haven, CT, USA
- 4 Medical Research Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, UK
- 5 Centre for Molecular Medicine, Ninewells Hospital and Medical School, Dundee, UK
- 6 College of Medical, Veterinary and Life Sciences, University of Glasgow, Glasgow, UK
- 7 Division of Psychology, University of Edinburgh, Edinburgh, UK
Hyperpolarization-activated cyclic nucleotide-gated (HCN) channels are encoded by four genes (HCN1–4) and, through activation by cyclic AMP (cAMP), represent a point of convergence for several psychosis risk genes. On the basis of positive preliminary data, we sought to test whether genetic variation in HCN1–4 conferred risk of depression or cognitive impairment in the Generation Scotland: Scottish Family Health Study. HCN1, HCN2, HCN3, and HCN4 were genotyped for 43 haplotype-tagging SNPs and tested for association with DSM-IV depression, neuroticism, and a battery of cognitive tests assessing cognitive ability, memory, verbal fluency, and psychomotor performance. No association was found between any HCN channel gene SNP and risk of depression, neuroticism, or on any cognitive measure. The current study does not support a genetic role for HCN channels in conferring risk of depression or cognitive impairment in individuals from the Scottish population.
Introduction
Mood disorders and schizophrenia are common and disabling conditions in which stressful life events and genetic risk factors are known to contribute to the risk of illness (Murray and Lopez, 1997; Sullivan et al., 2000). Impairment in cognitive function is associated with a variety of psychiatric disorders (Aleman et al., 1999; Austin et al., 2001; Bearden et al., 2001; McIntosh et al., 2005). Cognitive functions such as memory and planning appear to be particularly important and have been shown to contribute significantly to disability, diminished quality of life, and the ability to live independently (Alexopoulos et al., 2000; Dimitris et al., 2000). However, unlike positive psychotic symptoms and pathological mood states, these deficits are still largely irremediable (Galletly, 2009; Goldberg and Chengappa, 2009).
The relationship of cognitive performance to stress is described by the inverted U-shaped Yerkes–Dodson curve (Yerkes and Dodson, 1908). This relationship is partially mediated by release of dopamine and/or noradrenaline, with downstream effects on cyclic AMP (cAMP) signaling through Gs protein coupled receptors (Wang et al., 2007; Robbins and Arnsten, 2009). Although cAMP has many downstream effects, the time course of impaired executive function is perhaps most consistent with the rapid downstream effects of cAMP on hyperpolarization-activated cyclic nucleotide-gated (HCN) channels (Wang et al., 2007). Indeed, HCN channels have been shown to have a significant role in the development of stress-induced cognitive impairment (Arnsten, 2007). Stimulation by cAMP facilitates rapid opening of these receptors and an inwardly hyperpolarizing current referred to as Ih (Wahl-Schott and Biel, 2009). These currents, in turn, cause perturbations in neuronal signaling through reduced dendritic summation of incoming currents.
Several lines of evidence in animal studies are in support of the hypothesis that HCN channels may have a role in mediating the deleterious effects of stress. This work includes research demonstrating the possible efficacy of the compound guanfacine in improving attention and working memory, a drug which inhibits one of the pathways activating HCN channels (Wang et al., 2007).
Genetic variants DISC1, PDE4B, DGKH, and RGS4, each of which affect cAMP signaling, have risk modifying effects in mood disorders and schizophrenia (Thomson et al., 2005; Talkowski et al., 2006; Baum et al., 2007; Pickard et al., 2007). None to date show clear diagnostic specificity, although there is some evidence that they are associated with impairments in cognition common to several psychiatric disorders (Porteous et al., 2006). Since cAMP activates HCN channels, variants in their genes would therefore also be good candidates for genetic risk factors to a spectrum of psychiatric conditions. Indeed, we have previously reported preliminary evidence in support of an association between rs12905211 in HCN4 and risk of a broad psychiatric phenotype consisting of depression, anxiety disorders, and obsessive-compulsive disorder (Kelmendi et al., 2011). The original study (N = 285 cases, 384 controls) and the strength of the association were modest in size (OR = 1.35–1.5) but warranted further investigation. Here, we seek further evidence for an association of HCN channel genes and common variants with depression, neuroticism, and cognitive function in a large independent cohort. In addition, we used neuroticism as a proxy for stress sensitivity and examined its interaction with HCN1–4, whose effects on risk of MDD may be mediated through stress-induced cognitive impairment.
Materials and Methods
The Generation Scotland: the Scottish Family Health Study (GS:SFHS) is a multi-institution study of the genetic contributions to health and disease in the Scottish population (Smith et al., 2006). The study identified potential participants from general practices throughout Scotland, along with as many first- and second-degree relatives as could be recruited; volunteers and their relatives were also recruited. Participants completed a pre-assessment questionnaire, attended a clinic where physical measurements were made and undertook detailed measures of neuropsychological function. A sample of blood or saliva was taken from which genomic DNA was extracted.
The neuropsychological test battery used included the Mill Hill Vocabulary Test (Raven et al., 1977; general cognitive ability), the Logical Memory Test (Wechsler, 1998; mnemonic function), the Digit Symbol Substitution test from the WAIS IIIR (Wechsler, 1998; psychomotor performance), and an estimate of verbal fluency (Lezak, 1995; executive function) which measured the numbers of words beginning with the letters C, F, or L generated verbally within 1 min. Individuals also completed the NEO five-factor inventory (NEO-FFI; Costa and McCrae, 1989) in order to provide measures of neuroticism and extraversion, traits previously linked to susceptibility to depression (Kendler et al., 1993, 2006).
A total of 23,960 individuals have provided informed consent of which 21,476 underwent clinical and cognitive assessments and provided DNA. Participants were screened for emotional and psychiatric disorder using the screening module from Structured Clinical Interview for DSM-IV (SCID, axis I) with a researcher trained in its use. An affirmative answer to at least one screening question [Question (1) “Have you ever seen anybody for emotional or psychiatric problems?” Question (2) “Was there ever a time when you, or someone else, thought you should see someone because of the way you were feeling or acting?”] was interpreted as a positive screen. Those who screened positive were invited to continue the interview, which focused on major depression and bipolar disorder. Lifetime diagnoses were made according to DSM-IV criteria and information on the lifetime number of depressive episodes was also provided. Researchers received field training and reliability assessments involving an independent, blind rating of the interviews by a trained psychiatrist (Donald J. MacIntyre). Inter-rater reliability for presence or absence of a lifetime diagnosis of major depressive disorder was determined: in the sample (N = 58) Kappa = 0.86 (p < 0.001), 95% CI (0.7, 1.0).
A total of 1501 confirmed cases of DSM-IV major depressive disorder were identified in the full GS:SFHS cohort, using the SCID. Screened negative controls (N = 4499) were matched to cases for age and sex as far as possible. The sample size had >80% power to detect an association between the HCN4 SNP rs12905211, previously reported to show an association with a broad psychiatric phenotype, and MDD (calculation performed using online software: http://pngu.mgh.harvard.edu/~purcell/gpc/).
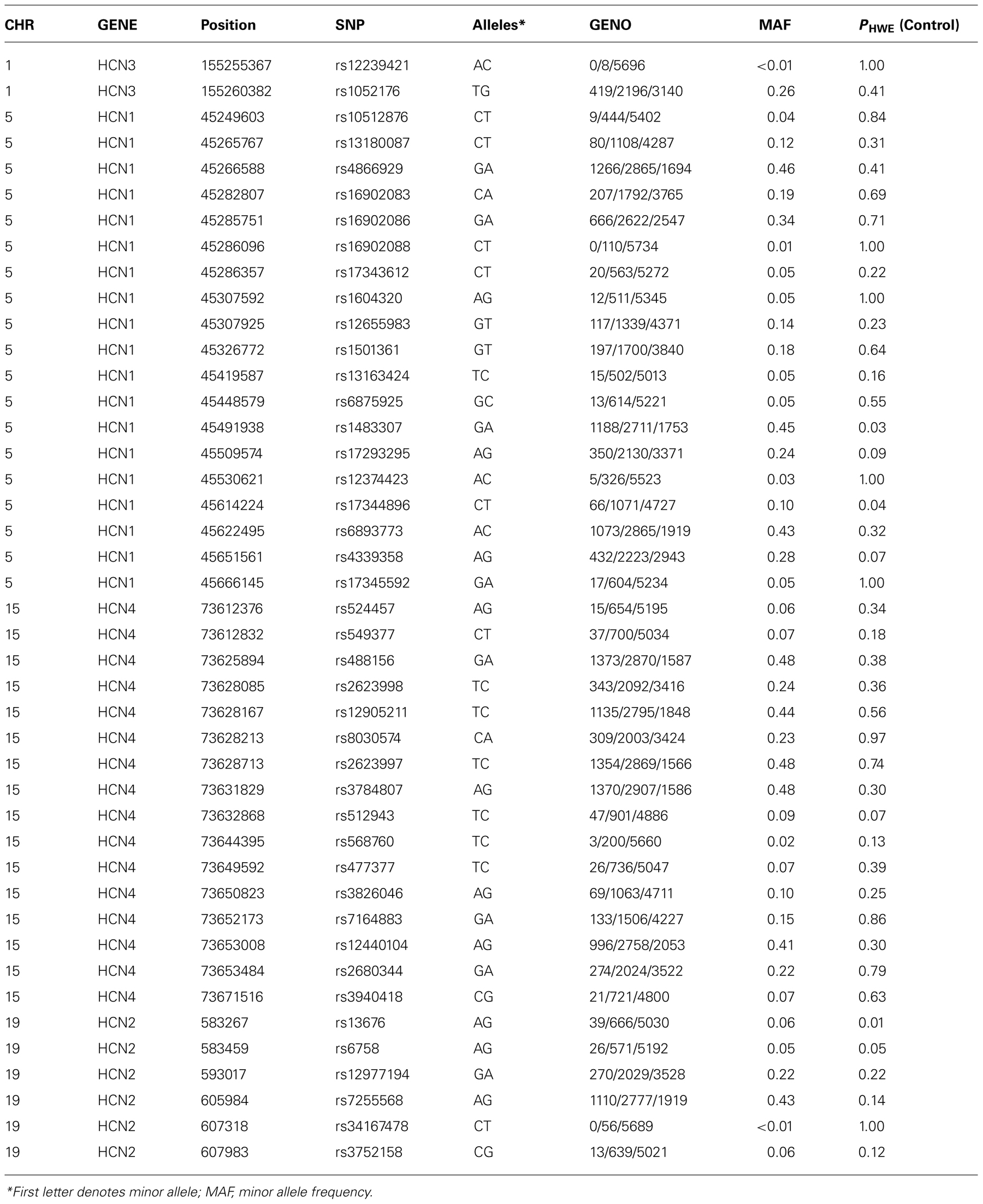
Genotyping of DNA was performed on the Applied Biosystems OmniExpress Array using TaqMan SNP genotyping assays. SNP assays were selected for HCN1 (5p12), HCN2 (19p13.3), HCN3 (1q22), and HCN4 (15q24.1) using Tagger with HaploView software using a pairwise allele tagging threshold of r2= 0.8 and a minimum minor allele frequency (MAF) of 1%. Details of the genomic regions spanned by each SNP set are provided in Table 2 and included the promoter and 5′ regions of each gene 2 Kb upstream from the transcription start site and 2 Kb downstream from the 3′ UTR.
Statistical Analysis
All clinical, cognitive, and genotyping data was imported into the R software package for analysis of demographic and historical information. The mean age of controls, individuals with an episode of MDD, and those with recurrent MDD were then compared using univariate ANOVA. These variables differed between the three groups, therefore subsequent analyses of the cognition, neuroticism, and extraversion are conducted controlling for these potential confounds. Post hoc tests were conducted using Tukey's method. Depression, neuroticism, extraversion, and all cognitive variables were then exported to the PLINK (Purcell et al., 2007) genetic software analysis package (version 1.07, http://pngu.mgh.harvard.edu/purcell/plink/) for further analysis.
Forty-three SNPs out of 44 SNPs genotyped passed quality control procedures (>95% genotype success rate, no significant deviation from Hardy–Weinberg equilibrium, PHWE <0.005 in control participants). Genetic association of HCN1–4 SNPs with depression was tested by conducting a logistic regression analysis, adjusted for age and sex, followed by a likelihood ratio-based test of association for each individual SNP. We analyzed the association with all MDD (single episode and recurrent MDD) initially followed by an analysis of recurrent MDD only on the basis that recurrent MDD may represent a more heritable form of the disorder (Levinson, 2006). Genetic association with continuous traits was conducted using a linear regression model. All analyses were adjusted for age, any diagnosis of MDD, and sex. Sixty-five individuals of non-European ancestry were excluded from the genetic analyses on the basis of self reported ethnicity and family history. Finally, to examine whether there was an association between each SNP and each measure based on a trait liability to stress, we repeated the above linear models examining HCN1–4 SNP effects on cognition including the SNP × neuroticism interaction term (as well as the SNP and neuroticism main effects), whilst retaining “any episode of depression,” age and sex as covariates in the model. All of the analyses with quantitative traits were then repeated without MDD as a covariate, with no changes in any of the conclusions. We then reported any associations that survived a false discovery rate correction (Benjamini and Hochberg, 2000) of p < 0.05 as significant, although nominally significant results are reported in the tables.
Results
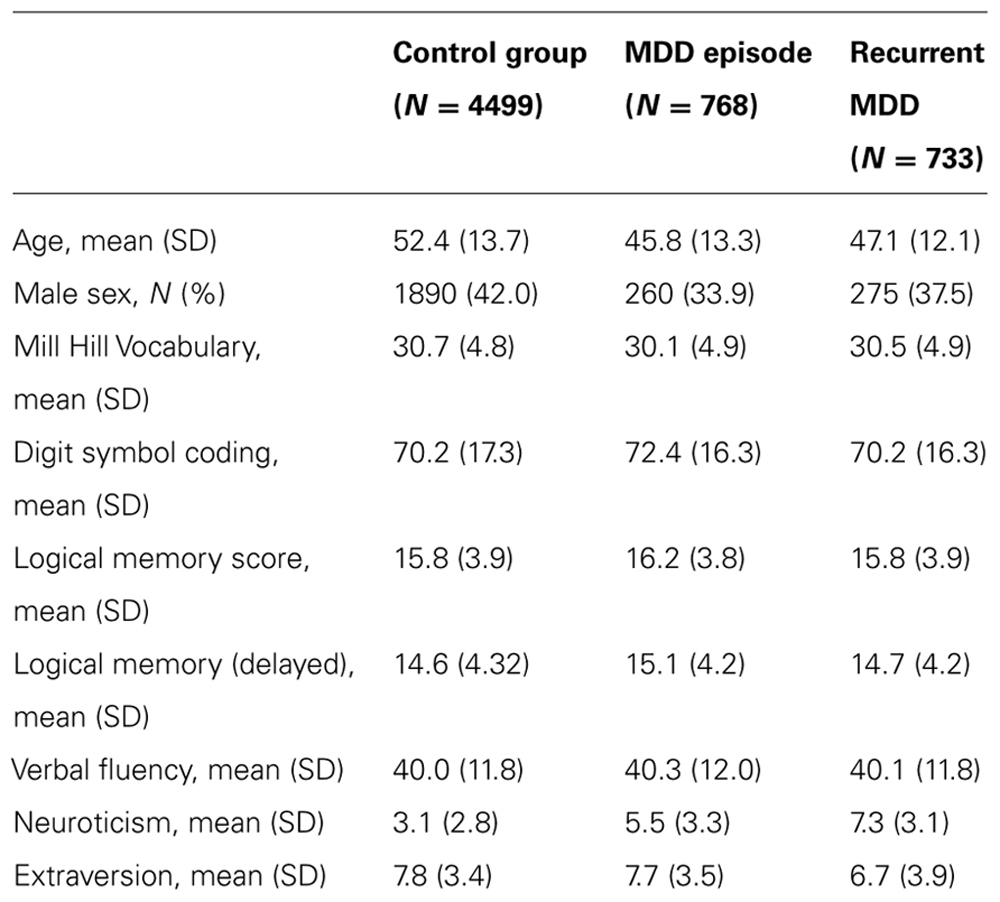
At the start of the study 6000 individuals provided useable clinical and cognitive data of which 4499 had no psychiatric diagnosis and 1501 met criteria for DSM-IV major depressive disorder (N = 733 recurrent, N = 768 with a single episode). The rate of depression in the GS:SFHS cohort as a whole reflects that in the Scottish population, but since this study was limited to 6000 samples, we maximized statistical power by over sampling individuals with MDD initially before filling the remaining spaces with control participants. Demographic and historical details of the diagnostic groups are shown in Table 1 and details of the genotyping in Table 2.
There were significant differences in age and sex between the groups, with both depressed groups being younger and more frequently female than controls. Unadjusted mean values of neuroticism, extraversion, and the cognitive variables are presented in Table 1. After adjustment for age and sex, there were significant differences in Mill Hill Vocabulary (a higher mean vocabulary score was found in individuals with recurrent depression than in controls), Digit symbol coding (better performance in controls than in either depressed group), neuroticism scores (recurrent MDD > single episode MDD > controls), and extraversion score (recurrent MDD < single episode MDD < controls).
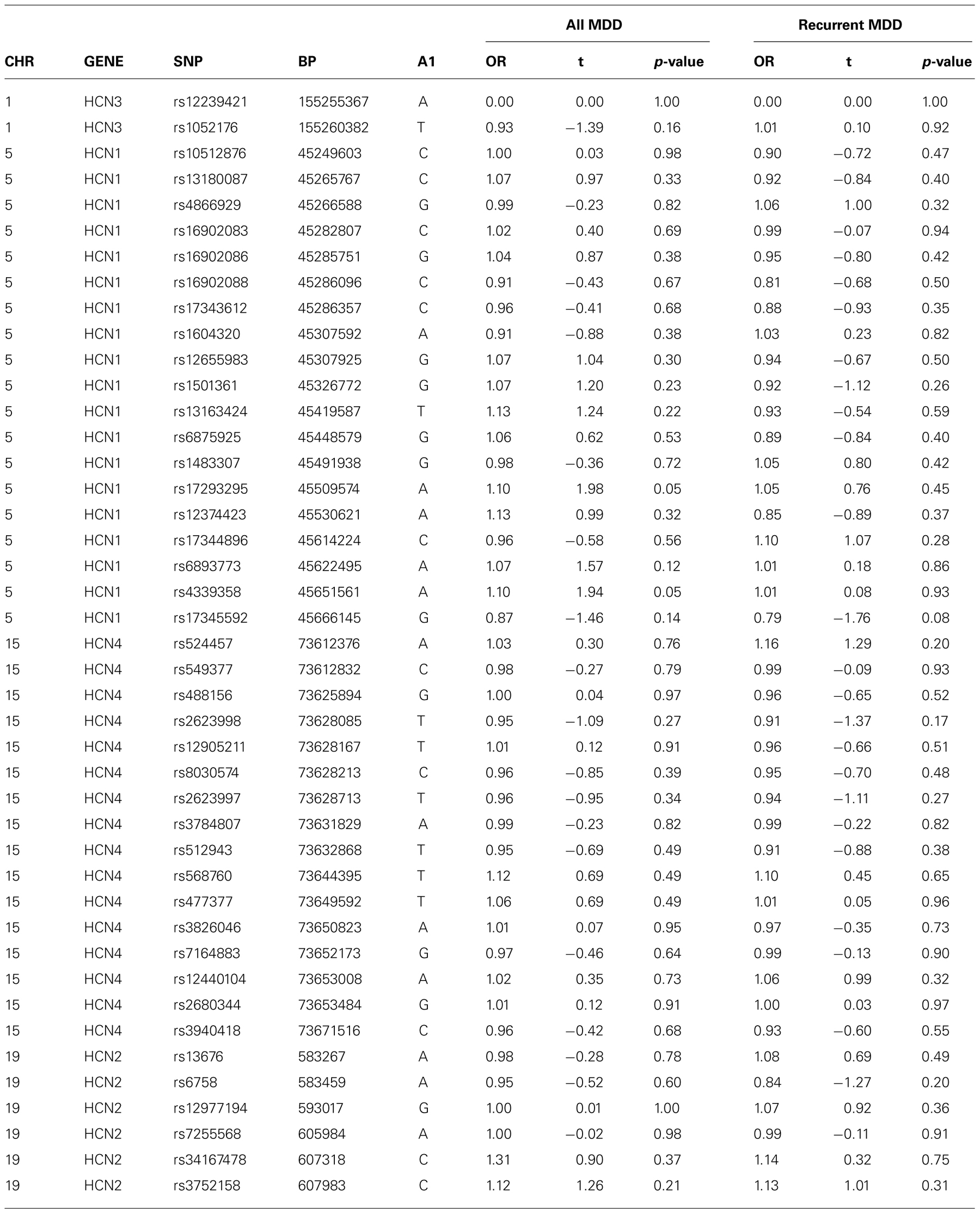
There was no evidence of an association between any SNP in HCN1–4 and major depression (recurrent and singe episodes combined) in the GS:SFHS sample (Table 3) after adjustment for age and sex (data before adjustment available on request). Similarly, there were no significant associations between SNPs in HCN1–4 and neuroticism, extraversion or any domain of cognition. The smallest nominally significant associations between SNPs in HCN1–4 and individual traits are shown in Table 4. We then repeated the analyses examining HCN1–4 SNP effects on neuroticism, extraversion, and cognition in the absence of MDD as a covariate, and all of the associations remained non-significant. Finally, we found that there was no statistically significant interaction between SNPs in HCN1–4 and neuroticism for the dependent variables considered in the current study.
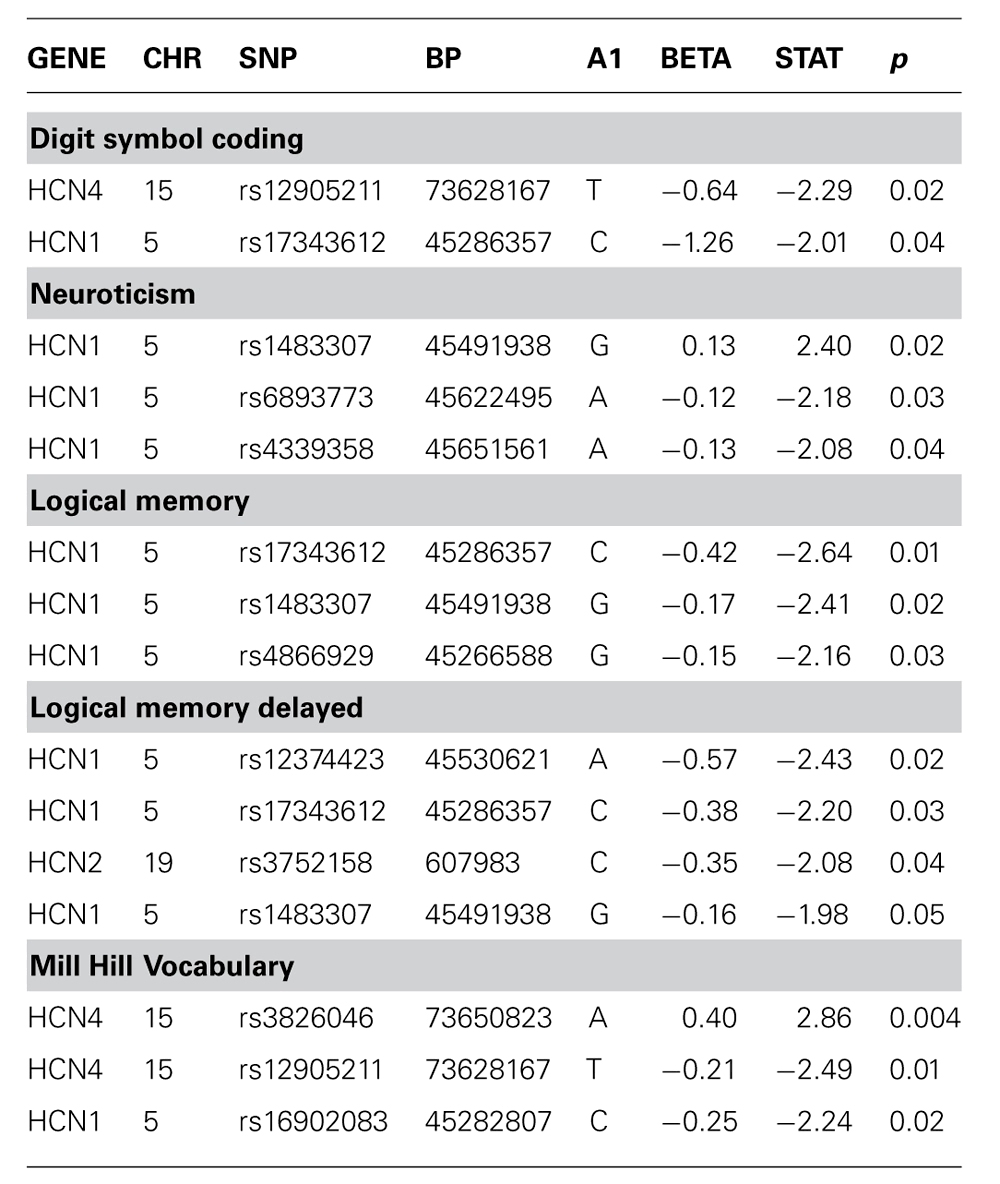
TABLE 4. Association of HCN1–4 with cognition, neuroticism or extraversion where unadjusted p-value <0.05.
Conclusion
In the current study, no evidence could be found to support an association between genetic variants in HCN1–4 and liability to depression, including measures of neuroticism or extraversion. Furthermore, no evidence was found to support an association between these four HCN channel genes and cognitive impairment, either as main effects or as interacting risk factors with neuroticism.
Whilst a role for HCN channels in mediating these effects cannot be excluded, common variants in HCN1–4 appear to have no effect on risk of depression or on cognitive performance in Caucasian individuals of presumed Scottish descent. The current study can also not exclude the possibility that rare functional variants in HCN1–4 may affect risk of depression or cognition.
The results of the present investigation differ from our previous study where we reported a significant association between rs12905211 in HCN4 and a phenotype comprised of unipolar and bipolar depression and/or anxiety disorder. The two studies differed in a number of important ways. Notably, the present study utilized a population sample whereas we previously studied a particularly ill sample of patients recruited from a tertiary care setting. The present sample was also composed almost exclusively of Caucasian non-Hispanic British participants whereas the previous sample was composed of Hispanic American individuals with mixed backgrounds. Finally, the present study was much larger than the previous sample of less than 700 participants. However, the finding of a significant association in one study and a subsequent failure to replicate in another has frequently been referred to in the literature as “winner's curse” and underlines the importance of independent replication.
Certain limitations of the current study should be taken into account when interpreting these results. Firstly, the psychiatric and psychological assessments within GS:SFHS were not conducted by a trained psychiatrist or post-doctoral psychologist. All assessments were however conducted by a research assistant trained and validated in the use of the cognitive instruments or the SCID after a period of training with an experienced psychiatrist (Donald J. MacIntyre) or psychologist (Ian J. Deary). The cognitive and personality assessments were also conducted blind to genotype and with ongoing quality assurance, reliability assessments and after extensive field-testing. SCID-based interviews took into account both current and lifetime histories of mood disorder. The reliability of the SCID ratings compared with that of a blind, independent psychiatrist were excellent (ICC = 0.86, 95% CI 0.7–1.0) and are unlikely to have obscured any positive findings. A further consideration in interpreting the current findings is the fact that four HCN1–4 SNP assays initially failed manufacture, three of which were in the HCN2 gene. Replacements were subsequently found in HCN1 although two further proxy SNPs failed manufacture in HCN2 after a second account and at least eight SNPs in HapMap 2 (release 2) could not be captured using the current assays. It is therefore important to note that a proportion of common genetic variation in HCN2 was not captured in the current study.
It is also possible that the current study, although it is the largest by far to examine this issue to date, was still nonetheless of insufficient size to detect an important genetic effect of HCN1–4. It is therefore possible that these may emerge as significant risk factors for depression or cognitive impairment in larger samples, although the sample size of 6000 is exceptionally large for a single case–control association study.
It is also important to note that, whereas preclinical research has employed measures of attention and working memory in elucidating a role for HCN channels in cognition, we measured verbal fluency. Although such measures are very useful clinically it is possible that the role of HCN channels is specific to certain sub-domains of executive functioning, and future studies should therefore include measures of attention and working memory.
In the current study we found no evidence to support a role for HCN channel genes in the etiology of depression. However, the evidence from basic neuroscience still supports a role for these channels in conveying the effects of stress-related cognitive impairment. Future studies might seek to address their functional role in larger samples, in individuals with rare variants, or using in vitro methods.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The Generation Scotland: Scottish Family Health Study (GS:SFHS) funded by the Chief Scientist Office of the Scottish Government Health Department (CZD/016/006). Access details can be found at http://www.generationscotland.co.uk/. The work described herein was also supported by the Chief Scientists Office of the Scottish Government Health Department (Reference: ETM/84). We are grateful to all the families who took part in the Generation Scotland: Scottish Family Health Study, the general practitioners and Scottish School of Primary Care for their help in recruiting them, and the whole Generation Scotland team, which includes academic researchers, clinic staff, laboratory technicians, clerical workers, IT staff, statisticians and research managers. Andrew M. McIntosh and Jeremy Hall are supported by Scottish Funding Council Senior Clinical Fellowships.
References
Aleman, A., Hijman, R., de Haan, E. H. F., and Kahn, R. S. (1999). Memory impairmnet in schizophrenia: a meta-analysis. Am. J. Psychiatry 156, 1358–1366.
Alexopoulos, G. S., Meyers, B. S., Young, R. C., Kalayam, B., Kakuma, T., Gabrielle, M., Sirey, J. A., and Hull, J. (2000). Executive dysfunction and long-term outcomes of geriatric depression. Arch. Gen. Psychiatry 57, 285–290.
Arnsten, A. F. (2007). Catecholamine and second messenger influences on prefrontal cortical networks of “representational knowledge”: a rational bridge between genetics and the symptoms of mental illness. Cereb. Cortex 17(Suppl. 1), i6–15.
Austin, M. P., Mitchell, P., and Goodwin, G. M. (2001). Cognitive deficits in depression: possible implications for functional neuropathology. Br. J. Psychiatry 178, 200–206.
Baum, A. E., Akula, N., Cabanero, M., Cardona, I., Corona, W., Klemens, B., Schulze, T. G., Cichon, S., Rietschel, M., and Nöthen, M. M. (2007). A genome-wide association study implicates diacylglycerol kinase eta (DGKH) and several other genes in the etiology of bipolar disorder. Mol. Psychiatry 13, 197–207.
Bearden, C. E., Hoffman, K. M., and Cannon, T. D. (2001). The neuropsychology and neuroanatomy of bipolar affective disorder: a critical review. Bipolar Disord. 3, 106–150.
Benjamini, Y., and Hochberg, Y. (2000). On the adaptive control of the false discovery fate in multiple testing with independent statistics. J. Educ. Behav. Statist. 25, 60–83.
Costa, P. T., and McCrae, R. R. (1989). The NEO-PI/NEO-FFI Manual supplement. Odessa, FL: Psychological Assessemnt Rescources.
Dimitris, N. K., George, S. A., and Christopher, M. (2000). Symptoms of striatofrontal dysfunction contribute to disability in geriatric depression. Int. J. Geriatr. Psychiatry 15, 992–999.
Galletly, C. (2009). Recent advances in treating cognitive impairment in schizophrenia. Psychopharmacology 202, 259–273.
Goldberg, J. F., and Chengappa, K. N. (2009). Identifying and treating cognitive impairment in bipolar disorder. Bipolar Disord. 11(Suppl. 2), 123–137.
Kelmendi, B., Holsbach-Beltrame, M., McIntosh, A. M., Hilt, L., George, E. D., Kitchen, R. R., Carlyle, B. C., Pittenger, C., Coric, V., Nolen-Hoek-sema, S., Sanacora, G., and Simen, A. A. (2011). Association of polymorphisms in HCN4 with mood disorders and obsessive compulsive disorder. Neurosci. Lett. 496, 195–199.
Kendler, K. S., Gatz, M., Gardner, C. O., and Pedersen, N. L. (2006). Personality and major depression: a Swedish longitudinal, population-based twin study. Arch. Gen. Psychiatry 63, 1113–1120.
Kendler, K. S., Neale, M. C., Kessler, R. C., Heath, A. C., and Eaves, L. J. (1993). A longitudinal twin study of personality and major depression in women. Arch. Gen. Psychiatry 50, 853–862.
McIntosh, A. M., Harrison, L. K., Forrester, K., Lawrie, S. M., and Johnstone, E. C. (2005). Neuropsychological impairments in people with schizophrenia or bipolar disorder and their unaffected relatives. Br. J. Psychiatry 186, 378–385.
Murray, C. J., and Lopez, A. D. (1997). Global mortality, disability, and the contribution of risk factors: Global Burden of Disease Study. Lancet 349, 1436–1442.
Pickard, B. S., Thomson, P. A., Christoforou, A., Evans, K. L., Morris, S. W., Porteous, D. J., Blackwood, D. H., and Muir, W. J. (2007). The PDE4B gene confers sex-specific protection against schizophrenia. Psychiatr. Genet. 17, 129–133.
Porteous, D. J., Thomson, P., Brandon, N. J., and Millar, J. K. (2006). The genetics and biology of DISC1 – an emerging role in psychosis and cognition. Biol. Psychiatry 60, 123–131.
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M. A., Bender, D., Maller, J., Sklar, P., de Bakker, P. I., Daly, M. J., and Sham, P. C. (2007). PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575.
Raven, J. C., Court, J. H., and Raven, J. (1977). Manual for Raven's Progressive Matrices and Vocabulary Scales. London, UK: H. K. Lewis.
Robbins, T. W., and Arnsten, A. F. (2009). The neuropsychopharmacology of fronto-executive function: monoaminergic modulation. Annu. Rev. Neurosci. 32, 267–287.
Smith, B. H., Campbell, H., Blackwood, D., Connell, J., Connor, M., Deary, I. J., Dominiczak, A. F., Fitzpatrick, B., Ford, I., Jackson, C., Haddow, G., Kerr, S., Lindsay, R., McGilchrist, M., Morton, R., Murray, G., Palmer, C. N., Pell, J. P., Ralston, S. H., St Clair, D., Sullivan, F., Watt, G., Wolf, R., Wright, A., Porteous, D., and Morris, A. D. (2006). Generation Scotland: the Scottish Family Health Study; a new resource for researching genes and heritability. BMC Med. Genet. 7, 74. doi: 10.1186/1471-2350-7-74
Sullivan, P. F., Neale, M. C., and Kendler, K. S. (2000). Genetic epidemiology of major depression: review and meta-analysis. Am. J. Psychiatry 157, 1552–1562.
Talkowski, M. E., Seltman, H., Bassett, A. S., Brzustowicz, L. M., Chen, X., Chowdari, K. V., Collier, D. A., Cordeiro, Q., Corvin, A. P., Deshpande, S. N., Egan, M. F., Gill, M., Kendler, K. S., Kirov, G., Heston, L. L., Levitt, P., Lewis, D. A., Li, T., Mirnics, K., Morris, D. W., Norton, N., O'Donovan, M. C., Owen, M. J., Richard, C., Semwal, P., Sobell, J. L., St Clair, D., Straub, R. E., Thelma, B. K., Vallada, H., Weinberger, D. R., Williams, N. M., Wood, J., Zhang, F., Devlin, B., and Nimgaonkar, V. L. (2006). Evaluation of a susceptibility gene for schizophrenia: genotype based meta-analysis of RGS4 polymorphisms from thirteen independent samples. Biol. Psychiatry 60, 152–162.
Thomson, P. A., Wray, N. R., Millar, J. K., Evans, K. L., Le Hellard, S., Condie, A., Muir, W. J., Blackwood, D. H. R., and Porteous, D. J. (2005). Association between the TRAX/DISC locus and both bipolar disorder and schizophrenia in the Scottish population. Mol. Psychiatry 10, 657–668.
Wahl-Schott, C., and Biel, M. (2009). HCN channels: structure, cellular regulation and physiological function. Cell. Mol. Life Sci. 66, 470–494.
Wang, M., Ramos, B. P., Paspalas, C. D., Shu, Y., Simen, A., Duque, A., Vijayraghavan, S., Brennan, A., Dudley, A., Nou, E., Mazer, J. A., McCormick, D. A., and Arnsten, A. F. (2007). Alpha2A-adrenoceptors strengthen working memory networks by inhibiting cAMP-HCN channel signaling in prefrontal cortex. Cell 129, 397–410.
Wechsler, D. (1998). WMS-IIIUK Administration and Scoring Manual. London, UK: Psychological Corporation.
Keywords: stress, depression, HCN channel, genetics, association, cognition, neuroticism
Citation: Mcintosh AM, Simen AA, Evans KL, Hall J, MacIntyre DJ, Blackwood D, Morris AD, Smith BH, Dominiczak A, Porteous D, Deary IJ and Thomson PA (2012) Genetic variation in hyperpolarization-activated cyclic nucleotide-gated channels and its relationship with neuroticism, cognition, and risk of depression. Front. Gene. 3:116. doi: 10.3389/fgene.2012.00116
Received: 25 March 2012; Accepted: 05 June 2012;
Published online: 02 July 2012.
Edited by:
Jenae Neiderhiser, The Pennsylvania State University, USAReviewed by:
Jaime Derringer, University of Colorado, USAAndrew McQuillin, University College London, UK
Copyright: © 2012 Mcintosh, Simen, Evans,Hall,MacIntyre, Blackwood,Morris, Smith, Dominiczak, Porteous, Deary and Thomson. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Andrew M. McIntosh, Division of Psychiatry, Royal Edinburgh Hospital, University of Edinburgh, Kennedy Tower, Edinburgh EH10 5HF, UK. e-mail: andrew.mcintosh@ed.ac.uk