- 1Department of Clinical and Molecular Sciences, Università Politecnica delle Marche, Ancona, Italy
- 2Center of Clinical Pathology and Innovative Therapy, I.N.R.C.A. National Institute, Ancona, Italy
Evidence on circulating microRNAs (miRNAs) is indisputably opening a new era in systemic and tissue-specific biomarker research, highlighting new inter-cellular and inter-organ communication mechanisms. Circulating miRNAs might be active messengers eliciting a systemic response as well as non-specific “by-products” of cell activity and even of cell death; in either case they have the potential to be clinically relevant biomarkers for a number of physiopathological processes, including inflammatory responses and inflammation-related conditions. A large amount of evidence indicates that miRNAs can exert two opposite roles, activating as well as inhibiting inflammatory pathways. The inhibitory action probably relates to the need for activating anti-inflammatory mechanisms to counter potent proinflammatory signals, like the nuclear factor kappaB (NF-κB) pathway, to prevent cell and tissue destruction. MiRNA-based anti-inflammatory mechanisms may acquire a crucial role during aging, where a chronic, low-level proinflammatory status is likely sustained by the cell senescence secretome and by progressive activation of immune cells over time. This process entails age-related changes, especially in extremely old age, in those circulating miRNAs that are capable of modulating the inflammatory status (inflamma-miRs). Interestingly, a number of such circulating miRNAs seem to be promising biomarkers for the major age-related diseases that share a common chronic, low-level proinflammatory status, such as cardiovascular disease (CVD), type 2 diabetes mellitus (T2DM), Alzheimer Disease (AD), rheumatoid arthritis (RA), and cancers.
Inflamma-miRs
The inflammatory response comprises complex biological reactions that require a fine-tuned integration between a range of immune system cell classes and an extensive network of biomolecules, which until recently had been thought to be largely cytokines. However, identification of a vast repertoire of non-coding microRNA (miRNA) in the mammalian genome has completely revolutionized our understanding of most biological processes, including inflammation (Nilsen, 2007). MiRNAs play a significant role in gene regulation acting as repressors as well as activators, mainly at the post-transcriptional level (Breving and Esquela-Kerscher, 2010). Since a single miRNA can target several genes, and multiple miRNAs share common targets, miRNAs are particularly suited for regulating processes and pathways at the “network” level (Inukai and Slack, 2013).
Sensing of dangerous signals by the innate immune system involves a number of germline-encoded pattern recognition receptors (PRRs) that can detect both conserved pathogen-associated molecular profiles (PAMPs) expressed on micro-organisms and altered endogenous ligands, mostly released by necrotic, senescent, and/or damaged cells (DAMPs). Among PRRs, toll-like receptors (TLRs) play a central role, since their engagement activates a potent proinflammatory pathway (Kawai and Akira, 2011). TLR signaling initiates from different adaptor proteins, such as myeloid differentiation factor 88 (MyD88) or TIR-domain-containing adapter protein-inducing interferon-β, which in turn activate several downstream pathways, leading to activation of transcription factor nuclear factor kappaB (NF-κB), mitogen-activated protein kinases (MAPKs), and members of the interferon regulatory factor family (Dunne and O'Neill, 2005). Fine tuning of TLR signaling prevents generation of harmful and inappropriate inflammatory responses without lowering the surveillance for potentially dangerous signals. Deregulation of the whole network can have destructive effects and lead to tissue damage: this is a hallmark of chronic inflammation, which is often associated with age-related diseases (Olivieri et al., 2013a).
A mounting body of evidence has been documenting a relatively small number of miRNAs that are involved in regulating inflammation: their prototypes are miR-155, miR-21, and miR-146a (Quinn and O'Neill, 2011), hereinafter referred to as inflamma-miRs.
In physiological conditions transcription of miR-155, miR-21, and miR-146a is at baseline levels; however, initiation of proinflammatory TLR signaling immediately results in strong co-induction of their expression through a mechanism that is largely NF-κB-dependent (Boldin and Baltimore, 2012).
Although the importance of inflamma-miRs in innate immune response regulation is widely accepted, the molecular mechanism of their action has proved to be highly complex. Early investigations disclosed that miR-146a acts as a negative regulator of TLR signaling by targeting both tumor necrosis factor receptor-associated factor 6 (TRAF6) and IL-1 receptor-associated kinase 1 (IRAK-1) (Taganov et al., 2006). MiR-21 was found to down-regulate the expression of IRAK and MyD88 (Chen et al., 2013), as well as of programmed cell death protein 4 (PDCD4), switching the cell program from proinflammatory to anti-inflammatory, mainly as reflected by IL-10 production (Sheddy et al., 2010).
Additional findings showing that miR-155 can also negatively modulate TLR signaling by targeting MyD88, Tak1-binding protein 2 (TAB2) (Ceppi et al., 2009), and the I-κ-B kinase ε (IKKε), (Liang et al., 2011) have generated a model where inflamma-miRs operate as a negative feedback loop to protect the organism against overwhelming inflammation. However, subsequent research has disclosed that inflamma-miRs could play a dual function, inhibiting as well as inducing TLR signaling (Kondo et al., 2012). This is the case of miR-155, which both suppresses and enhances TLR signaling by silencing Src homology 2 domain-containing protein tyrosine phosphatase-1 (SHP-1), a negative regulator of IRAK activity (O'Connell et al., 2009); suppressor of cytokine signaling 1 (SOCS1); and B-cell lymphoma 6 protein (BCL6), a transcription factor that attenuates NF-κB signaling (Nazari-Jahantigh et al., 2012). A similar action is also exerted by miR-21, which can function as an agonist of single-stranded RNA-binding TLRs and can therefore induce NF-κB activation and secretion of inflammatory molecules (Fabbri et al., 2012).
Although inflamma-miRs are co-induced during TLR signaling, several observations suggest that they do not act redundantly and simultaneously, but rather cooperate to control TLR signaling through functionally different performances. At least for miR-146a and miR-155 this has recently been confirmed by Schulte and colleagues, who found different induction behaviors and mRNA target profiles in the response to microbial lipopolysaccaride (LPS), (Schulte et al., 2013). Such coordination of inflamma-miRs could go beyond suppression of TLR signaling and contribute to regulating the immunity/inflammation balance, as proposed by Akira's group (Kondo et al., 2012).
Knockout mouse models have been instrumental in shedding light on the role of inflamma-miRs in inflammation and inflammation-related diseases. Transcriptome analysis of bic/miR-155-deficient CD4+ T cells in mice identified a wide spectrum of miR-155-regulated genes, including cytokines, chemokines, and transcription factors, suggesting that bic/miR-155 plays a key role in immune system homeostasis and function (Rodriguez et al., 2007). Knockout of miR-146a gene in C57BL/6 mice involves increased transcription of NF-κB-regulated genes (Zhao et al., 2011). These animals also develop myeloid sarcomas and lymphomas as well as chronic myeloproliferation in bone marrow. Genetic ablation of NF-κB p50 suppresses the myeloproliferation, demonstrating that NF-κB dysregulation is responsible for the myeloproliferative disease (Zhao et al., 2011).
Details on the role of inflamma-miRs in controlling TLR signaling are just beginning to be explored, and further investigations are warranted to gain insights not only into their individual contribution to the homeostasis of the innate immune response, but also into the consequences of their deregulation in conditions characterized by chronic inflammation.
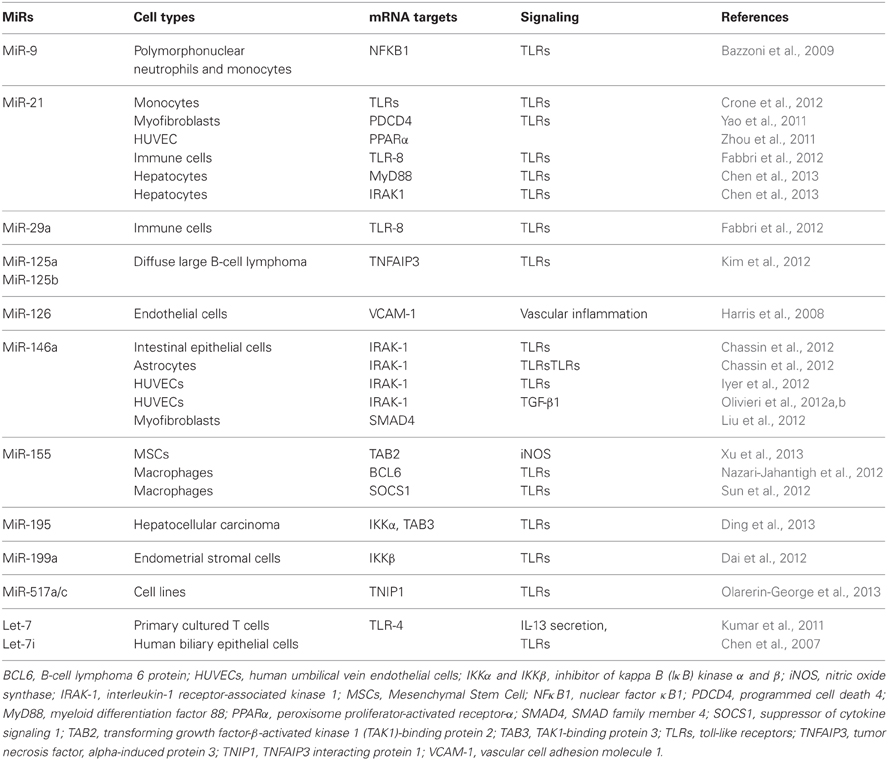
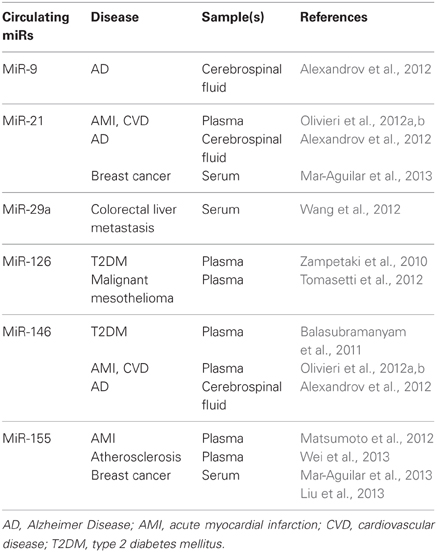
The most recent evidence for the involvement of inflamma-miRs in modulating the proinflammatory response at the cell level is summarized in Table 1. Few cellular inflamma-miRs have been reported to be modulated in plasma/serum samples in different physiopathological conditions, as depicted in Table 2.
Relationship between Tissue and Circulating Inflamma-miRs
It is becoming increasingly clear that multiple miRNAs can be deregulated in several diseases (Reid et al., 2011). Detection and identification of stable miRNAs in body fluids has opened a new era in systemic biomarker research directed at improving clinical diagnosis/prognosis within translational medicine. The numerous reports of changes in miRNA expression have often failed to clarify whether they are the cause or the effect of malfunction. Identification of multiple miRNA changes in plasma is important, because specific miRNA combinations unique to a normal physiological or pathological state can provide a useful reference. These findings also raise several questions: do circulating miRNA levels match tissue expression levels? If this is the case, does it also apply to pathological conditions? A greater understanding of the relationship between the level of tissue and plasma miRNAs should help uncover the origin and/or function of circulating miRNAs.
So far only a limited number of studies, mostly in oncology, have addressed the issue. Findings have been controversial: some researchers described a similar trend of alteration both in circulating and tissue miRNAs (Brase et al., 2011), whereas others reported that only a subset of circulating miRNAs reflect tissue cellular abundance (e.g., mammary epithelial tumor), and suggested that cells might have developed a mechanism that selects specific miRNAs for release or retention (Pigati et al., 2010). Studies of animal models found distinct roles for circulating and tissue miRNAs (Waters et al., 2012).
The way circulating miRNAs are delivered into the bloodstream is also poorly understood. Evidence from several studies indicates that miRNAs exist freely in the systemic circulation despite their susceptibility to degradation by extracellular RNAses, raising the issue of which mechanisms underpin their unexpected stability. On the one hand, different studies have revealed that miRNAs are secreted into the extracellular space or the bloodstream either in microvesicles or in exosomes (Valadi et al., 2007; Collino et al., 2010; Hosoda et al., 2011). Secretory exosomes are promising candidates for intercellular miRNAs transfer not only because they provide a protected environment, but also because they may directly transfer internal components to target cells by receptor-mediated interactions. This is consistent with the observation that plasma exosomes can deliver exogenous short interfering RNA to monocytes and lymphocytes (Wahlgren et al., 2012). On the other hand, a number of reports have shown that a significant fraction of extracellular miRNAs reside outside vesicles and act in an exosome-independent manner, protected by RNA-binding proteins such as nucleophosmin 1 (NPM1) (Wang et al., 2010) or argonaute protein 2 (Ago2), (Turchinovich et al., 2011). Whether circulating miRNAs are found in soluble free form or are predominantly transported via secreted microvesicles/exosomes is still unclear, since evidence has been found for both mechanisms. However, it has recently been reported that the form of delivery of circulating miRNAs could depend on the type of tissue injury, suggesting a different role for each mode of systemic transport. Bala and co-workers documented this mechanism for circulating inflamma-miRs using different liver disease mouse models: in inflammatory liver injury and alcoholic liver disease (ALD) serum/plasma miR-122 and miR-155 were predominantly associated with the exosome-rich fraction, whereas in drug (acetaminophen, APAP)-induced liver injury (DILI/APAP) the same miRNAs were found mainly in protein-rich, soluble free form (Bala et al., 2012).
Evidence has also been found that blood cells are the major contributors to circulating miRNAs (Pritchard et al., 2012), However, analysis of inflamma-miRs from patients with acute coronary syndrome (ACS) documented that their levels in plasma and peripheral blood mononuclear cells (PBMCs) did not correlate in all subjects (Yao et al., 2011). This suggests that other cells may contribute to circulating inflamma-miRs. Endothelial cells could exert such a role, especially during normal aging and in age related-diseases. Indeed human endothelial cells and circulating progenitor endothelial cells can acquire the senescence-associated secretory phenotype (SASP) during replicative senescence in vitro (Olivieri et al., 2012b, 2013c). The accumulation of endothelial senescent cells in vivo, and stimulation of immune cells over time, could strongly contribute to promoting and maintaining low-level chronic systemic inflammation (Orjalo et al., 2009; Davalos et al., 2010; Campisi et al., 2011).
Circulating Inflamma-miRs in Aging
A progressive increase in circulating acute-phase proteins and proinflammatory mediators, i.e., proteases, cytokines, chemokines, and growth factors, has been described as a general feature of the aging process and has been denominated inflamm-aging (Franceschi et al., 2000). Age-associated chronic inflammation has mainly been attributed to progressive activation of immune cells over time and to accumulation of senescent cells with a proinflammatory secretory phenotype (Olivieri et al., 2013b). The complex inflamma-ging phenotype is the result of age-related cell/tissue adaptation and remodeling interacting with genetic/epigenetic factors.
Even though TLR family members do not show consistent age-dependent changes across model systems, there is evidence for impaired downstream signaling events during aging, including inhibition of positive effectors and activation of negative modulators of TLR signaling (Olivieri et al., 2013a). Therefore, during aging inflamma-miR levels progressively increase in order to stem the cell and tissue damage induced by the low-level chronic inflammation, also likely sustained by the cell senescence secretome (Murray and Smale, 2012).
We have recently described an increased expression of miR-146a in human umbilical vein endothelial cells (HUVECs) and in aortic and coronary endothelial cells (respectively HAECs and HCAECs) during replicative senescence, thus demonstrating that TLR/NF-κB activation and cell senescence can be modulated by the same miRNAs (Olivieri et al., 2012b). MiR-146a is highly expressed also in aged mice (Jiang et al., 2012). In addition, lack of response by aged mouse macrophages to stimulation with LPS and proinflammatory cytokines indicates interruption of the negative feedback loop of miR-146a (Jiang et al., 2012). Altogether, these data lend support to the hypothesis that cellular senescence and TLR signaling activation may be closely interconnected and share common regulators.
Given the involvement of miRNAs in gene expression regulation, a peculiar modulation of their expression might contribute to efficient homeostasis in human aging. It is worth stressing that exceptionally long survival requires dynamic preservation of optimal levels of physiological variables, and that the mean levels of many biomarkers of aging are not stable, but change in the course of life (Spazzafumo et al., 2013). Only four studies have compared the miRNA expression profile of centenarians and younger subjects. They have shown a significant overlap between the miRNA profiles of centenarians and young individuals and a different profile in octogenarians, supporting the hypothesis that achievement of extreme longevity probably requires a special gene expression regulation (ElSharawy et al., 2012; Gombar et al., 2012; Olivieri et al., 2012a; Serna et al., 2012). Interestingly, all these studies showed miR-21 deregulation in centenarians compared with younger subjects. Age-related changes in the expression of miR-21 and miR-21* have recently been reported also in mouse heart, the major changes occurring from middle to old age (Zhang et al., 2012).
Modulation of miR-21 expression in plasma, circulating cells, and tissues of very old subjects and animals is not surprising to those who believe that miR-21 lies at the intersection of senescence, inflammation, and age-related diseases. Our group showed a positive correlations between circulating miR-21 and two important biomarkers of inflammation: C-reactive protein (CRP) and fibrinogen (Olivieri et al., 2012a,b). These data suggest that centenarians may have a better balance of their systemic inflammatory status compared with elderly subjects. Notably, we also found age-related changes in circulating miR-146a levels that were quite similar to those described for miR-21 (Olivieri et al., 2012a,b, our unpublished data). Interestingly, analysis of global miRNA expression in the peripheral blood of adult women has shown that miR-155 is one of the most up-regulated miRNAs among older women (Sredni et al., 2011).
Circulating Inflamma-miRs in Age-Related Diseases
Abundant data continue to support the hypothesis that progressive up-regulation of inflammatory gene expression and high levels of inflammatory signaling facilitate the development and progression of the major age-related diseases, such as cardiovascular disease (CVD), type 2 diabetes mellitus (T2DM), Alzheimer Disease (AD), rheumatoid arthritis (RA), and cancers. Patients suffering from such diseases show subclinical/clinical inflammation and, interestingly, deregulation of most circulating inflamma-miRs (Wang et al., 2012). Since multiple co-expressed miRNAs can cooperatively regulate a given biological process by targeting common components of that process, the development and progression of human diseases could be associated with abnormal regulation of multiple miRNAs functioning cooperatively. Some of the most recent data showing that cellular and circulating inflamma-miR deregulation is shared by the major human age-related diseases are summarized below.
Inflamma-miRs and T2DM
Recently, some of the miRNAs that we have designated inflamma-miRs, including miR-21 and miR-146a, were identified as novel players in β-cell failure elicited by proinflammatory cytokines in both in vitro and in vivo animal models (Roggli et al., 2010). Reduced miR-146a levels were also reported in PBMCs from Asian Indian patients with T2DM, in association with insulin resistance, poor glycaemia control, specific proinflammatory cytokine gene variants, and high levels of plasma TNFα and IL-6 (Balasubramanyam et al., 2011). Importantly, glucose concentrations were also reported to correlate negatively with circulating miR-126 in endothelial apoptotic bodies (Zampetaki et al., 2010); similarly, in patients with diabetes the reduction in miR-126 was seen to be confined to circulating vesicles in plasma (Zampetaki et al., 2010). Interestingly, miR-126 can modulate endothelial cell activation in response to systemic inflammatory stimuli, and can down-regulate the expression of IKBα, an important inhibitor of NF-κB signaling (Asgeirsdóttir et al., 2012; Feng et al., 2012).
Inflamma-miRs and CVD
In patients with CVD activation of innate immunity leads to an acute inflammatory reaction. There is evidence that inflamma-miR may contribute to the development/restraint of this inflammatory response: circulating inflamma-miRs could thus be clinically relevant diagnostic/prognostic biomarkers in CVD patients. MiR-21 plays important roles in cardiovascular and pulmonary disorders, including cardiac and pulmonary fibrosis and myocardial infarction, and also regulates various immunological and developmental processes (Kumarswamy et al., 2011).
We recently reported significantly higher circulating miR-21 and miR-146a levels in elderly patients with acute myocardial infarction (AMI) and/or heart failure compared with healthy subjects (Olivieri et al., 2012c). Circulating miR-155 were higher in those post-AMI patients who experienced cardiac death within 1 year (Matsumoto et al., 2012). Elevated miR-155 levels are also found in proinflammatory macrophages and atherosclerotic lesions, even though the effects of miR-155 seem to be different in early vs. advanced atherosclerosis (Wei et al., 2013). Recent studies suggest that miRNA deregulation may limit cardiovascular repair responses and result in an altered function and differentiation of cardiovascular progenitor cells and endothelial progenitor cells (EPCs), modulating endothelial regeneration and cardiomyocyte homeostasis and playing a crucial role in CVD (Jakob and Landmesser, 2012). Aging-associated senescence results in reduced EPC number and function, contributing to enhanced cardiac risk, reduced angiogenic capacity, and impaired cardiac repair effectiveness. Mounting evidence supports a role for miRNAs in vascular homeostasis, and miR-21 was found to regulate EPC senescence (Zhu et al., 2013).
Overall, circulating miR-146a, miR-155, and miR-21 are up-regulated in plasma of CVD patients.
Inflamma-miRs and AD
Increasing evidence supports a major role for TLRs in brain injury and their involvement in neurodegenerative disorders including AD. Components of this inflammatory pathway are known to contribute to AD, in part through overexpression of IL-1α and promotion of 42-amino acid amyloid β 42 (Aβ42) peptide generation; in turn IL-1α and amyloid β induce transcription of the proinflammatory prostaglandin synthase cyclooxygenase-2 (COX-2) gene and stimulate apoptotic brain cell death and neural tissue degeneration (Mrak and Griffin, 2001). Consistent with these findings many inflamma-miRs, including miR-9, miR-146a, and miR-155, are strongly expressed in human cerebrospinal fluid (CSF)- and brain tissue-derived extracellular fluid (ECF) from AD patients and are significantly up-regulated compared with age-matched controls, suggesting that they may be involved in modulation or promotion of miRNA-triggered pathogenic signaling throughout the brain and the CNS (Alexandrov et al., 2012). In AD patients miR-125b, miR-146a, and miR-155 have been shown to down-regulate complement Factor H (CFH), an important repressor of innate immunity acting on the cerebral inflammation response (Lukiw and Alexandrov, 2012; Lukiw et al., 2012a).
MiR-146a is among the more extensively investigated miRNAs in AD: it is up-regulated in response to IL-1, Aβ42 and oxidative stress in cultured human neuronal glial cells (Lukiw et al., 2008; Cui et al., 2010; Holohan et al., 2012; Lukiw and Alexandrov, 2012). In line with these data it is over-expressed in temporal cortices and in Aβ42-stressed human astroglial cells, where it down-regulates IRAK1 while inducing compensatory up-regulation of IRAK2 (Cui et al., 2010).
Interestingly, miR-146a and miR-155 were detected in the secretion of stressed human primary neural cells, and the conditioned medium containing miR-146a and miR-155 was found to induce Alzheimer-type gene expression changes in control brain cells (Lukiw et al., 2012b). These data suggest that paracrine transfer of genetic information between cells—either within the local brain environment or in the cerebrospinal or systemic circulation—may be the source of both beneficial and detrimental signals that further modulate the amyloidogenic, inflammatory, or neurotrophic aspects of the AD process (Lukiw, 2012).
Further confirmation that inflammatory pathways can contribute to AD development comes from the recent identification of a role for triggering receptor expressed on myeloid cells 2 (TREM2) in modulating the risk of AD onset (Guerreiro et al., 2013; Jonsson et al., 2013). TREM2 is an innate immune receptor expressed on the cell surface of microglia, macrophages, immature dendritic cells, and white matter in the hippocampus and neocortex, two areas that partially overlap with those affected by AD (Jiang et al., 2013). It controls two signaling pathways, one enhancing phagocytosis and another suppressing inflammatory reactivity. Reduced TREM2 expression is associated with an increase in microgliosis and neurodegeneration, suggesting that it modulates AD by enhancing inflammation (Jiang et al., 2013).
Although the analysis of miRNAs in AD is a relatively new research area, the data reported so far strongly indicate that inflamma-miRs could play a key role also in this condition.
Inflamma-miRs and RA
Altered miRNA expression has been demonstrated in the inflamed joints of RA patients (Blüml et al., 2011). Interestingly, miR-155 up-regulation has been documented in synovial membrane and synovial fluid (SF) macrophages from RA patients in association with reduced expression of its target, SHIP-1, an inhibitor of inflammation (Kurowska-Stolarska et al., 2011). Moreover miR-155(−/−) mice show significantly reduced local bone destruction, attributed to reduced osteoclast generation, although the severity of the joint inflammation is similar to the one seen in wild-type mice. These data demonstrate that miR-155 is critically involved in the adaptive and innate immune reactions leading to autoimmune arthritis (Blüml et al., 2011). In addition up-regulation of miR-146a, detected in PBMCs from RA patients, suggests that this miRNA could be a useful marker of disease activity (Abou-Zeid et al., 2011; Xie et al., 2013). Interestingly, circulating miR-21 is increased in the blood of patients with systemic lupus erythematosus (SLE) or RA, whereas circulating miR-146a and miR-155 show a trend toward significantly reduced levels only in SLE (Carlsen et al., 2013).
Inflamma-miRs and Cancers
MiRNAs are increasingly being recognized as oncogenes or oncosuppressors, since they contribute to cell transformation. Solid tumors are infiltrated by different stromal cells, e.g., fibroblasts, endothelial cells, lymphocytes and macrophages, that strongly influence neoplastic processes through a continuous heterotypic cross-talk (Squadrito et al., 2013). It is therefore not surprising that inflamma-miRs have been recognized as modulators of cancer development and progression (Williams et al., 2008). The contribution of miR-146a deregulation has been widely documented (Hurst et al., 2009; Li et al., 2010; Labbaye and Testa, 2012). However, its mechanism of action remains elusive, since both raised and decreased levels have been described depending on the type of cancer (Williams et al., 2008). Since miR-146a acts as a negative feedback loop of inflammation, dynamic changes in its expression can be expected in cancer tissues depending on the context of the heterotypic cross-talk.
Interestingly, mice with miR-146a deletion spontaneously develop subcutaneous flank tumors (Zhao et al., 2011). MiR-146a has been reported to suppress metastatic activity (Hou et al., 2012; Hwang et al., 2012), in particular its up-regulation inhibits cancer cell invasion and metastasis in vitro and in vivo (Hou et al., 2012). Altogether these findings show that by counteracting the inflammatory state associated with cell senescence, miR-146a can exert a general tumor suppressing action by inhibiting cancer development and cancer cell invasion and metastasis.
MiR-21 is frequently up-regulated in cancer and is implicated in practically every stage of the cancer process: promotion of cell proliferation, invasion, and metastasis; genome instability and mutation; inflammation; replicative immortalization;abnormal metabolism; angiogenesis; evasion of apoptosis; immune destruction and growth suppression. This suggests that miR-21 is an oncogene with a key role in resisting programmed cell death in cancer cells (Buscaglia and Li, 2011). MiR-155 is also an established “oncomiR” in breast cancer and regulates several pro-oncogenic pathways including angiogenesis (Czyzyk-Krzeska and Zhang, 2013).
Conclusion
Recent data on inflammation-related miRNAs in normal and pathological aging outline a complex scenario characterized by an altered expression of specific miRNAs that we have named inflamma-miRs, which mainly target the TLRs/NF-κB pathway. Up-regulation of inflammation markers is a general feature of the aging process and has been named inflamm-aging. The inflamm-aging phenotype results from age-related cell and tissue adaptation/remodeling interacting with the genetic/epigenetic background. It is a complex phenotype involving not only innate but also adaptive immunity and affecting a range of tissues and organs such as gut, fat, liver, muscle and brain. Importantly, inflamm-aging appears to be accelerated in a variety of age-associated diseases. Tissue and circulating inflamma-miRs could contribute to restrain the activity of the senescent cell secretome and to check the destruction induced by activation of the inflammatory response (Murray and Smale, 2012; Olivieri et al., 2013a). Inflamma-miRs have been implicated in regulation of the immune and inflammatory response, and their abnormal expression may contribute to the low-level chronic inflammation that has been documented both in normal aging and in the major age-related diseases. Circulating inflamma-miRs could thus have diagnostic/prognostic relevance in human diseases, e.g., CVD, T2DM, AD, RA, and cancer, which share a common inflammatory background. Recent data show up-regulation of inflamma-miRs in the circulation of healthy elderly and old individuals: the increase is less pronounced in centenarians and greater in patients with CVD, AD, or cancer. Notably, T2DM patients show decreased levels of circulating inflamma-miRs, but these data need to be confirmed in patients with diabetic complications. It is conceivable that the main sources of circulating inflamma-miRs in aging and age-related diseases are immunity circulating/tissue cells and endothelial circulating/resident cells. Cell senescence and inflammatory stimulation can contribute to induce and perpetuate systemic inflammation over time, inducing up-regulation of inflamma-miRs to stem the excessive activation of inflammatory pathways.
Since this field of research is new and growing, it would not be surprising if in the near future novel miRNAs were recognized as fine tuners of inflammation and thus added to the list of inflamma-miRs.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors are grateful to Dr. Silvia Modena for the language revision (www.silviamodena.com). This work has received funding from Grande Oriente d'Italia (GOI), Massoneria Italiana, Collegio delle Marche; and Università Politecnica delle Marche to Antonio D. Procopio and Fabiola Olivieri.
References
Abou-Zeid, A., Saad, M., and Soliman, E. (2011). MicroRNA 146a expression in rheumatoid arthritis: association with tumor necrosis factor-alpha and disease activity. Genet. Test Mol. Biomarkers 15, 807–812. doi: 10.1089/gtmb.2011.0026
Alexandrov, P. N., Dua, P., Hill, J. M., Bhattacharjee, S., Zhao, Y., and Lukiw, W. J. (2012). MicroRNA (miRNA) speciation in Alzheimer's disease (AD) cerebrospinal fluid (CSF) and extracellular fluid (ECF). Int. J. Biochem. Mol. Biol. 3, 365–373.
Asgeirsdóttir, S. A., van Solingen, C., Kurniati, N. F., Zwiers, P. J., Heeringa, P., van Meurs, M., et al. (2012). MicroRNA-126 contributes to renal microvascular heterogeneity of VCAM-1 protein expression in acute inflammation. Am. J. Physiol. Renal Physiol. 302, F1630–F1639. doi: 10.1152/ajprenal.00400.2011
Bala, S., Petrasek, J., Mundkur, S., Catalano, D., Levin, I., Ward, J., et al. (2012). Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology 56, 1946–1957. doi: 10.1002/hep.25873
Balasubramanyam, M., Aravind, S., Gokulakrishnan, K., Prabu, P., Sathishkumar, C., Ranjani, H., et al. (2011). Impaired miR-146a expression links subclinical inflammation and insulin resistance in Type 2 diabetes. Mol. Cell. Biochem. 351, 197–205. doi: 10.1007/s11010-011-0727-3
Bazzoni, F., Rossato, M., Fabbri, M., Gaudiosi, D., Mirolo, M., Mori, L., et al. (2009). Induction and regulatory function of miR-9 in human monocytes and neutrophils exposed to proinflammatory signals. Proc. Natl. Acad. Sci. U.S.A. 106, 5282–5287. doi: 10.1073/pnas.0810909106
Blüml, S., Bonelli, M., Niederreiter, B., Puchner, A., Mayr, G., Hayer, S., et al. (2011). Essential role of microRNA-155 in the pathogenesis of autoimmune arthritis in mice. Arthritis Rheum. 63, 1281–1288. doi: 10.1002/art.30281
Boldin, M. P., and Baltimore, D. (2012). MicroRNAs, new effectors and regulators of NF-kB. Immunol. Rev. 246, 205–220. doi: 10.1111/j.1600-065X.2011.01089.x
Brase, J. C., Johannes, M., Schlomm, T., Falth, M., Haese, A., Steuber, T., et al. (2011). Circulating miRNAs are correlated with tumor progression in prostate cancer. Int. J. Cancer 128, 608–616. doi: 10.1002/ijc.25376
Breving, K., and Esquela-Kerscher, A. (2010). The complexities of microRNA regulation: mirandering around the rules. Int. J. Biochem. Cell Biol. 42, 1316–1329. doi: 10.1016/j.biocel.2009.09.016
Buscaglia, L. E., and Li, Y. (2011). Apoptosis and the target genes of microRNA-21. Chin. J. Cancer 30, 371–380.
Campisi, J., Andersen, J. K., Kapahi, P., and Melov, S. (2011). Cellular senescence: a link between cancer and age-related degenerative disease? Semin. Cancer Biol. 21, 354–359. doi: 10.1016/j.semcancer.2011.09.001
Carlsen, A. L., Schetter, A. J., Nielsen, C. T., Lood, C., Knudsen, S., and Voss, A. (2013). Circulating microRNA expression profiles associated with systemic lupus erythematosus. Arthritis Rheum. 65, 1324–1334. doi: 10.1002/art.37890
Ceppi, M., Pereira, P. M., Dunand-Sauthier, I., Barras, E., Reith, W., Santos, M. A., et al. (2009). MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells. Proc. Natl. Acad. Sci. U.S.A. 104, 2735–2740. doi: 10.1073/pnas.0811073106
Chen, X. M., Splinter, P. L., O'Hara, S. P., and LaRusso, N. F. (2007). A cellular micro-RNA, let-7i, regulates Toll-like receptor 4 expression and contributes to cholangiocyte immune responses against Cryptosporidium parvum infection. J. Biol. Chem. 282, 28929–28938. doi: 10.1074/jbc.M702633200
Chen, Y., Chen, J., Wang, H., Shi, J., Wu, K., Liu, S., et al. (2013). HCV-induced miR-21 contributes to evasion of host immune system by targeting MyD88 and IRAK1. PLoS Pathog. 9:e1003248. doi: 10.1371/journal.ppat.1003248
Chassin, C., Hempel, C., Stockinger, S., Dupont, A., Kübler, J. F., Wedemeyer, J., et al. (2012). MicroRNA-146a-mediated downregulation of IRAK1 protects mouse and human small intestine against ischemia/reperfusion injury. EMBO Mol. Med. 4, 1308–1319. doi: 10.1002/emmm.201201298
Collino, F., Deregibus, M. C., Bruno, S., Sterpone, L., Aghemo, G., Viltono, L., et al. (2010). Microvesicles derived from adult human bone marrow and tissue specific mesenchymal stem cells shuttle selected pattern of miRNAs. PLoS ONE 5:e11803. doi: 10.1371/journal.pone.0011803
Crone, S. G., Jacobsen, A., Federspiel, B., Bardram, L., Krogh, A., Lund, A. H., et al. (2012). microRNA-146a inhibits G protein-coupled receptor-mediated activation of NF-κB by targeting CARD10 and COPS8 in gastric cancer. Mol. Cancer 11, 71.
Cui, J. G., Li, Y. Y., Zhao, Y., Bhattacharjee, S., and Lukiw, W. J. (2010). Differential regulation of interleukin-1 receptor-associated kinase-1 (IRAK-1) and IRAK-2 by microRNA-146a and NF-kappaB in stressed human astroglial cells and in Alzheimer disease. J. Biol. Chem. 285, 38951–38960. doi: 10.1074/jbc.M110.178848
Czyzyk-Krzeska, M. F., and Zhang, X. (2013). MiR-155 at the heart of oncogenic pathways. Oncogene. doi: 10.1038/onc.2013.26. [Epub ahead of print].
Dai, L., Gu, L., and Di, W. (2012). MiR-199a attenuates endometrial stromal cell invasiveness through suppression of the IKKβ/NF-κB pathway and reduced interleukin-8 expression. Mol. Hum. Reprod. 18, 136–145. doi: 10.1093/molehr/gar066
Davalos, A. R., Coppe, J. P., Campisi, J., and Desprez, P. Y. (2010). Senescent cells as a source of inflammatory factors for tumor progression. Cancer Metastasis Rev. 29, 273–283. doi: 10.1007/s10555-010-9220–9
Ding, J., Huang, S., Wang, Y., Tian, Q., Zha, R., Shi, H., et al. (2013). Genome-wide screening revealed that miR-195 targets the TNF-α/NF-κB pathway by downregulating IKKα and TAB3 in hepatocellular carcinoma. Hepatology. doi: 10.1002/hep.26378. [Epub ahead of print].
Dunne, A., and O'Neill, L. (2005). Adaptor usage and Toll-like receptor signalling specificity. FEBS Lett. 579, 3330–3335. doi: 10.1016/j.febslet.2005.04.024
ElSharawy, A., Keller, A., Flachsbart, F., Wendschlag, A., Jacobs, G., Kefer, N., et al. (2012). Genome-wide miRNA signatures of human longevity. Aging Cell 11, 607–616. doi: 10.1111/j.1474-9726.2012.00824.x
Fabbri, M. (2012). TLRs as miRNA Receptors. Cancer Res. doi: 10.1158/0008-5472.CAN-12-3229. [Epub ahead of print].
Fabbri, M., Paone, A., Calore, F., Galli, R., Gaudio, E., Santhanam, R., et al. (2012). MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. Proc. Natl. Acad. Sci. U.S.A. 109, E2110–E2116. doi: 10.1073/pnas.1209414109
Feng, X., Wang, H., Ye, S., Guan, J., Tan, W., Cheng, S., et al. (2012). Up-regulation of microRNA-126 may contribute to pathogenesis of ulcerative colitis via regulating NF-kappaB inhibitor IκBα. PLoS ONE 7:e52782. doi: 10.1371/journal.pone.0052782
Franceschi, C., Bonafè, M., Valensin, S., Olivieri, F., De Luca, M., Ottaviani, E., et al. (2000). Inflamm-aging. An evolutionary perspective on immunosenescence. Ann. N.Y. Acad. Sci. 908, 244–254. doi: 10.1111/j.1749-6632.2000.tb06651.x
Guerreiro, R., Wojtas, A., Bras, J., Carrasquillo, M., Rogaeva, E., Majounie, E., et al. (2013). TREM2 variants in Alzheimer's disease. N. Engl. J. Med. 368, 117–127. doi: 10.1056/NEJMoa1211851
Gombar, S., Jung, H. J., Dong, F., Calder, B., Atzmon, G., Barzilai, N., et al. (2012). Comprehensive microRNA profiling in B-cells of human centenarians by massively parallel sequencing. BMC Genomics 13:353. doi: 10.1186/1471-2164-13-353.
Harris, T. A., Yamakuchi, M., Ferlito, M., Mendell, J. T., and Lowenstein, C. J. (2008). MicroRNA-126 regulates endothelial expression of vascular cell adhesion molecule 1. Proc. Natl. Acad. Sci. U.S.A. 105, 1516–1521. doi: 10.1073/pnas.0707493105
Holohan, K. N., Lahiri, D. K., Schneider, B. P., Foroud, T., and Saykin, A. J. (2012). Functional microRNAs in Alzheimer's disease and cancer: differential regulation of common mechanisms and pathway. Front. Genet. 3:323. doi: 10.3389/fgene.2012.00323
Hosoda, T., Zheng, H., Cabral-da-Silva, M., Sanada, F., Ide-Iwata, N., Ogórek, B., et al. (2011). Human cardiac stem cell differentiation is regulated by a mircrine mechanism. Circulation 123, 1287–1296. doi: 10.1161/CIRCULATIONAHA.110.982918
Hou, Z., Yin, H., Chen, C., Dai, X., Li, X., Liu, B., et al. (2012). microRNA-146a targets the L1 cell adhesion molecule and suppresses the metastatic potential of gastric cancer. Mol. Med. Rep. 6, 501–506. doi: 10.3892/mmr.2012.946
Hurst, D. R., Edmonds, M. D., Scott, G. K., Benz, C. C., Vaidya, K. S., and Welch, D. R. (2009). Breast cancer metastasis suppressor 1 up-regulates miR-146, which suppresses breast cancer metastasis. Cancer Res. 69, 1279–1283. doi: 10.1158/0008-5472.CAN-08–3559
Hwang, S. J., Seol, H. J., Park, Y. M., Kim, K. H., Gorospe, M., Nam, D. H., et al. (2012). MicroRNA-146a suppresses metastatic activity in brain metastasis. Mol. Cells 34, 329–334. doi: 10.1007/s10059-012-0171-6
Inukai, S., and Slack, F. (2013). MicroRNAs and the genetic network in aging. J. Mol. Biol. pii: S0022-2836(13)00039-9. doi: 10.1016/j.jmb.2013.01.023.
Iyer, A., Zurolo, E., Prabowo, A., Fluiter, K., Spliet, W. G., van Rijen, P. C., et al. (2012). MicroRNA-146a: a key regulator of astrocyte-mediated inflammatory response. PLoS ONE 7:e44789. doi: 10.1371/journal.pone.0044789
Jakob, P., and Landmesser, U. (2012). Role of microRNAs in stem/progenitor cells and cardiovascular repair. Cardiovasc. Res. 93, 614–622. doi: 10.1093/cvr/cvr311
Jiang, M., Xiang, Y., Wang, D., Gao, J., Liu, D., Liu, Y., et al. (2012). Dysregulated expression of miR-146a contributes to age-related dysfunction of macrophages. Aging Cell 11, 29–40. doi: 10.1111/j.1474-9726.2011.00757.x
Jiang, T., Yu, J. T., Zhu, X. C., and Tan, L. (2013). TREM2 in Alzheimer's disease. Mol. Neurobiol. doi: 10.1007/s12035-013-8424-8. [Epub ahead of print].
Jonsson, T., Stefansson, H., Steinberg, S., Jonsdottir, I., Jonsson, P. V., Snaedal, J., et al. (2013). Variant of TREM2 associated with the risk of Alzheimer's disease. N. Engl. J. Med. 368, 107–116. doi: 10.1056/NEJMoa1211103
Kawai, T., and Akira, S. (2011). Toll-like receptors and their crosstalk with other innate receptors in infection and immunity. Immunity 34, 637–650. doi: 10.1016/j.immuni.2011.05.006
Kim, S. W., Ramasamy, K., Bouamar, H., Lin, A. P., Jiang, D., and Aguiar, R. C. (2012). MicroRNAs miR-125a and miR-125b constitutively activate the NF-κB pathway by targeting the tumor necrosis factor alpha-induced protein 3 (TNFAIP3, A20). Proc. Natl. Acad. Sci. U.S.A. 109, 7865–7870. doi: 10.1073/pnas.1200081109
Kondo, T., Kawai, T., and Akira, S. (2012). Dissecting negative regulation of Toll-like receptor signaling. Trends Immunol. 33, 449–458. doi: 10.1016/j.it.2012.05.002
Kumar, M., Ahmad, T., Sharma, A., Mabalirajan, U., Kulshreshtha, A., Agrawal, A., et al. (2011). Let-7 microRNA-mediated regulation of IL-13 and allergic airway inflammation. J. Allergy Clin. Immunol. 128, 1077–1085.e1–e10. doi: 10.1016/j.jaci.2011.04.034
Kumarswamy, R., Volkmann, I., and Thum, T. (2011). Regulation and function of miRNA-21 in health and disease. RNA Biol. 8, 706–713. doi: 10.4161/rna.8.5.16154
Kurowska-Stolarska, M., Alivernini, S., Ballantine, L. E., Asquith, D. L., Millar, N. L., Gilchrist, D. S., et al. (2011). MicroRNA-155 as a proinflammatory regulator in clinical and experimental arthritis. Proc. Natl. Acad. Sci. U.S.A. 108, 11193–11198. doi: 10.1073/pnas.1019536108
Labbaye, C., and Testa, U. (2012). The emerging role of MIR-146A in the control of hematopoiesis, immune function and cancer. J. Hematol. Oncol. 5, 13. doi: 10.1186/1756-8722-5–13
Li, Y., Vandenboom, T. G. 2nd, Wang, Z., Kong, D., Ali, S., Philip, P. A., et al. (2010). miR-146a suppresses invasion of pancreatic cancer cells. Cancer Res. 70, 1486–1495. doi: 10.1158/0008-5472.CAN-09–2792
Liang, D., Gao, Y., Lin, X., He, Z., Zhao, Q., Deng, Q., et al. (2011). A human herpesvirus miRNA attenuates interferon signalling and contributes to mainteinanceof viral latency by targeting IKK epsilon. Cell Res. 21, 793–806. doi: 10.1038/cr.2011.5
Liu, J., Mao, Q., Liu, Y., Hao, X., Zhang, S., and Zhang, J. (2013). Analysis of miR-205 and miR-155 expression in the blood of breast cancer patients. Chin. J. Cancer Res. 25, 46–54. doi: 10.3978/j.issn.1000-9604.2012.11.04
Liu, Z., Lu, C. L., Cui, L. P., Hu, Y. L., Yu, Q., Jiang, Y., et al. (2012). MicroRNA-146a modulates TGF-β1-induced phenotypic differentiation in human dermal fibroblasts by targeting SMAD4. Arch. Dermatol. Res. 304, 195–202. doi: 10.1007/s00403-011-1178–0
Lukiw, W. J., Zhao, Y., and Cui, J. G. (2008). An NF-κB-sensitive micro RNA-146a-mediated inflammatory circuit in Alzheimer disease and in stressed human brain cells. J. Biol. Chem. 283, 31315–31322. doi: 10.1074/jbc.M805371200
Lukiw, W. J. (2012). NF-κB-regulated, proinflammatory miRNAs in Alzheimer's disease. Alzheimers Res. Ther. 4, 47. doi: 10.1186/alzrt150
Lukiw, W. J., and Alexandrov, P. N. (2012). Regulation of complement factor H (CFH) by multiple miRNAs in Alzheimer's disease (AD) brain. Mol. Neurobiol. 46, 11–19. doi: 10.1007/s12035-012-8234-4
Lukiw, W. J., Andreeva, T. V., Grigorenko, A. P., and Rogaev, E. I. (2012a). Studying micro RNA function and dysfunction in Alzheimer's disease. Front. Genet. 3:327. doi: 10.3389/fgene.2012.00327
Lukiw, W. J., Alexandrov, P. N., Zhao, Y., Hill, J. M., and Bhattacharjee, S. (2012b). Spreading of Alzheimer's disease inflammatory signaling through soluble micro-RNA. Neuroreport 23, 621–626. doi: 10.1097/WNR.0b013e32835542b0
Mar-Aguilar, F., Mendoza-Ramírez, J. A., Malagón-Santiago, I., Espino-Silva, P. K., Santuario-Facio, S. K., Ruiz-Flores, P., et al. (2013). Serum circulating microRNA profiling for identification of potential breast cancer biomarkers. Dis. Markers 34, 163–169. doi: 10.3233/DMA-120957
Matsumoto, S., Sakata, Y., Nakatani, D., Suna, S., Mizuno, H., Shimizu, M., et al. (2012). A subset of circulating microRNAs are predictive for cardiac death after discharge for acute myocardial infarction. Biochem. Biophys. Res. Commun. 427, 280–284. doi: 10.1016/j.bbrc.2012.09.039
Mrak, R. E., and Griffin, W. S. (2001). Interleukin-1, neuroinflammation, and Alzheimer's disease. Neurobiol. Aging 22, 903–908. doi: 10.1016/S0197-4580(01)00287-1
Murray, P. J., and Smale, S. T. (2012). Restraint of inflammatory signaling by interdependent strata of negative regulatory pathways. Nat. Immunol. 13, 916–924. doi: 10.1038/ni.2391
Nazari-Jahantigh, M., Wei, Y., Noels, H., Akhtar, S., Zhou, Z., Koenen, R. R., et al. (2012). MicroRNA-155 promotes atherosclerosis by repressing Bcl6 in macrophages. J. Clin. Invest. 122, 4190–4202. doi: 10.1172/JCI61716
Nilsen, T. W. (2007). Mechanisms of microRNA-mediated gene regulation in animal cells. Trends Genet. 23, 243–249. doi: 10.1016/j.tig.2007.02.011
O'Connell, R. M., Chaudhuri, A. A., Rao, D. S., and Baltimore, D. (2009). Inositol phosphatase SHIP1 is a primary target of miR-155. Proc. Natl. Acad. Sci. U.S.A. 106, 7113–7118. doi: 10.1073/pnas.0902636106
Olarerin-George, A. O., Anton, L., Hwang, Y. C., Elovitz, M. A., and Hogenesch, J. B. (2013). A functional genomics screen for microRNA regulators of NF-kappaB signaling. BMC Biol. 11:19. doi: 10.1186/1741-7007-11-19
Olivieri, F., Rippo, M. R., Prattichizzo, F., Babini, L., Graciotti, L., Recchioni, R., et al. (2013a). Toll like receptor signaling in “inflammaging”: microRNA as new players. Immun. Ageing 10, 11. doi: 10.1186/1742-4933-10-11
Olivieri, F., Recchioni, R., Marcheselli, F., Abbatecola, A. M., Santini, G., Borghetti, G., et al. (2013b). Cellular senescence in cardiovascular diseases: potential age-related mechanisms and implications for treatment. Curr. Pharm. Des. 19, 1710–1719.
Olivieri, F., Antonicelli, R., Recchioni, R., Mariotti, S., Marcheselli, F., Lisa, R., et al. (2013c). Telomere/telomerase system impairment in circulating angiogenic cells of geriatric patients with heart failure. Int. J. Cardiol. 164, 99–105. doi: 10.1016/j.ijcard.2011.06.091
Olivieri, F., Spazzafumo, L., Santini, G., Lazzarini, R., Albertini, M. C., Rippo, M. R., et al. (2012a). Age-related differences in the expression of circulating microRNAs: miR-21 as a new circulating marker of inflammaging. Mech. Ageing Dev. 133, 675–685. doi: 10.1016/j.mad.2012.09.004
Olivieri, F., Lazzarini, R., Recchioni, R., Marcheselli, F., Rippo, M. R., Di Nuzzo, S., et al. (2012b). MiR-146a as marker of senescence-associated pro-inflammatory status in cells involved in vascular remodelling. Age (Dordr.). doi: 10.1007/s11357-012-9440-8. [Epub ahead of print].
Olivieri, F., Antonicelli, R., Lorenzi, M., D'Alessandra, Y., Lazzarini, R., Santini, G., et al. (2012c). Diagnostic potential of circulating miR-499-5p in elderly patients with acute non ST-elevation myocardial infarction. Int. J. Cardiol. doi: 10.1016/j.ijcard.2012.01.075. [Epub ahead of print].
Orjalo, A. V., Bhaumik, D., Gengler, B. K., Scott, G. K., and Campisi, J. (2009). Cell surface-bound IL-1alpha is an upstream regulator of the senescence-associated IL-6/IL-8 cytokine network. Proc. Natl. Acad. Sci. U.S.A. 106, 17031–17036. doi: 10.1073/pnas.0905299106
Pigati, L., Yaddanapudi, S. C., Iyengar, R., Kim, D. J., Hearn, S. A., Danforth, D., et al. (2010). Selective release of microRNA species from normal and malignant mammary epithelial cells. PLoS ONE 5:e13515. doi: 10.1371/journal.pone.0013515
Pritchard, C. C., Kroh, E., Wood, B., Arroyo, J. D., Dougherty, K. J., Miyaji, M. M., et al. (2012). Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer Prev. Res. (Phila.) 5, 492–497. doi: 10.1158/1940-6207.CAPR-11–0370
Quinn, S., and O'Neill, L. A. (2011). A trio of microRNAs that control toll-like receptor signalling. Int. Immunol. 23, 421–425. doi: 10.1093/intimm/dxr034
Reid, G., Kirschner, M. B., and van Zandwijk, N. (2011). Circulating microRNAs: association with disease and potential use as biomarkers. Crit. Rev. Oncol. Hematol. 80, 193–208. doi: 10.1016/j.critrevonc.2010.11.004
Rodriguez, A., Vigorito, E., Clare, S., Warren, M. V., Couttet, P., Soond, D. R., et al. (2007). Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611. doi: 10.1126/science.1139253
Roggli, E., Britan, A., Gattesco, S., Lin-Marq, N., Abderrahmani, A., Meda, P., et al. (2010). Involvement of microRNAs in the cytotoxic effects exerted by proinflammatory cytokines on pancreatic beta-cells. Diabetes 59, 978–986. doi: 10.2337/db09–0881
Schulte, L. N., Westermann, A. J., and Vogel, J. (2013). Differential activation and functional specialization of miR-146 and miR-155 in innate immune sensing. Nucleic Acids Res. 41, 542–553. doi: 10.1093/nar/gks1030
Serna, E., Gambini, J., Borras, C., Mohammed, K., Belenguer, A., Sanchis, P., et al. (2012). Centenarians, but not octogenarians, up-regulate the expression of microRNAs. Sci. Rep. 2, 961. doi: 10.1038/srep00961
Sheddy, F. J., Palsson-McDermott, E., Hennessy, E. J., Martin, C., O'Leary, J. J., Ruan, Q., et al. (2010). Negative regulation of TLR4 via targeting of the proinflammatory tumor suppressor PDCD4 by the microRNA miR-21. Nat. Immunol. 11, 141–147. doi: 10.1038/ni.1828
Spazzafumo, L., Olivieri, F., Abbatecola, A. M., Castellani, G., Monti, D., Lisa, R., et al. (2013). Remodelling of biological parameters during human ageing: evidence for complex regulation in longevity and in type 2 diabetes. Age (Dordr.) 35, 419–429. doi: 10.1007/s11357-011-9348–8
Squadrito, M. L., Etzrodt, M., De Palma, M., and Pittet, M. J. (2013). MicroRNA-mediated control of macrophages and its implications for cancer. Trends Immunol. pii: S1471-4906(13)00029-X. doi: 10.1016/j.it.2013.02.003
Sredni, S. T., Gadd, S., Jafari, N., and Huang, C. C. (2011). A parallel study of mRNA and microRNA profiling of peripheral blood in young adult women. Front. Genet. 2:49. doi: 10.3389/fgene.2011.00049
Sun, Y., Cai, J., Ma, F., Lü, P., Huang, H., and Zhou, J. (2012). miR-155 mediates suppressive effect of progesterone on TLR3, TLR4-triggered immune response. Immunol. Lett. 146, 25–30. doi: 10.1016/j.imlet.2012.04.007
Taganov, K. D., Boldin, M. P., Chang, K. J., and Baltimore, D. (2006). NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signalling proteins of innate immune responses. Proc. Natl. Acad. Sci. U.S.A. 103, 12481–12486. doi: 10.1073/pnas.0605298103
Tomasetti, M., Staffolani, S., Nocchi, L., Neuzil, J., Strafella, E., Manzella, N., et al. (2012). Clinical significance of circulating miR-126 quantification in malignant mesothelioma patients. Clin. Biochem. 45, 575–581. doi: 10.1016/j.clinbiochem.2012.02.009
Turchinovich, A., Weiz, L., Langheinz, A., and Burwinkel, B. (2011). Characterization of extracellular circulating microRNA. Nucleic Acids Res. 39, 7223–7233. doi: 10.1093/nar/gkr254
Valadi, H., Ekström, K., Bossios, A., Sjöstrand, M., Lee, J. J., and Lötvall, J. O. (2007). Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659. doi: 10.1038/ncb1596
Wahlgren, J., De L Karlson, T., Brisslert, M., Vaziri Sani, F., Telemo, E., Sunnerhagen, P., et al. (2012). Plasma exosomes can deliver exogenous short interfering RNA to monocytes and lymphocytes. Nucleic Acids Res. 40, e130. doi: 10.1093/nar/gks463
Wang, K., Zhang, S., Weber, J., Baxter, D., and Galas, D. J. (2010). Export of microRNAs and microRNA-protective protein by mammalian cells. Nucleic Acids Res. 38, 7248–7259. doi: 10.1093/nar/gkq601
Wang, L. L., Huang, Y., Wang, G., and Chen, S. D. (2012). The potential role of microRNA-146 in Alzheimer's disease: biomarker or therapeutic target. Med. Hypotheses 78, 398–401. doi: 10.1016/j.mehy.2011.11.019
Waters, P. S., McDermott, A. M., Wall, D., Heneghan, H. M., Miller, N., and Newell, J. (2012). Relationship between circulating and tissue microRNAs in a murine model of breast cancer. PLoS ONE 7:e50459. doi: 10.1371/journal.pone.0050459
Wei, Y., Nazari-Jahantigh, M., Neth, P., Weber, C., and Schober, A. (2013). MicroRNA-126, -145, and -155: a therapeutic triad in atherosclerosis. Arterioscler. Thromb. Vasc. Biol. 33, 449–454. doi: 10.1161/ATVBAHA.112.300279
Williams, A. E., Perry, M. M., Moschos, S. A., Larner-Svensson, H. M., and Lindsay, M. A. (2008). Role of miRNA-146a in the regulation of the innate immune response and cancer. Biochem. Soc. Trans. 36, 1211–1125. doi: 10.1042/BST0361211
Xie, Q., Wang, S. C., Zhong, J., and Li, J. (2013). MicroRNA-146a, a good biomarker, and potential therapeutic target for rheumatoid arthritis. Genet. Test Mol. Biomarkers 17, 91–92. doi: 10.1089/gtmb.2012.0333
Xu, C., Ren, G., Cao, G., Chen, Q., Shou, P., Zheng, C., et al. (2013). MiR-155 Regulates Immune modulatory properties of mesenchymal stem cells by targeting TAK1-binding protein 2. J. Biol. Chem. 288, 11074–11079. doi: 10.1074/jbc.M112.414862
Yao, R., Ma, Y., Du, Y., Liao, M., Li, H., Liang, W., et al. (2011). The altered expression of inflammation-related microRNAs with microRNA-155 expression correlates with Th17 differentiation in patients with acute coronary syndrome. Cell. Mol. Immunol. 8, 486–495. doi: 10.1038/cmi.2011.22
Yao, Q., Cao, S., Li, C., Mengesha, A., Kong, B., and Wei, M. (2011). Micro-RNA-21 regulates TGF-β-induced myofibroblast differentiation by targeting PDCD4 in tumor-stroma interaction. Int. J. Cancer 128, 1783–1792. doi: 10.1002/ijc.25506
Zampetaki, A., Kiechl, S., Drozdov, I., Willeit, P., Mayr, U., Prokopi, M., et al. (2010). Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ. Res. 107, 810–817. doi: 10.1161/CIRCRESAHA.110.226357
Zhang, X., Azhar, G., and Wei, J. Y. (2012). The expression of microRNA and microRNA clusters in the aging heart. PLoS ONE 7:e34688. doi: 10.1371/journal.pone.0034688
Zhao, J. L., Rao, D. S., Boldin, M. P., Taganov, K. D., O'Connell, R. M., and Baltimore, D. (2011). NF-kappaB dysregulation in microRNA-146a deficient mice drives the development of myeloid malignancies. Proc. Natl. Acad. Sci. U.S.A. 108, 9184–9189. doi: 10.1073/pnas.1105398108
Zhou, J., Wang, K. C., Wu, W., Subramaniam, S., Shyy, J. Y., Chiu, J. J., et al. (2011). MicroRNA-21 targets peroxisome proliferators-activated receptor-alpha in an autoregulatory loop to modulate flow-induced endothelial inflammation. Proc. Natl. Acad. Sci. U.S.A. 108, 10355–10360. doi: 10.1073/pnas.1107052108
Keywords: inflammation mediators, circulating miRs, age-related diseases, NF-κB signaling, cellular senescence
Citation: Olivieri F, Rippo MR, Procopio AD and Fazioli F (2013) Circulating inflamma-miRs in aging and age-related diseases. Front. Genet. 4:121. doi: 10.3389/fgene.2013.00121
Received: 29 March 2013; Accepted: 06 June 2013;
Published online: 26 June 2013.
Edited by:
Andrey Turchinovich, German Cancer Research Center, GermanyReviewed by:
Janaiah Kota, Indiana University School of Medicine, USAPhilippe Georgel, Strasbourg University, France
Evgeny I. Rogaev, University of Massachusetts Medical School, USA
Copyright © 2013 Olivieri, Rippo, Procopio and Fazioli. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in other forums, provided the original authors and source are credited and subject to any copyright notices concerning any third-party graphics etc.
*Correspondence: Fabiola Olivieri, Department of Clinical and Molecular Sciences, Università Politecnica delle Marche, Via Tronto 10/A, 60020 Ancona, Italy e-mail: f.olivieri@univpm.it