The Me31B DEAD-box helicase localizes to postsynaptic foci and regulates expression of a CaMKII reporter mRNA in dendrites of Drosophila olfactory projection neurons
- 1 Smurfit Institute of Genetics and Trinity College Institute for Neuroscience, Trinity College Dublin, Dublin, Ireland
- 2 Department of Electronic and Electrical Engineering, Trinity College Dublin, Dublin, Ireland
- 3 Department of Biological Sciences, University of Denver, Denver, CO, USA
- 4 Department of Molecular and Cellular Biology, University of Arizona, Tucson, AZ, USA
- 5 Howard Hughes Medical Institute, University of Arizona, Tucson, AZ, USA
mRNP granules at adult central synapses are postulated to regulate local mRNA translation and synapse plasticity. However, they are very poorly characterized in vivo. Here, in Drosophila olfactory synapses, we present early observations and characterization of candidate synaptic mRNP particles, one of which contains a widely conserved, DEAD-box helicase, Me31B. In Drosophilas, Me31B is required for translational repression of maternal and miRNA-target mRNAs. A role in neuronal translational control is primarily suggested by Me31B’s localization, in cultured primary neurons, to neuritic mRNP granules that contain: (i) various translational regulators; (ii) CaMKII mRNA; and (iii) several P-body markers including the mRNA hydrolases, Dcp1, and Pcm/Xrn-1. In adult neurons, Me31B localizes to P-body like cytoplasmic foci/particles in neuronal soma. In addition it is present to synaptic foci that may lack RNA degradative enzymes and localize predominantly to dendritic elements of olfactory sensory and projection neurons (PNs). MARCM clones of PNs mutant for Me31B show loss of both Me31B and Dcp1-positive dendritic puncta, suggesting potential interactions between these granule types. In PNs, expression of validated hairpin-RNAi constructs against Me31B causes visible knockdown of endogenous protein, as assessed by the brightness and number of Me31B puncta. Knockdown of Me31B also causes a substantial elevation in observed levels of a translational reporter of CaMKII, a postsynaptic protein whose mRNA has been shown to be localized to PN dendrites and to be translationally regulated, at least in part through the miRNA pathway. Thus, neuronal Me31B is present in dendritic particles in vivo and is required for repression of a translationally regulated synaptic mRNA.
Introduction
The formation of long-term memory (LTM) and the underlying persistent synaptic changes require protein synthesis at stimulated synapses (Steward and Schuman, 2001; Martin and Zukin, 2006; Sutton and Schuman, 2006; Costa-Mattioli et al., 2009; Richter and Klann, 2009). For this local translation, mRNAs have to be transported in a repressed state from soma to synapse, and then stored at sites where translational activation must occur. Several studies have implicated a role for ribonucleoprotein (RNP) particles in this process (Martin and Zukin, 2006; Sossin and DesGroseillers, 2006; Bramham and Wells, 2007; Banerjee et al., 2009; Mikl et al., 2010). RNP particles, which contain proteins involved in mRNA translation, transport, repression, and decay, can form large complexes that have been identified in neurons of different species by light microscopy (Krichevsky and Kosik, 2001; Barbee et al., 2006; Kiebler and Bassell, 2006; Cougot et al., 2008; Zeitelhofer et al., 2008; Miller et al., 2009; Tubing et al., 2010).
Staufen- and FMRP-containing RNPs in the neurites of cultured Drosophila neurons contain proteins found in P-bodies, somatic RNPs that are sites of mRNA degradation and translational control (Barbee et al., 2006; Parker and Sheth, 2007). The discovery that neuritic RNPs are related to P-bodies has suggested a hypothesis that P-body components may play an important role in the regulation of synaptic mRNAs and thereby in the regulation of synaptic plasticity (Barbee et al., 2006; Cougot et al., 2008). Consistent with this hypothesis, some components of these large neuritic mRNPs including Staufen, Pumilio, Gld-2 and FMRP are required in Drosophila for the formation of LTM (Dubnau et al., 2003; Bolduc et al., 2008; Kwak et al., 2008) and/or dendrite morphogenesis (Lee et al., 2003; Ye et al., 2004). However, for most granule proteins, their localization, traffic, and functions are poorly understood in mature nervous systems.
One such granule protein found in P-bodies, maternal RNA granules, and neuritic RNPs, is a DEAD-box helicase with conserved orthologs from S. cerevisiae (Dhh1), to mammals (RCK/DDX-6) (Navarro et al., 2001; Coller et al., 2002; Cougot et al., 2004; Weston and Sommerville, 2006). In yeast and mammalian cells, the protein is required not only for translational repression and mRNA decay (Coller et al., 2002; Minshall and Standart, 2004), but also for P-body formation (Andrei et al., 2005; Coller and Parker, 2005). In contrast to these findings, which point to function as a negative regulator of mRNAs, observations in the C. elegans germline and in P. falciparum, indicates that the orthologs CGH1/DOZI proteins may have additional roles in stabilizing subsets of mRNAs and protecting them from degradation (Mair et al., 2006; Boag et al., 2008).
The Drosophila homolog Me31B is associated with maternal sponge bodies where it appears to repress translation of many granule-associated mRNAs (Nakamura et al., 2001). Me31B is associated with dFMR1/FMRP in neuritic mRNPs, regulates dendritic morphogenesis in sensory neurons, and, like its human homolog RCK, is required for optimal miRNA function (Barbee et al., 2006; Chu and Rana, 2006). Together, these observations point to a role for Me31B in the repression of dendritically localized mRNAs, their aggregation into transport mRNPs, and in control of locally translated synaptic mRNAs such as CaMKII, which regulate synaptic plasticity (Mayford et al., 1996; Aakalu et al., 2001; Ashraf et al., 2006).
However, many aspects of the above model have not been experimentally tested. It remains to be shown that mRNP aggregates exist in differentiated adult synapses in vivo. Therefore the composition and characteristics of such mRNP particles is unclear. In addition how Me31B in particular, and neuronal mRNP components in general, regulate target dendritic mRNAs also remains largely unknown in vivo.
To address these outstanding questions, we studied the neuronal expression and localization of Me31B in the adult Drosophila brain and the effect of Me31B mutations on the expression of synaptic CaMKII reporter protein (Mayford et al., 1996; Aakalu et al., 2001; Ashraf et al., 2006). We chose to analyze neurons and synapses in the olfactory circuit, which is one of the best-understood and genetically accessible neuronal networks in the adult brain (Davis, 2005; Vosshall and Stocker, 2007; Fiala, 2008). Not only is it well characterized in terms of its connectivity and function, but also the plasticity of synapses on projection neurons (PNs), as well as local translation of CaMKII mRNA in PNs, has been associated with olfactory memory (Ashraf et al., 2006; Keene and Waddell, 2007; Fiala, 2008).
Our data show: (i) that Me31B is expressed widely in the nervous system; (ii) that it is present on a postsynaptic particle in vivo; and (iii) that Me31B is required in PNs for the repression of CaMKII reporter mRNA. These observations represent an early step in the process of linking translational control proteins, synaptic mRNP dynamics and local mRNA translation with synaptic plasticity in a well-characterized behavioral circuit.
Materials and Methods
Drosophila Stocks
Fly stocks were raised at 25°C on standard cornmeal and agar media. Wild-type stocks (Oregon-R), UAS-Me31B strains, me31BΔ2, FRT40A (Barbee et al., 2006), and UAS-myrGFPCaMKII3′UTR were from the Ramaswami stock collection. MZ-317-Gal4, tub-Gal80ts 2x, and GH146-Gal4 (on X) were from Liqun Luo, V. Rodrigues, and S. G. Gajendra (NCBS, Bangalore). NP3529-Gal4 was from Rachel Wilson and Kei Ito; NP225-Gal4 from Kei Ito; UAS-Me31B RNAi 4916R-1 (on X; Barbee et al., 2006) was from the NIG-FLY collection; UAS-Me31B RNAi 49379 (on 2nd; Dietzl et al., 2007) was from the VDRC stock collection; UAS-SytGFP, Or83B-Gal4, GH146-Gal4, UAS-mCD8-GFP, hsFLP, FRT40A, FRT82B, tubGal80, FRT82B and tubGal80, FRT40a were from the Bloomington Stock Centre.
Immunohistochemistry
Preparations of Drosophila adult brains were performed after R. Stocker (Stocker et al., 1990). Decapitated fly heads were fixed in 4% PFA in 1× PBS + 0.2% Triton for 3 h on ice, washed with 1× PBS + 0.2% Triton and brains were dissected in blocking solution (PBS + 0.2% Triton and 5% NGS). Brains were incubated with primary antibodies overnight on 4°C diluted in blocking solution, followed by incubation with secondary antibodies for 3 h at room temperature diluted in blocking solution.
For preparations of Drosophila antenna whole flies were fixed in 4% PFA in PBS + 3% Triton for 1 h on ice. Antenna were cut off and fixed for an additional 4 h on ice. The antenna were washed quickly with 1× PBS + 3% Triton for 3× 15 min, followed by a 2-day wash on 4°C. Incubation with primary antibodies in 1× PBS + 0.1% triton over night at 4°C and incubation with secondary antibody as well in 1× PBS + 0.1% triton over night at 4°C.
Primary antibodies used: mouse anti-Me31B (Nakamura et al., 2001), rabbit anti-GFP (Molecular Probes), goat anti-GFP FITC conjugated and chicken anti-GFP (Abcam), rabbit anti-Pacman (S. Newbury; Barbee et al., 2006; Hillebrand et al., 2007; Zabolotskaya et al., 2008), rabbit anti-Dcp1 (J. Wilhelm; Barbee et al., 2006; Hillebrand et al., 2007; Lin et al., 2008; Zabolotskaya et al., 2008), rabbit anti-Staufen (D. St Johnston; St Johnston et al., 1991), mouse anti-NC82 (Developmental Studies Hybridoma Bank), rabbit anti-CaMKII (L. Griffith; Koh et al., 1999), and mouse anti-CaMKII (S. Ohsako; Takamatsu et al., 2003). Secondary antibodies used: Alexa 488-, Alexa 555-, and Alexa 647-conjugated anti rabbit and anti mouse IgG (Molecular Probes).
Preparations were mounted in Vectashield Mounting Medium (Vecta Labs) and imaged on a Zeiss LSM 510-meta confocal microscope.
Image Acquisition
All images were taken on a Zeiss LSM 510 confocal microscope. To image mRNP particles in fixed brain tissue, we used a 63× objective (Zeiss Plan-Apochromat, 1.4 Oil Ph3) and the “digital zoom” software feature of the LSM 510 software (zoom factor between 3 and 4) that allows to scan a region of interest at a higher pixel resolution, constrained of course by the normal ∼300 nm resolution limit of light microscopy. Regions of particular interest were selected in Photoshop and presented in a way to allow a better evaluation by the reader.
MARCM
For the induction of MARCM clones Drosophila crosses of the respective genotype were raised at 25°C and the flies were transferred to a new vial every 4 h for timed egg collections. A heat-shock pulse at 37°C in a water bath for 1 h was given at optimal time points to induce single-cell clones in PNs projecting to the DL-1 glomerulus. The time points (between 0 and 60 h after larval hatching) were chosen after G. Jefferis (Jefferis et al., 2001). Heat-shocked larvae were transferred back to 25°C, raised to adulthood and the adult flies were dissected 1–4 days after eclosion.
Automated Spot Colocalization Analysis
We created “Spotnik” a MatLab plug-in which allowed us to automate the quantitative analysis of spot colocalization across red and green channels of a double-stained image. The details of the signal-processing algorithms used have been described elsewhere (Pan et al., 2010). However, the main steps and logic were as follows.
Spot region extraction
Given that spots can be seen in regions of the image with highly variable levels of local background labeling, a direct image thresholding tool does not allow spots to be identified. Thus, we created a spot extraction tool that sharpened and enhanced details of the image, using a Wiener filter coupled with a gamma operation, and then modified the contrast of the image such that local features have the same contrast independent of their local background. The image was now thresholded with a user-defined threshold, to create a mask that included regions well-contrasted from the background. This mask was then used to select regions of pixels from the original sharpened image for subsequent processing.
Spot detection and analysis
Individual regions defined by the mask are first normalized to remove the background (by fitting a local plane to the background). A subsequent modeling step identified Gaussian parameters for each spot in each identified region using a Split-Merge Estimation-Maximization technique (Ueda et al., 2000) and synthesized Gaussians at their modeled positions. The center of these spots and their variances (widths) are then recorded for analysis. At each step, we ensured that manually detected spots were not lost and that computationally detected spots were not artifactual.
Colocalization analysis
Several different types of colocalization analysis are possible. However, for each spot imaged in one spectral channel, a simple MatLab tool allowed us to compute the distance of the closest spot in the other channel. Plots were then created in which the percent of spots that have corresponding closest spots closer than a given distance was plotted against distance.
Results
Me31B is Expressed in the Adult Brain
The expression of neuronal Me31B/DDX6/RCK/Dhh1/Cgh1 and the composition of associated neuronal mRNPs has been previously examined in dendrites of primary cultured neurons (Barbee et al., 2006; Cougot et al., 2008; Zeitelhofer et al., 2008; Miller et al., 2009). In order to extend these studies to an in vivo system, we used confocal microscopy and immunohistochemistry to examine Me31B expression in adult Drosophila brains, focusing primarily on the well-studied olfactory system (Figure 1A).
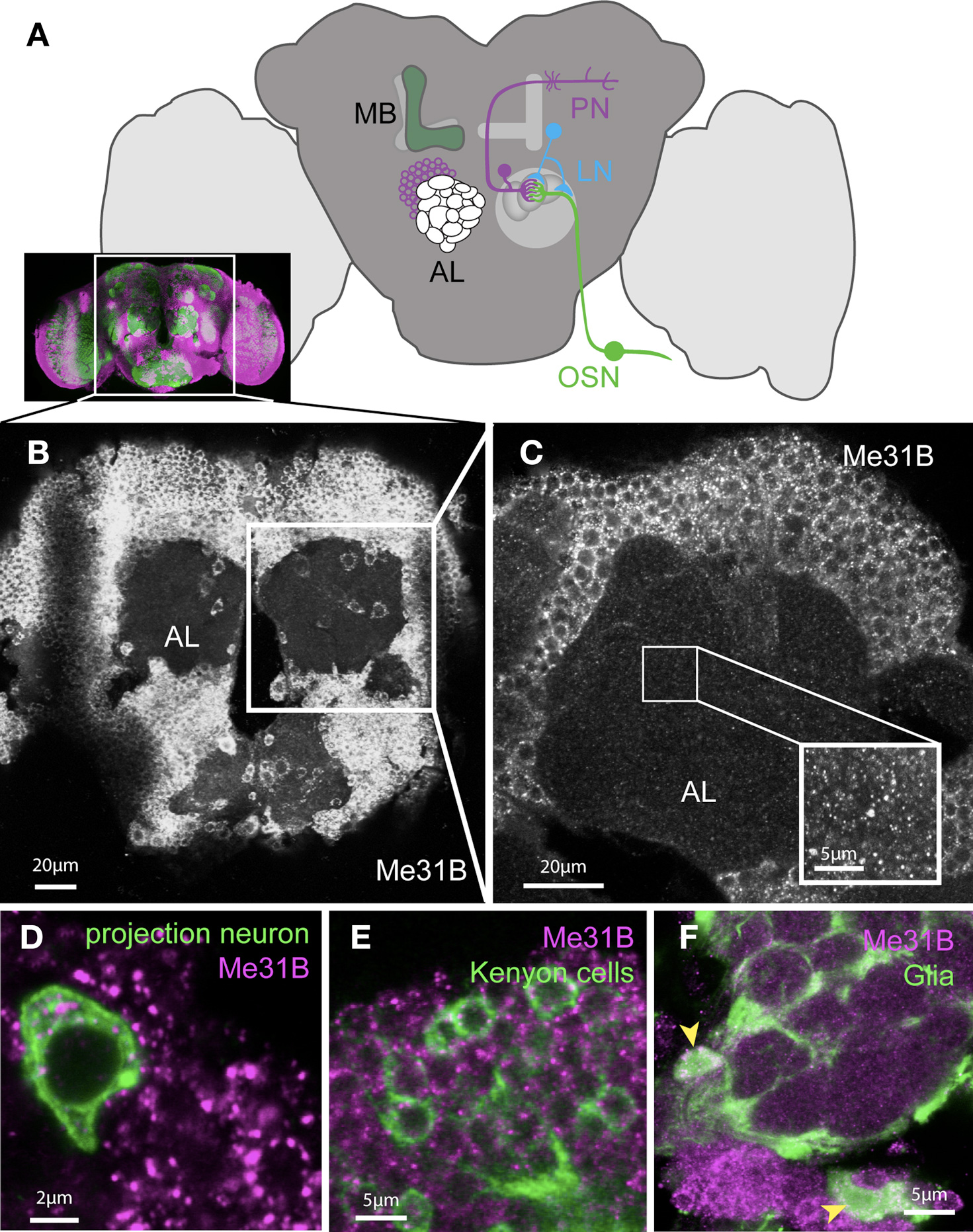
Figure 1. Me31B is ubiquitously expressed in the adult Drosophila brain. (A) Cartoon schematic illustrating the main components of the olfactory system in the adult Drosophila brain (the inset shows a real stained brain; co-immunostaining for the presynaptic marker Nc82 (green) and Me31B (magenta)). Sensory information from the antenna is sent to the brain via olfactory sensory neurons (OSNs) which synapse in the antennal lobes (AL) to dendrites of projection neurons (PNs). The axonal terminals of OSNs and dendrites of PNs form dense synaptic areas called glomeruli. PN axons project into higher brain regions and form synapses in the mushroom-body (MB) calyx and the lateral horn. Olfactory information is believed to be stored in the MB. The sensory information can be regulated in addition by local interneurons (LNs), which form synapses with PNs and OSNs. (B) Projection of confocal images of an adult Drosophila brain (showing the region of the brain indicated in the inset above) stained with an Me31B antibody, showing apparently uniform Me31B expression in neuronal cell bodies throughout the brain. (C) A single confocal slice at a higher magnification shows the punctate expression of Me31B in cell bodies and as well as fainter foci in the synaptic neuropil (AL); inset shows an example of Me31B expression in the neuropil in higher magnification (gain adjusted for better visualization of foci in the neuropil). Foci are absent in Me31B mutant tissue (Figures 6B,C,G). (D,E) Expression of Me31B (magenta) in genetically marked neural subpopulations: in (D) projection neurons; (E) mushroom-body Kenyon cells, as well as (F) glia cells (cell bodies marked by arrowheads). Single-cells, or small clones of the indicated cell type, are respectively marked by the expression of mCD8-GFP (green) in the pattern of specific Gal4 driver lines; (D,E) NP3529; (F) MZ-317 using the MARCM technique (Lee and Luo, 2001).
Me31B is ubiquitously expressed throughout the brain (Figure 1B). Closer examination of Me31B expression in the antennal lobes (AL) area showed punctate expression in cell bodies and in the neuropil (synaptic) area of the AL (Figure 1C). Combining anatomical information with genetic markers for specific neuronal cell types, we directly confirmed Me31B expression in olfactory PNs (Figure 1D), local interneurons (LNs), and mushroom-body (MB) Kenyon cells (Figure 1E). In addition to neurons, Me31B was also strongly expressed in glial cells, which sent processes into the AL (Figure 1F). In control experiments, we confirmed that the anti-Me31B antibodies did not label cell bodies or synapses of me31B null mutant neurons (see Figure 6; Figure S1 in Supplementary Material). Thus, Me31B is present in both somatic and synaptic particles.
We further attempted to examine the composition of these particles and their relationship to P-bodies, cytoplasmic mRNP aggregates that mediate specific forms of translational repression and mRNA decay (Eulalio et al., 2007a; Parker and Sheth, 2007).
Somatic and Synaptic Me31B Foci Differ in their Composition
In cultured Drosophila neurons, Me31B-positive mRNPs in neurites are related to P-bodies based on the high frequency at which they contain various P-body markers, including the RNA decapping enzymes Dcp1 and Pcm (the Drosophila homologue of the 5′–3′ RNA exonulease Xrn-1) (Barbee et al., 2006). We asked how these neuritic mRNPs compared with Me31B containing particles in vivo, by examining Me31B-positive foci in the adult Drosophila brain for presence of the P-body markers Dcp1 and Pcm as well as the neuronal granule/transport mRNP marker Staufen, which may also be a component of P-bodies (Anderson and Kedersha, 2006; Eulalio et al., 2007a). In neuronal cell bodies, double-immunostained to visualize Me31B and P-body markers, we observed that a very large fraction of the Me31B-positive, cytoplasmic foci also contained each of the three proteins (Figure 2). This indicates that the Me31B foci in neuronal soma are similar to P-bodies and to mRNP granules in neurites observed in primary cell culture (Barbee et al., 2006).
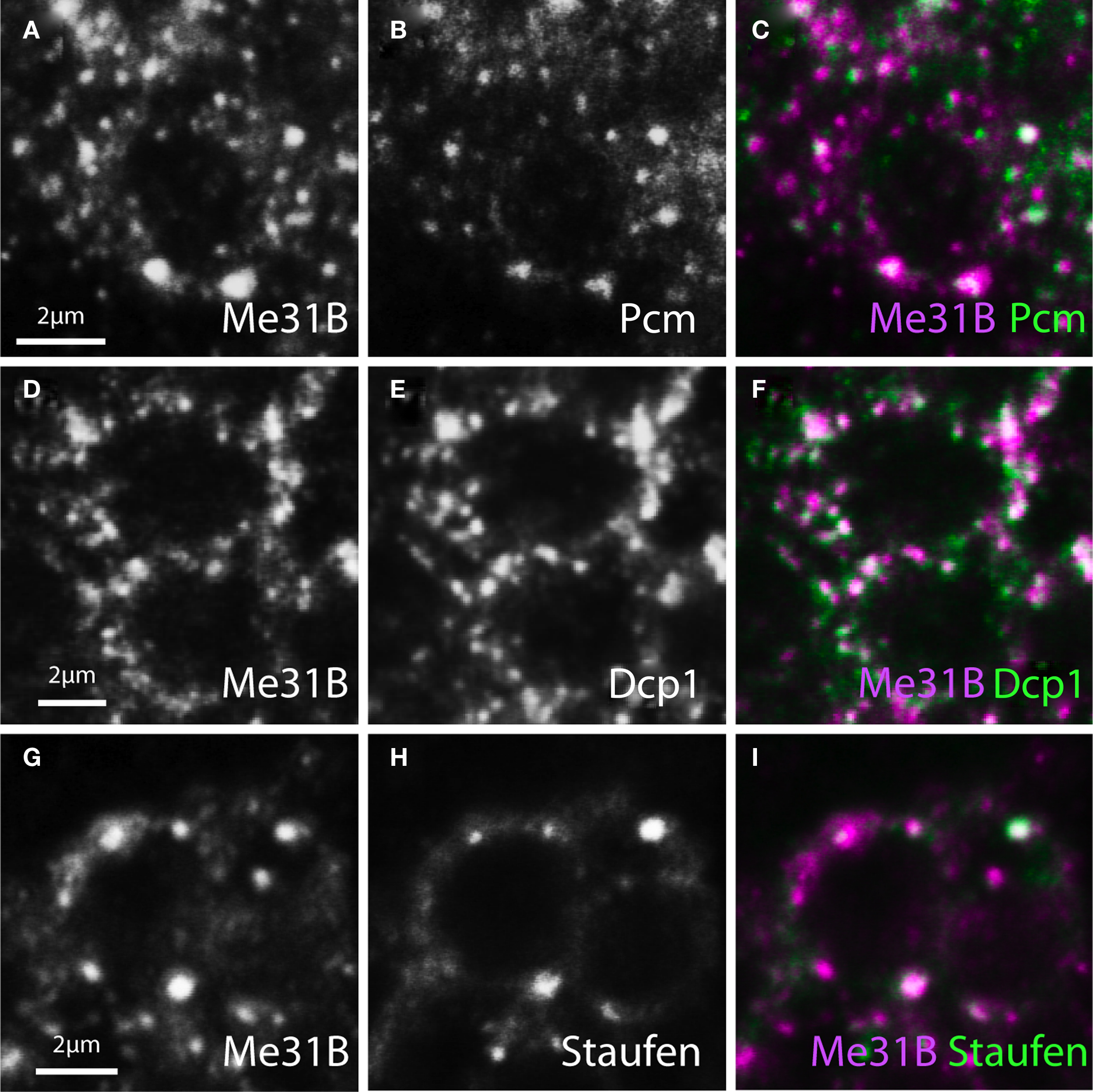
Figure 2. Me31B foci in neuronal soma contain Staufen and P-body associated mRNA hydrolases. Image pairs and merged channels of neuronal soma from adult brain, double-stained for Me31B (left panels or magenta) and three mRNP components (middle panels or green) show substantial colocalization. (A–C) Cells double-labeled for Me31B and Pacman (Pcm, the Drosophila homologue of the 5′–3′ RNA exonuclease Xrn1). (D–F) Neurons double-stained for Me31B and Dcp1 (an mRNA decapping factor). (G–I) Cells similarly double-labeled for Me31B and Staufen (a conserved mRNA binding protein). The determined levels of colocalization are presented in Figure 4.
In contrast to particles in the soma, Me31B foci in the synaptic neuropil areas appeared to be distinct and to only rarely contain the P-body markers tested (Figure 3). Thus, although Dcp1, Pcm, or Stau-labeled foci were present in the neuropil, these showed relatively weak colocalization with Me31B.

Figure 3. Synaptic Me31B foci largely exclude Pcm, Dcp1, and Staufen. Confocal images of olfactory glomeruli in the Drosophila antennal lobe (AL) double-stained for Me31B (left panels or magenta) and P-body markers (middle panels or green). In contrast to Me31B-containing foci in the soma (Figure 2), synaptic Me31B particles show only weak colocalization with P-body proteins. (A–C) Single confocal image pairs and overlay (right) of synaptic neuropil double-stained for Me31B and Pcm. (D–F) Similar confocal sections double-stained for Me31B and Dcp1; (G–I). Similar confocal sections of Me31B and Staufen foci in a double-stained glomerulus. Computationally analyzed levels of colocalization are presented Figure 4.
In order to rigorously assess levels of colocalization, we developed an algorithm, tentatively named Spotnik that allowed us to efficiently identify foci in one spectral channel and to measure the distance to the closest foci in the other spectral channel. Two key features of this program were: (a) local field thresholding, which allowed faint spots to be selected as efficiently as bright ones; and (b) Gaussian modeling of spots, which importantly allowed foci to be distinguished from shapeless or other local background signals. As can be seen from the image processing steps shown in Figure 4, the Gaussian modeling step also allowed two or three adjacent and overlapping foci to be resolved (see Materials and Methods; Pan et al., 2010). The distance between the centers of Gaussians (foci) in one channel to the closest Gaussian peak in the other were measured and the percentage of foci within a distance (x) of a foci in the other channel was plotted (Figures 4B–D).
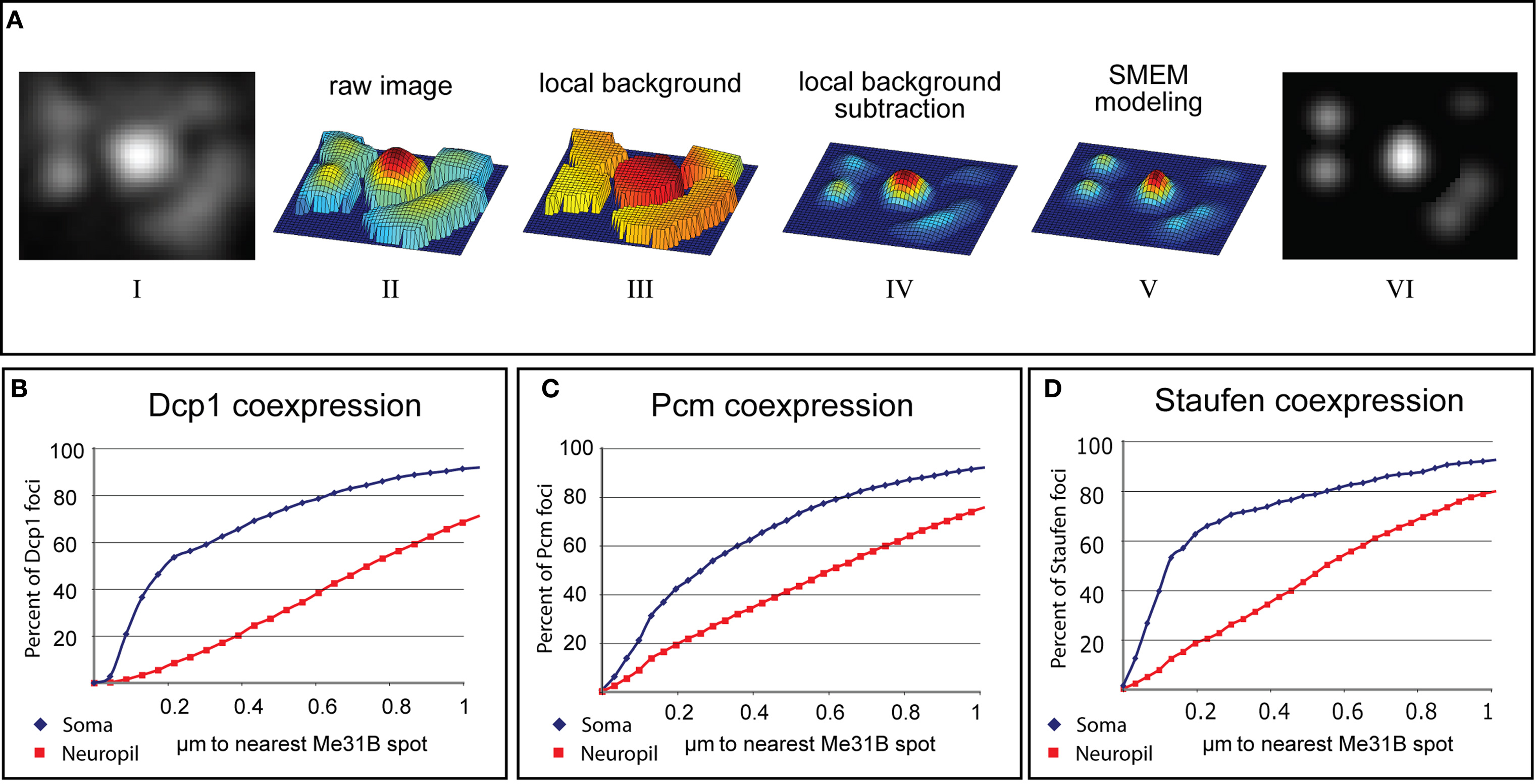
Figure 4. Computational analysis of Me31B colocalization with P-body proteins in somatic and synaptic foci. (A) Stages of image processing during automated detection of immunostained puncta (see Materials and Methods). The original image (i) is transformed into a mesh-plot (ii). The local background (iii) is subtracted from this image to yield image (iv). Each object in this background-subtracted image is fit to Gaussian parameters using an SMEM (Split-Merge Estimation-Maximization) technique and the result (v) is used as a mask to analyze underlying spots in the original image (vi). (B–D) Quantified colocalization of Dcp1, Pcm, and Staufen with Me31B. The graphs plot cumulative fraction of Dcp1, Pcm, or Stau positive particles found within a specific distance from the closest Me31B containing particle (fraction on the “y” axis is plotted versus distance on the “x”). Blue curves correspond to somatic puncta and red curves to synaptic ones (for each curve 20 images and about 5000 spots-per-channel, were analyzed). Distances are as measured by Spotnik from the computationally determined centroids for each spot, as described by the Gaussian modeling technique. However the actual position of these centroids are naturally limited by the resolution of light microscopy. Thus, spots separated by less than about 0.30 μ should be assumed to colocalize exactly. (B) Cumulative plot showing the fraction of Dcp1 foci (y) that are within (x) micrometers from an Me31B foci described using Gaussian modeling. The quick rise of the blue curve (somatic foci) compared to the red curve (synaptic foci) indicates substantially tighter colocalization of Me31B with P-body markers in somatic particles compared to synaptic ones. (C,D) Plots similar to (B) for Pcm and Staufen.
The rigorous colocalization analysis enabled by Spotnik confirmed that particles labeled with Me31B colocalized much more efficiently with Dcp1, Pcm, and Staufen immunostained foci in cell bodies than it did at synapses. For example, based on using a 200-nm cut-off for perfect colocalization, Me31B showed 58% colocalization with Dcp1 in soma, but only 7% at the synapse. Thus, the data indicate that Me31B is present in a class of synaptic particle that lacks RNA degradative enzymes, reminiscent of a class of storage mRNP described recently in the C. elegans germline (Boag et al., 2008; Gallo et al., 2008). Alternative interpretations of these data are considered in the Section “Discussion”.
Me31B Foci are Predominantly Dendritic In OSNs And PNs
The observations above suggest an obvious hypothesis in which synaptic Me31B foci represent sites where mRNAs are held repressed until released for stimulation-induced, local translation. To ask whether Me31B particles were enriched in either the presynaptic terminal or dendrites, we performed a series of experiments, which examined the localization of Me31B foci relative to synaptic element or cell-type specific markers. In some experiments, we used the nc82 antibody to label Bruchpilot (Brp), a protein specific to presynaptic active zones. In others, we used cell-type specific expression of either a synaptotagmin-GFP (Syt-GFP) fusion protein to label presynaptic terminals, or a membrane associated myristoylated-GFP under control of the 3′UTR of Drosophila CaMKII (myr-GFPCaMKII3′UTR) mRNA, shown to localize to dendrites (Ashraf et al., 2006). Figures 5A–C show that the latter reporter, when expressed in PNs, marks PN dendrites that show only minimal overlap with presynaptic elements in AL labeled with nc82.
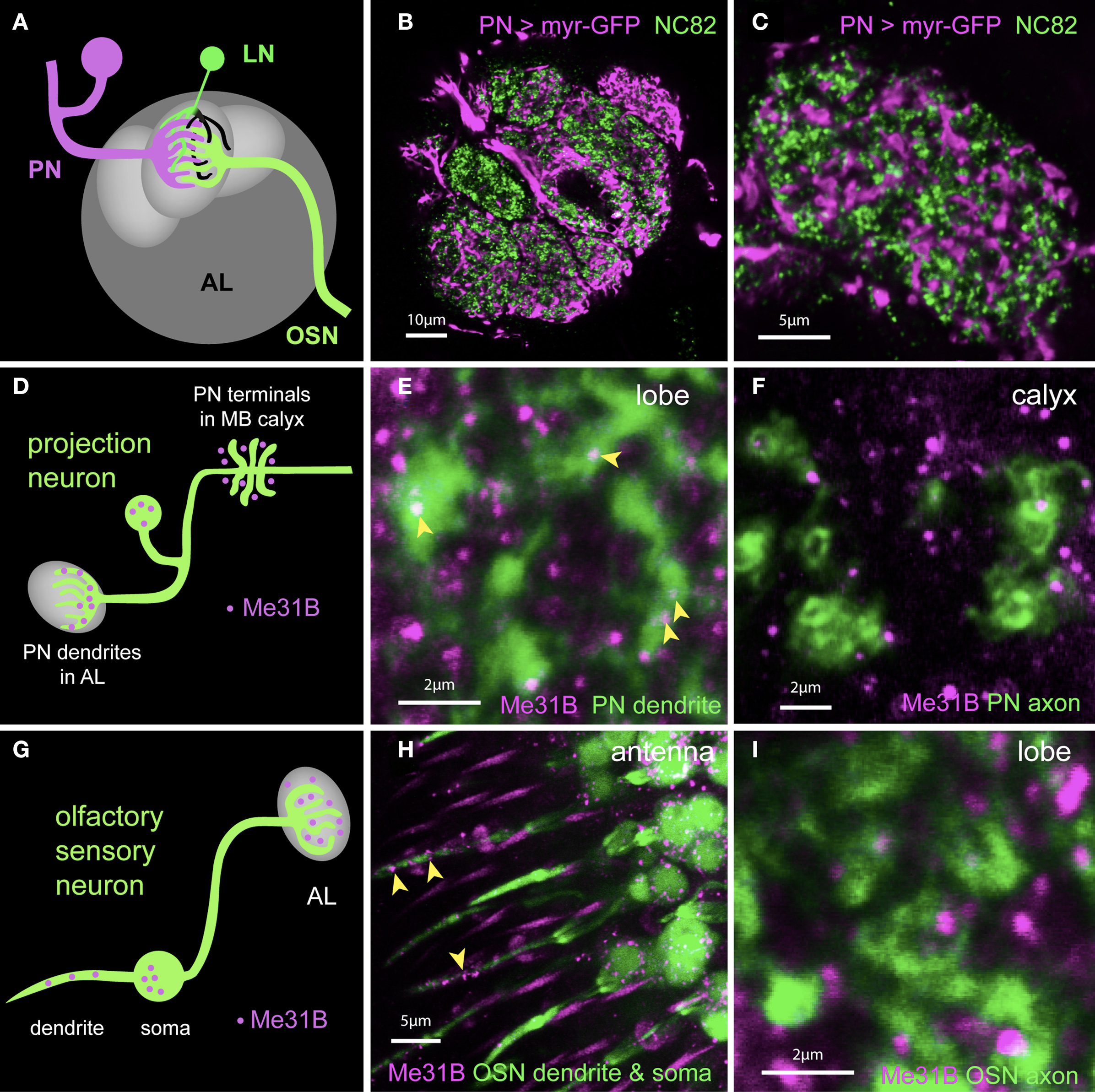
Figure 5. Me31B foci are highly enriched in postsynaptic dendrites in vivo. (A) Schematic showing OSNs (green) synapsing onto dendrites of PNs (magenta) and LNs (local interneurons in green and black) in a glomerulus. Presynaptic (green) as well as postsynaptic (black) elements of LNs are present in the glomerulus. (B) Image of a single confocal slice of the antennal lobe; Within a single glomerulus, presynaptic active zones labeled by an anti-Bruchpilot (Brp) antibody (nc82, green) lie in spatial domains distinct from those of PN dendrites (magenta) marked by expression of a myr-GFP construct under control of the NP225 promoter. (C) A high magnification image of an optical section through a glomerulus more clearly shows the separation of PN dendrites and presynaptic terminals. The NP225 promoter is expressed in 60% of all PNs. myrGFP coding sequences are fused to 3′UTR sequences of the dendritic mRNA CaMKII. (D) Schematic illustrating the observed expression of Me31B foci in a PN (green). Me31B foci (magenta) are seen within dendrites in the glomerulus (left), but not within nerve terminals (right), which are instead surrounded by Me31B foci. (E) Expression of Me31B in dendritic regions of projection neurons. Me31B foci (magenta; arrowheads) are present within PN dendrites (green) marked by myrGFP expression in NP225-positive PNs (green). Additional Me31B spots outside PN dendrites are likely to be in glial processes and/or local interneurons. (F) Axonal projections of PNs (green) in the mushroom-body calyx visualized by GFP expression. Me31B foci (magenta) in the calyx do not overlap with axonal endings of PNs. PN dendrites in the AL have 0.22 Me31B foci/μm2 (see Figure 6); in contrast, only 0.026 Me31B particles/μm2 are seen in PN terminals in the calyx (this small number is potentially due to the limitations of the technique used). (G) Schematic illustrating predominantly dendritic localization of Me31B foci in OSNs (olfactory sensory neurons). (H) Me31B foci (magenta) are present in cell bodies of OSNs (green) and in OSN dendrites within the sensillum (arrowheads). Or83b-Gal4 driven GFP is used to reveal OSN soma and dendrites (green); and Me31B (magenta) is visualized by immunocytochemistry. (I) Presynaptic areas of OSNs (green) in the antennal lobes are marked by Or83b-Gal4 driven expression of synaptotagmin-GFP (Syt-GFP). Me31B foci (magenta) are often adjacent to, but rarely, if ever, within OSN presynaptic endings. This is consistent with predominantly somatodendritic localization of Me31B particles.
In PNs, synaptic Me31B foci are predominantly found in postsynaptic dendrites (Figures 5D–F). Thus, in AL of animals expressing the myr-GFPCaMKII3′UTR reporter in PNs, Me31B foci showed considerable overlap with reporter GFP (Figure 5E). In contrast, Me31B foci were almost completely excluded from presynaptic endings of PNs visualized in the MB calyx (Figure 5F; 0.22 Me31B foci/μm2 in PN dendrites in the AL versus 0.026 foci/μm2 in PN terminals in the calyx). Me31B foci observed within AL glomeruli, but outside the areas labeled by the myr-GFPCaMKII3′UTR reporter in Figure 5E, are probably contained in processes that other cell types (e.g., LNs or glia) send to the AL.
In additional control experiments, we used the MARCM technique to analyze Me31B localization in single PNs as well as the specificity of Me31B staining in PN dendrites (Lee and Luo, 2001). The MARCM technique allows one to visualize individual, homozygous-mutant cells in otherwise heterozygous animals. We used the Gal4 lines NP3529, which among other tissues marks PNs specifically projecting into the glomerulus DL-1, and GH146, which marks about 60% of all PNs. Dendrites of individually imaged DL-1 PNs contained Me31B foci (Figure 6A). Loss of these foci in MARCM-generated me31BD2/me31BD2 mutant PNs confirmed the specificity of Me31B staining (Figure 6B).
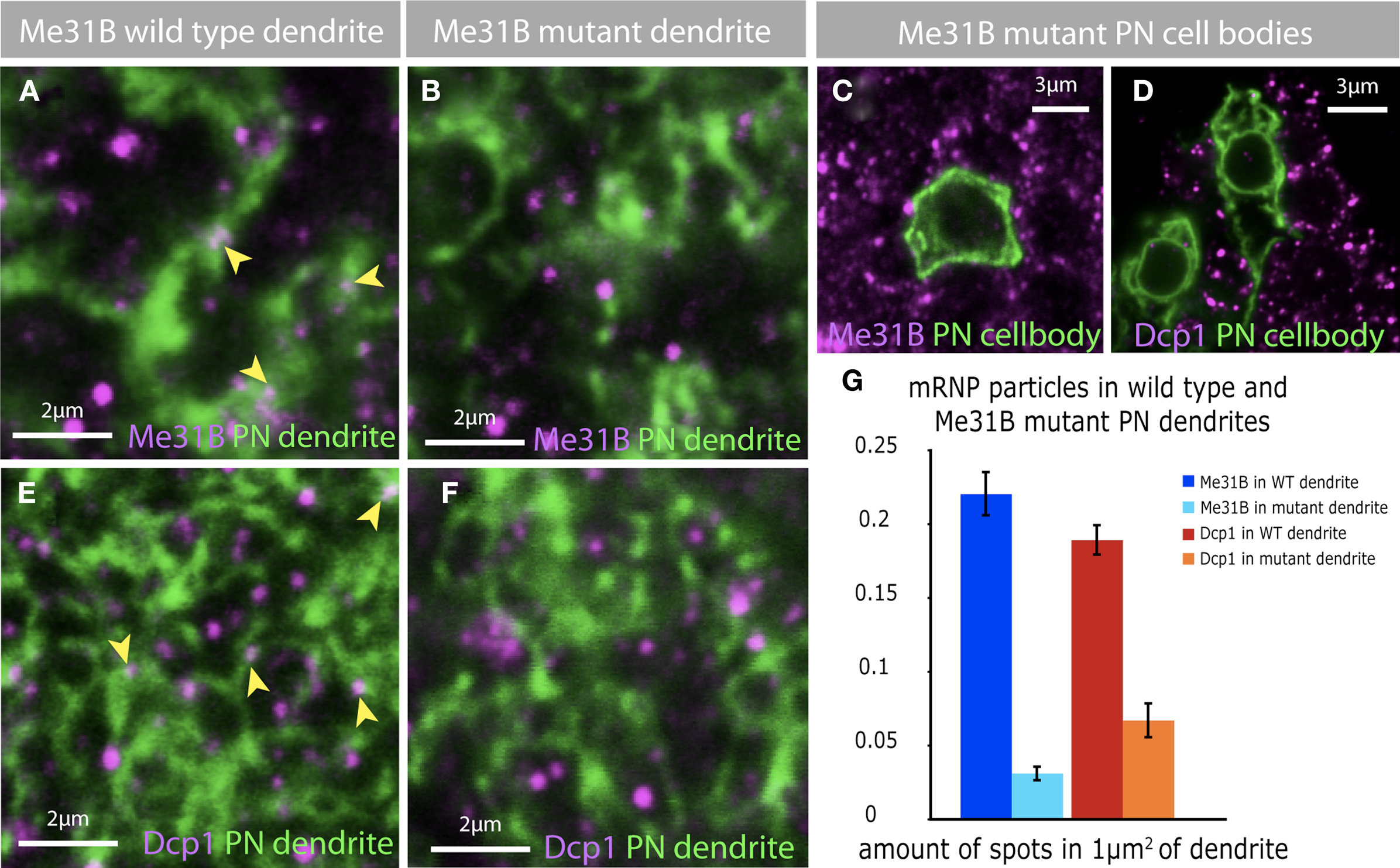
Figure 6. Depletion of Me31B in projection neurons effect mRNP particles. (A) Me31B foci (magenta, arrowheads) are localized to dendrites of PNs. The dendrites of a single PN (green) was marked using the MARCM technique. (B,C) The depletion of Me31B in single PNs, by inducing Me31B mutant MARCM clones, leads to a loss of Me31B foci in mutant dendrites (B) and cell bodies (C). (D–F) In addition the loss of Me31B in PNs effects the distribution of Dcp1 foci in PN cell bodies (D) and dendrites (F). Dcp1 foci are present in wild-type PN dendrites of single-cell MARCM clones (E, magenta, arrowhead), but fewer Dcp1 foci are observed in Me31B mutant PN dendrites (F). (G) Quantification of Me31B and Dcp1 foci expression in wild-type and Me31B mutant PN dendrites shows a significant decrease in the number of foci in mutant PN dendrites. The graph shows the average amount of foci in an area of 1 μm2 dendrite for wt and mutant PN dendrites, coimmuno-labeled with Me31B or Dcp1. Eight to ten single slices of a confocal stack of five independent PN MARCM clones for each genotype were analyzed for the expression of Me31B and Dcp1 foci in PN dendrites. Genotype of imaged flies: wt MARCM: hsFLP, mCD8-GFP; NP225; FRT82B/tubGal80,FRT82B Me31B MARCM: GH146/hsFLP, mCD8-GFP; Me31BΔ2, FRT40A/tubGal80, FRT40A.
Me31B foci were similarly enriched in dendrites of olfactory sensory neurons (OSNs; Figures 5G–I). OSN soma and sensory dendrites in the antenna visualized in Or83bGal4-UAS-GFP (Or83b > GFP) animals contained several Me31B particles. However, OSN presynaptic terminals imaged in the AL of Or83b > SytGFP animals, showed very little overlap with Me31B foci. Indeed, we observed many instances where presynaptic endings revealed by Syt-GFP appeared to surround Me31B labeled particles.
Thus, in at least two of the major classes of AL neurons, Me31B foci are predominantly localized to dendrites.
Me31B Regulates Dcp1 Positive Particles in PN Dendrites
In non-neuronal cells, Me31B and its non-fly orthologs are required for the normal assembly of P-body like particles which contain not only Me31B, but also RNA hydrolases such as Dcp1 and Pcm (Barbee et al., 2006; Hillebrand et al., 2007). The observation that most dendritic Me31B-positive particles are not marked by anti-Dcp1 pointed to the possibility that dendritic Dcp1 particles may assemble in an Me31B independent fashion. To address this specific question, as well as the more general issue of whether different classes of synaptic mRNPs interact and/or share common mechanisms for biogenesis, we examined the effect of loss of Me31B protein, on the brightness and number of Me31B or Dcp1 positive particles in olfactory PNs in vivo (Figure 6).
We examined Me31B and Dcp1 in single-cell clones of wild-type or me31bD2/me31bD2 PNs generated using the MARCM technique. CD8-GFP marked, wild-type MARCM clones showed both somatic and dendritic Me31B and Dcp1 particles within areas marked by CD8-GFP (Figures 6A,E and not shown). And as expected, me31bD2/me31bD2 mutant cells lacked Me31b-positive foci in soma and dendrites (Figures 6B,C). In addition, these cells showed a striking reduction of Dcp1 foci (Figures 6D,F), which is quantified in Figure 6G. Thus, although synaptic Dcp1 foci appear to be largely free of Me31B (or to have particularly low levels of the protein), the integrity of these foci is dependent on Me31B.
Me31B Represses a CaMKII Reporter mRNA in PN Dendrites
The presence of Me31B in postsynaptic particles suggested that the protein might repress synaptic mRNA translation, similar to its role in oskar and bicD regulation. We therefore asked whether loss of Me31B would result in significant upregulation of CaMKII translation in dendrites, much as its loss causes precocious translation of Oskar and BicD in oocytes (Nakamura et al., 2001; Ashraf et al., 2006).
We knocked down Me31B in PNs by coexpressing two double-stranded RNAi constructs that have been previously used to knockdown Me31B in non-neuronal cells (Barbee et al., 2006; Hillebrand et al., 2007) and analyzed the effects of this knockdown on expression of a translational reporter for the CaMKII mRNA (Ashraf et al., 2006) (Figures 7D,E). In the CaMKII reporter, GFP coding sequences are placed under control of the 3′UTR for CaMKII. When transcribed in PNs, the reporter mRNA is transported to PN dendrites where it is repressed by the miRNA pathway and activated by behavioral experience (Ashraf et al., 2006). Knockdown of Me31B in PNs caused a substantial increase in levels of CaMKII reporter protein in the AL (Figures 7D,E). Thus, Me31B is required in PNs for repression of a synaptically targeted, miRNA regulated mRNA. Me31B repression influences both dendritic and somatic levels of the CaMKII reporter protein; indeed, knockdown of Me31B is associated with roughly two-fold increase in levels of the reporter in PN cell bodies (not shown). This might be expected if Me31B is involved in the repression of CaMKII transcript in the soma in preparation for its dendritic targeting.
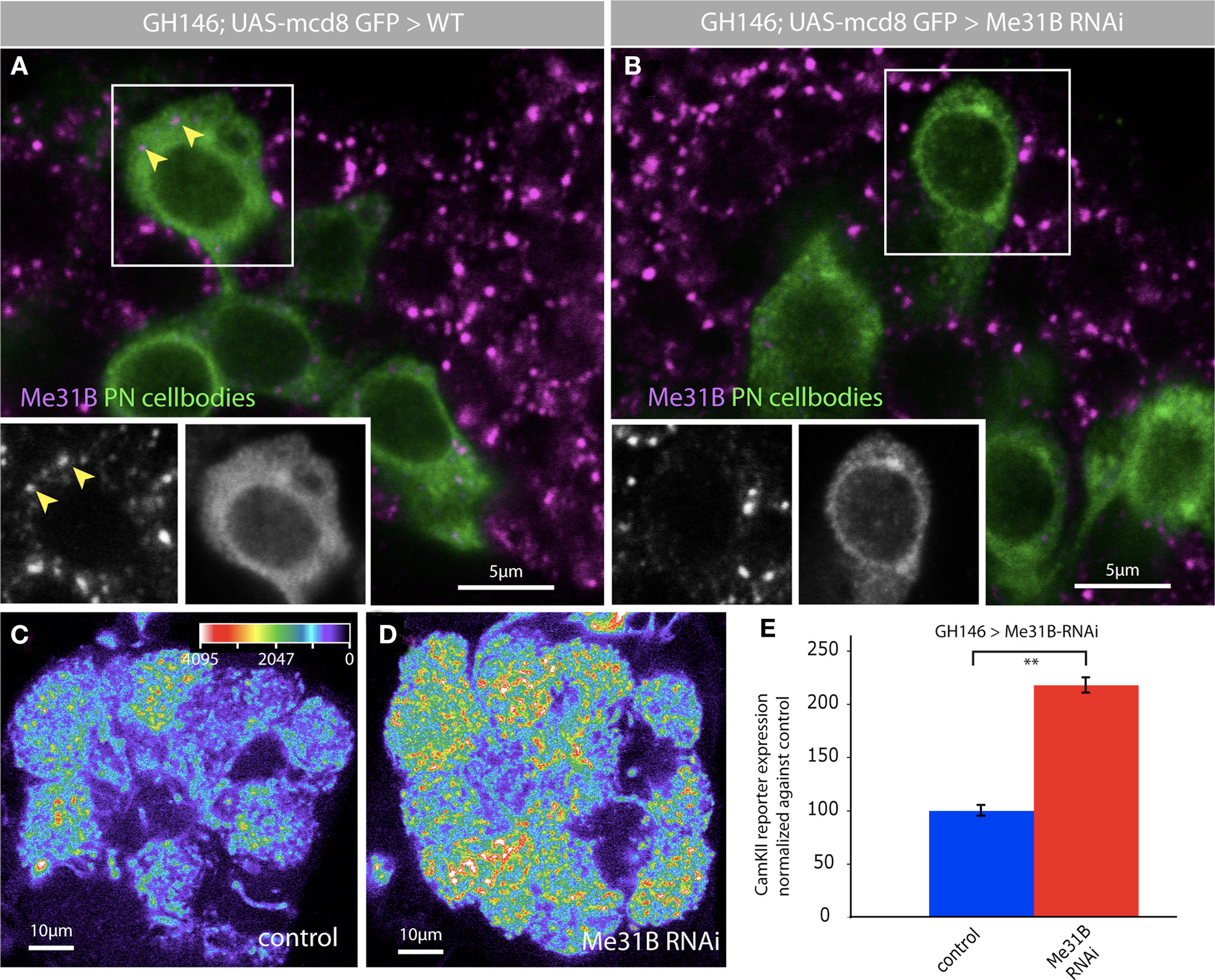
Figure 7. Me31B regulates CaMKII levels in projection neuron dendrites. The depletion of Me31B by RNAi leads to a reduction of Me31B foci in PN cell bodies. Two independent RNAi lines for Me31B were coexpressed with GH146-Gal4 in PNs, which were marked by the expression of UAS-myrGFPCamKII-3′UTR. The use of tub-Gal80ts allowed a specific temporal expression of the Me31B hairpin constructs in adult flies (crosses were kept on 18°C till late pupal stage and Gal4 expression was induced by switching the crosses to 30°C for 4 days). (A) Me31B foci in wt PN cell bodies (magenta, arrowheads); insets show both channels for the boxed cell; Genotype for wt: +/+; GH146-Gal4, UAS-myrGFPCaMKII-3′UTR/+; 2x tub-Gal80ts/+. (B) Reduction of Me31B foci after the expression of Me31B RNAi hairpin constructs in PNs; insets show both channels for the boxed cell; Genotype for RNAi: UAS-me31B-RNAi/+; GH146Gal4, UAS-myrGFPCaMKII-3′UTR/UAS-me31B-RNAi; 2x tub-Gal80ts/+. (C–E) Expression of two Me31B RNAi lines in PNs using GH146-Gal4 increases the level of CaMKII translational reporter protein expression (myrGFPCaMKII-3′UTR). (C) Representative image of myrGFPCaMKII-3′UTR expression driven in GH146 Gal4, presented as a false color image (the gray value intensity for the associated color is annotated); tub Gal80ts flies which restricts reporter expression to adults; Genotype: +/+; GH146-Gal4, UAS-myrGFPCaMKII-3′UTR/+; 2x tub-Gal80ts/+. (D) Representative image of myrGFPCaMKII-3′UTR expression in the antennal lobe of flies expressing two RNAi constructs for Me31B (intensity scale bar in C). Genotype: UAS-me31B-RNAi/+; GH146Gal4, UAS-myrGFPCaMKII-3′UTR/UAS-me31B-RNAi; 2x tub-Gal80ts/+. (E) myrGFPCaMKII-3′UTR fluorescence quantified in the DA1 glomerulus of antennal lobes of wt and experimental flies in which Me31B is depleted in ∼60% of all PNs. A two-fold increase of CaMKII reporter expression in the DA1 glomerulus is observed following Me31B depletion in adult flies: P < 0.001 (Student’s t-test). Measurements correspond to the average mean intensity of CamKII reporter expression through a confocal stack of the DA-1 glomerulus (8–10 slices) of 10 adult brains analyzed.
Discussion
Experimental observations presented here are of interest because they: (a) provide new information on neuronal mRNPs in vivo; (b) reveal a novel function for Me31B in the regulation of dendritic mRNA expression; and (c) together highlight important aspects of mRNA regulation at synapses.
Properties and Functional Implications of Synaptic ME31B Granules Observed In Vivo
The Me31B/Dhh1p/DDX6/CGH-1 class of DEAD box helicases is associated with many different kinds of mRNP aggregates, including maternal RNA storage granules, P-bodies, stress granules, as well as various granule subtypes observed during C. elegans germline development (Nakamura et al., 2001; Cougot et al., 2004; Andrei et al., 2005; Coller and Parker, 2005; Kedersha et al., 2005; Eulalio et al., 2007b; Boag et al., 2008; Jud et al., 2008; Noble et al., 2008). In addition it is required for the assembly of P-bodies in yeast, Drosophila and mammalian cells (Andrei et al., 2005; Coller and Parker, 2005; Eulalio et al., 2007b; Hillebrand et al., 2007) and as well for the formation of stress granules in mammals (Wilczynska et al., 2005). For these reasons, the punctate distribution of Me31B in postsynaptic dendrites is likely to indicate its presence in a specific type of synaptic mRNP particle. However, unlike Me31B-positive particles described in neurites of cultured Drosophila neurons, synaptic Me31B foci do not appear to contain the RNA hydrolases Dcp1 and Pcm/Xrn-1. Thus, they may be a distinct class of particle, which localize preferentially to postsynaptic dendrites. These represent early images of candidate mRNA storage particles at synapses in vivo (Christie et al., 2009). A paucity of antibodies and the challenging nature of such high-resolution immunocytochemistry in whole brain tissue has so far made it difficult to more completely characterize other components of synaptic Me31B particles as well as to establish whether Dcp1, Pcm, and Stau coexist on the same or different particle in the adult brain. Indeed even our conclusion that Me31B particles constitute a separate class must be qualified by the possibility that the visualization of two apparently distinct particle types arises from an artifact of incomplete antibody penetration into the neuropil.
It is possible that synaptic Me31B particles could be analogous to recently described granules in the C. elegans germline, which contain translationally controlled mRNAs and CGH-1/Me31B but exclude decapping enzymes and the P-body protein PATR1/PAT1 (Boag et al., 2008; Gallo et al., 2008). Immunoprecipitation and further colocalization studies suggest that these granules can also contain PAB-1, ATX-2, or TIA-1, markers of stress granules, which in other systems, contain translation initiation factors together with mRNAs stalled in translational initiation (Kedersha et al., 2005; Gallo et al., 2008). Thus, it is conceivable that Me31B/CGH-1-containing storage particles contain mRNAs stored in a stress-granule like state, in which the resident mRNAs are available for rapid activation (Rajyaguru and Parker, 2008).
The potential separation of storage and degradative particles leads to an attractive model in which individual mRNAs may transition from being available for translational activation in a storage granule, to being targeted for degradation in a P-body like particle. This is supported as well by observations in dendrites of cultured mammalian neurons where a distinct class of RNPs contain the degradative enzyme Xrn1, which is excluded from RNPs supposedly involved in storage (Cougot et al., 2008).
At synapses, a transition between storage and degradation particles may occur by three, non-exclusive, candidate mechanisms: (a) by the remodeling of a storage mRNP to a degradative one through protein exchange; (b) by the initial exit of mRNA from the storage RNP to a translating pool, followed by its subsequent targeting to a degradative particle; or (c) the fusion of the two particles. Recent studies in Drosophila provide a possible mechanism by which a change of proteins in RNP complexes could alter its function. Two related proteins of the Lsm-family, Enhancer of Decapping 3 (EDC3), which is implicated to play a role in mRNA decay, and Trailer Hitch (Tral), which supposedly is involved in mRNA repression, interact at the same domain with the Me31B protein. This suggests that the function of Me31B complexes might be determined by the interaction with specific binding partners (Tritschler et al., 2008, 2009). Some support for the second model is provided by the observation that the synaptically localized Arc mRNA is targeted for degradation after its translation is induced by synaptic activity (Bramham et al., 2008) and also by the observation that RCK-positive particles in dendrites of cultured hippocampal neurons are transiently disassembled following BDNF stimulation (Zeitelhofer et al., 2008). Further studies are required to understand how, when, and even whether these transitions of mRNA state occur in synapses and other biological contexts.
Functions of Me31B in Neuronal Translational Repression
Together with many analogous studies in yeast and mammalian cells, previous observations in Drosophila that Me31B is a repressor of maternal mRNA translation, a component of a repression pathway mediated by the bantam microRNA, and a repressor of growth of terminal dendrites, has led to a strong model that Me31B is a translational repressor protein (Nakamura et al., 2001; Coller et al., 2002; Barbee et al., 2006; Chu and Rana, 2006). In contrast, recent studies in C. elegans and P. falciparum have shown that Me31B orthologs, CGH-1 and DOZI, associate with specific mRNAs and protects them from degradation (Mair et al., 2006; Boag et al., 2008).
Our observations in neurons indicate a function for Me31B in repressing translation of a miRNA regulated, dendritically localized reporter mRNA in vivo. This is consistent with two related lines of data. First, it is consistent with the known function for Me31B in repression of miRNA-target genes in Drosophila wing imaginal cells as well as for its human homolog RCK in mammalian cultured cells (Barbee et al., 2006; Chu and Rana, 2006). Second, the correlation we observe between loss of synaptic Dcp1 puncta and upregulation of the CaMKII reporter, is consistent with observations in hippocampal cultured cells, where observed disassembly of “dendritic P-bodies” induced by synaptic stimulation has been proposed to underlie the temporally coincident translation of localized mRNAs (Zeitelhofer et al., 2008).
Thus, we suggest a simple model in which neuronal Me31B, as well as its homologs in other metazoa, mediates the formation of synaptic mRNP particles that contain locally repressed mRNAs. And that synaptic stimulation-induced disassembly of these particles is one aspect of the mechanism of local translational control.
Future Directions
One key goal of future studies will be to understand the composition and dynamics of dendritic mRNPs in vivo. This will be aided by genetic techniques to replace endogenous translational control molecules with genetically encoded, fluorescently tagged variants that retain functional and localization patterns of the endogenous proteins. When coupled with procedures to induce local protein synthesis in dendrites, such reagents will allow analysis of functionally relevant particle dynamics in vivo. In addition, by eliminating the need for antibodies whose use may be associated with artifacts of inclusion and exclusion, such reagents may provide more direct insight into the real nature of synaptic mRNPs in vivo.
A second goal is to understand the mechanism by which Me31B regulates the expression of CaMKII reporter levels in vivo. Although Me31B has been shown to be required for the miRNA pathway (Barbee et al., 2006; Chu and Rana, 2006) it is also required for other forms of translational repression, for example in S. cerevisiae that does not have miRNAs. Similarly, although the reporter used here is miRNA regulated, the same UTR also has binding sites for translational regulators that may operate independently of miRNAs (Ashraf et al., 2006). Thus, important and linked goals of future studies are to understand mechanisms by which the CaMKII UTR is regulated in dendrites and how Me31B engages with these mechanisms of neuronal translational control.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Prof Veronica Rodrigues and her lab at the NCBS for generous and extensive advice, Akira Nakamura, Eimear Holohan, Cathal McCann, and members of the Ramaswami lab for discussions, advice, and comments on the manuscript. We acknowledge the Kyoto, Szeged, VDRC, and Bloomington Stock Centers, Akira Nakamura, Veronica Rodrigues, S. G. Gajendra, Liqun Luo, and Rachel Wilson for Drosophila stocks; Shunji Ohsako, Leslie Griffith, Sarah Newbury, Jim Wilhelm, and Lynn Manseau for fly stocks and antibodies. This work, initiated with funds from the National Institute of Drug Abuse of the NIH, was funded by grants from the Science Foundation of Ireland to Mani Ramaswami and Anil Kokaram.
References
Aakalu, G., Smith, W. B., Nguyen, N., Jiang, C., and Schuman, E. M. (2001). Dynamic visualization of local protein synthesis in hippocampal neurons. Neuron 30, 489–502.
Andrei, M. A., Ingelfinger, D., Heintzmann, R., Achsel, T., Rivera-Pomar, R.,and Lührmann, R. (2005). A role for eIF4E and eIF4E-transporter in targeting mRNPs to mammalian processing bodies. RNA 11, 717–727.
Ashraf, S. I., McLoon, A. L., Sclarsic, S. M., and Kunes, S. (2006). Synaptic protein synthesis associated with memory is regulated by the RISC pathway in Drosophila. Cell 124, 191–205.
Banerjee, S., Neveu, P., and Kosik, K. S. (2009). A coordinated local translational control point at the synapse involving relief from silencing and MOV10 degradation. Neuron 64, 871–884.
Barbee, S. A., Estes, P. S., Cziko, A. M., Hillebrand, J., Luedeman, R. A., Coller, J. M., Johnson, N., Howlett, I. C., Geng, C., Ueda, R., Brand, A. H., Newbury, S. F., Wilhelm, J. E., Levine, R. B., Nakamura, A., Parker, R., and Ramaswami, M. (2006). Staufen- and FMRP-containing neuronal RNPs are structurally and functionally related to somatic P bodies. Neuron 52, 997–1009.
Boag, P. R., Atalay, A., Robida, S., Reinke, V., and Blackwell, T. K. (2008). Protection of specific maternal messenger RNAs by the P body protein CGH-1 (Dhh1/RCK) during Caenorhabditis elegans oogenesis. J. Cell Biol. 182, 543–557.
Bolduc, F. V., Bell, K., Cox, H., Broadie, K. S., and Tully, T. (2008). Excess protein synthesis in Drosophila fragile X mutants impairs long-term memory. Nat. Neurosci. 11, 1143–1145.
Bramham, C. R., and Wells, D. G. (2007). Dendritic mRNA: transport, translation and function. Nat. Rev. Neurosci. 8, 776–789.
Bramham, C. R., Worley, P. F., Moore, M. J., and Guzowski, J. F. (2008). The immediate early gene arc/arg3.1: regulation, mechanisms, and function. J. Neurosci. 28, 11760–11767.
Christie, S. B., Akins, M. R., Schwob, J. E., and Fallon, J. R. (2009). The FXG: a presynaptic fragile X granule expressed in a subset of developing brain circuits. J. Neurosci. 29, 1514–1524.
Chu, C. Y., and Rana, T. M. (2006). Translation repression in human cells by microRNA-induced gene silencing requires RCK/p54. PLoS Biol. 4, e210. doi: 10.1371/journal.pbio.0040210.
Coller, J. M., and Parker, R. (2005). General translational repression by activators of mRNA decapping. Cell 122, 875–886.
Coller, J. M., Tucker, M., Sheth, U., Valencia-Sanchez, M. A., and Parker, R. (2002). The DEAD box helicase, Dhh1p, functions in mRNA decapping and interacts with both the decapping and deadenylase complexes. RNA 7, 1717–1727.
Costa-Mattioli, M., Sossin, W. S., Klann, E., and Sonenberg, N. (2009). Translational control of long-lasting synaptic plasticity and memory. Neuron 61, 10–26.
Cougot, N., Babajko, S., and Seraphin, B. (2004). Cytoplasmic foci are sites of mRNA decay in human cells. J. Cell Biol. 165, 31–40.
Cougot, N., Bhattacharyya, S. N., Tapia-Arancibia, L., Bordonne, R., Filipowicz, W., Bertrand, E., and Rage, F. (2008). Dendrites of mammalian neurons contain specialized P-body-like structures that respond to neuronal activation. J. Neurosci. 28, 13793–13804.
Davis, R. L. (2005). Olfactory memory formation in Drosophila: from molecular to systems neuroscience. Annu. Rev. Neurosci. 28, 275–302.
Dietzl, G., Chen, D., Schnorrer, F., Su, K. C., Barinova, Y., Fellner, M., Gasser, B., Kinsey, K., Oppel, S., Scheiblauer, S., Couto, A., Marra, V., Keleman, K., and Dickson, B. J. (2007). A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156.
Dubnau, J., Chiang, A. S., Grady, L., Barditch, J., Gossweiler, S., McNeil, J., Smith, P., Buldoc, F., Scott, R., Certa, U., Broger, C., and Tully, T.(2003). The staufen/pumilio pathway is involved in Drosophila long-term memory. Curr. Biol. 13, 286–296.
Eulalio, A., Behm-Ansmant, I., and Izaurralde E. (2007a). P bodies: at the crossroads of post-transcriptional pathways. Nat. Rev. Mol. Cell Biol. 8, 9–22.
Eulalio, A., Behm-Ansmant, I., Schweizer, D., and Izaurralde E. (2007b). P-body formation is a consequence, not the cause of RNA-mediated gene silencing. Mol. Cell. Biol. 27, 3970–3981.
Fiala, A. (2008). Olfaction and olfactory learning in Drosophila: recent progress. Curr. Opin. Neurobiol. 17, 720–726.
Gallo, C. M., Munro, E., Rasoloson, D., Merritt, C., and Seydoux, G. (2008). Processing bodies and germ granules are distinct RNA granules that interact in C. elegans embryos. Dev. Biol. 323, 76–87.
Hillebrand, J., Barbee, S. A., and Ramaswami, M. (2007). P-body components, microRNA regulation, and synaptic plasticity. ScientificWorldJournal 7, 178–190.
Jefferis, G. S., Marin, E. C., Stocker, R. F., and Luo, L. (2001). Target neuron prespecification in the olfactory map of Drosophila. Nature 414, 204–208.
Jud, M. C., Czerwinski, M. J., Wood, M. P., Young, R. A., Gallo, C. M., Bickel, J. S., Petty, E. L., Mason, J. M., Little, B.A., Padilla, P.A., and Schisa, J. A. (2008). Large P body-like RNPs form in C. elegans oocytes in response to arrested ovulation, heat shock, osmotic stress, and anoxia and are regulated by the major sperm protein pathway. Dev. Biol. 318, 38–51.
Kedersha, N., Stoecklin, G., Ayodele, M., Yacono, P., Lykke-Andersen, J., Fritzler, M. J., Scheuner, D., Kaufman, R. J., Golan, D. E., and Anderson, P. (2005). Stress granules and processing bodies are dynamically linked sites of mRNP remodeling. J. Cell Biol. 169, 871–884.
Keene, A. C., and Waddell, S. (2007). Drosophila olfactory memory: single genes to complex neural circuits. Nat. Rev. Neurosci. 8, 341–354.
Kiebler, M. A., and Bassell, G. J. (2006). Neuronal RNA granules: movers and makers. Neuron 51, 685–690.
Koh, Y. H., Popova, E., Thomas, U., Griffith, L. C., and Budnik, V. (1999). Regulation of DLG localization at synapses by CaMKII-dependent phosphorylation. Cell 98, 353–363.
Krichevsky, A. M., and Kosik, K. S. (2001). Neuronal RNA granules: a link between RNA localization and stimulation-dependent translation. Neuron 32, 683–696.
Kwak, J. E., Drier, E., Barbee, S. A., Ramaswami, M., Yin, J. C., and Wickens, M. (2008). GLD2 poly(A) polymerase is required for long-term memory. Proc. Natl. Acad. Sci. U.S.A. 105, 14644–14649.
Lee, A., Li, W., Xu, K., Bogert, B. A., Su, K., and Gao, F. B. (2003). Control of dendritic development by the Drosophila fragile X-related gene involves the small GTPase Rac1. Development 130, 5543–5552.
Lee, T., and Luo, L. (2001). Mosaic analysis with a repressible cell marker (MARCM) for Drosophila neural development. Trends Neurosci. 24, 251–254.
Lin, M. D., Jiao, X., Grima, D., Newbury, S. F., Kiledjian, M., and Chou, T. B. (2008). Drosophila processing bodies in oogenesis. Dev. Biol. 322, 276–288.
Mair, G. R., Braks, J. A., Garver, L. S., Wiegant, J. C., Hall, N., Dirks, R. W., Khan, S. M., Dimopoulos, G., Janse, C. J., and Waters, A. P. (2006). Regulation of sexual development of Plasmodium by translational repression. Science 313, 667–669.
Martin, K. C., and Zukin, R. S. (2006). RNA trafficking and local protein synthesis in dendrites: an overview. J. Neurosci. 26, 7131–7134.
Mayford, M., Baranes, D., Podsypanina, K., and Kandel, E. R. (1996). The 3′-untranslated region of CaMKII alpha is a cis-acting signal for the localization and translation of mRNA in dendrites. Proc. Natl. Acad. Sci. U.S.A. 93, 13250–13255.
Mikl, M., Vendra, G., Doyle, M., and Kiebler, M. A. (2010). RNA localization in neurite morphogenesis and synaptic regulation: current evidence and novel approaches. J. Comp. Physiol. A Neuroethol. Sens. Neural. Behav. Physiol. 196, 321–334.
Miller, L. C., Blandford, V., McAdam, R., Sanchez-Carbente, M. R., Badeaux, F., DesGroseillers, L., and Sossin, W. S. (2009). Combinations of DEAD box proteins distinguish distinct types of RNA: protein complexes in neurons. Mol. Cell. Neurosci. 40, 485–495.
Minshall, N., and Standart, N. (2004). The active form of Xp54 RNA helicase in translational repression is an RNA-mediated oligomer. Nucleic Acids Res. 32, 1325–1334.
Nakamura, A., Amikura, R., Hanyu, K., and Kobayashi, S. (2001). Me31B silences translation of oocyte-localizing RNAs through the formation of cytoplasmic RNP complex during Drosophila oogenesis. Development 128, 3233–3242.
Navarro, R. E., Shim, E. Y., Kohara, Y., Singson, A., and Blackwell, T. K. (2001). cgh-1, a conserved predicted RNA helicase required for gametogenesis and protection from physiological germline apoptosis in C. elegans. Development 128, 3221–3232.
Noble, S. L., Allen, B. L., Goh, L. K., Nordick, K., and Evans, T. C. (2008). Maternal mRNAs are regulated by diverse P body-related mRNP granules during early Caenorhabditis elegans development. J. Cell Biol. 182, 559–572.
Pan, K., Kokaram, A., Hillebrand, J., and Ramaswami, M. (2010). “Gaussian mixture model for spot analysis in microscopy,” in Proceedings of the IEEE International Symposium on Biomedical Imaging. Rotterdam, Netherlands.
Parker, R., and Sheth, U. (2007). P bodies and the control of mRNA translation and degradation. Mol. Cell 25, 635–646.
Rajyaguru, P., and Parker, R. (2008). CGH-1 and the control of maternal mRNAs. Trends Cell Biol. 19, 24–28.
Richter, J. D., and Klann, E. (2009). Making synaptic plasticity and memory last: mechanisms of translational regulation. Genes Dev. 23, 1–11.
Sossin, W. S., and DesGroseillers, L. (2006). Intracellular trafficking of RNA in neurons. Traffic 7, 1581–1589.
Steward, O., and Schuman, E. M. (2001). Protein synthesis at synaptic sites on dendrites. Annu. Rev. Neurosci. 24, 299–325.
St Johnston, D., Beuchle, D., and Nusslein-Volhard, C. (1991). Staufen, a gene required to localize maternal RNAs in the Drosophila egg. Cell 66, 51–63.
Stocker, R. F., Lienhard, M. C., Borst, A., and Fischbach, K. F. (1990). Neuronal architecture of the antennal lobe in Drosophila melanogaster. Cell Tissue Res. 262, 9–34.
Sutton, M. A., and Schuman, E. M. (2006). Dendritic protein synthesis, synaptic plasticity, and memory. Cell 127, 49–58.
Takamatsu, Y., Kishimoto, Y., and Ohsako, S. (2003). Immunohistochemical study of Ca2+/calmodulin-dependent protein kinase II in the Drosophila brain using a specific monoclonal antibody. Brain Res. 974, 99–116.
Tritschler, F., Braun, J. E., Eulalio, A., Truffault, V., Izaurralde, E., and Weichenrieder, O. (2009). Structural basis for the mutually exclusive anchoring of P body components EDC3 and Tral to the DEAD box protein DDX6/Me31B. Mol. Cell 33, 661–668.
Tritschler, F., Eulalio, A., Helms, S., Schmidt, S., Coles, M., Weichenrieder, O., Izaurralde, E., and Truffault, V. (2008). A similar mode of interaction enables Trailer Hitch and EDC3 to associate with DCP1 and Me31B in distinct protein complexes. Mol. Cell. Biol. 28, 6695–6708.
Tubing, F., Vendra, G., Mikl, M., Macchi, P., Thomas, S.,and Kiebler, M. A. (2010). Dendritically localized transcripts are sorted into distinct ribonucleoprotein particles that display fast directional motility along dendrites of hippocampal neurons. J. Neurosci. 30, 4160–4170.
Ueda, N., Nakano, R., Ghahramani, Z., and Hinton, G. E. (2000). SMEM algorithm for mixture models. Neural Comput. 12, 2109–2128.
Vosshall, L. B., and Stocker, R. F. (2007). Molecular architecture of smell and taste in Drosophila. Annu. Rev. Neurosci. 30, 505–533.
Weston, A., and Sommerville, J. (2006). Xp54 and related (DDX6-like) RNA helicases: roles in messenger RNP assembly, translation regulation and RNA degradation. Nucleic Acids Res. 34, 3082–3094.
Wilczynska, A., Aigueperse, C., Kress, M., Dautry, F., and Weil, D. (2005). The translational regulator CPEB1 provides a link between dcp1 bodies and stress granules. J. Cell Sci. 118, 981–992.
Ye, B., Petritsch, C., Clark, I. E., Gavis, E. R., Jan, L. Y., and Jan, Y. N. (2004). Nanos and Pumilio are essential for dendrite morphogenesis in Drosophila peripheral neurons. Curr. Biol. 14, 314–321.
Zabolotskaya, M. V., Grima, D. P., Lin, M. D., Chou, T. B., and Newbury, S. F. (2008). The 5′–3′ exoribonuclease Pacman is required for normal male fertility and is dynamically localized in cytoplasmic particles in Drosophila testis cells. Biochem. J. 416, 327–335.
Keywords: Me13B, Drosophila, synaptic mRNPs, CamKII, translational control, olfactory circuitry
Citation: Hillebrand J, Pan K, Kokaram A, Barbee S, Parker R and Ramaswami M (2010) The Me31B DEAD-box helicase localizes to postsynaptic foci and regulates expression of a CaMKII reporter mRNA in dendrites of Drosophila olfactory projection neurons. Front. Neural Circuits 4:121. doi: 10.3389/fncir.2010.00121
Received: 30 April 2010;
Accepted: 27 September 2010;
Published online: 03 November 2010.
Edited by:
Leslie B. Vosshall, Rockefeller University, USAReviewed by:
Michael A. Sutton, University of Michigan, USAKelsey Martin, University of California at Los Angeles, USA
Jason Dictenberg, Hunter College, USA
Copyright: © 2010 Hillebrand, Pan, Kokaram, Barbee, Parker and Ramaswami. This is an open-access article subject to an exclusive license agreement between the authors and the Frontiers Research Foundation, which permits unrestricted use, distribution, and reproduction in any medium, provided the original authors and source are credited.
*Correspondence: Mani Ramaswami and Jens Hillebrand, Institute for Neuroscience, Trinity College Dublin, Lloyd Building, Dublin 2, Ireland. e-mail: mani@u.arizona.edu; jens.hillebrand@tcd.ie