Data sharing in neuroimaging research
- 1Neurospin, Commissariat à l'Energie Atomique et aux Energies Alternatives, Gif-sur-Yvette, France
- 2Brain Imaging Centre, University of California at Berkeley, Berkeley, CA, USA
- 3International Neuroinformatics Coordinating Facility, Karolinska Institute, Stockholm, Sweden
- 4McGovern Institute for Brain Research, Massachusetts Institute of Technology, Cambridge, MA, USA
- 5School of Informatics, University of Edinburgh, Edinburgh, UK
- 6Department of Psychological and Brain Sciences, Dartmouth College, Hanover, NH, USA
- 7Department of Experimental Psychology, Otto-von-Guericke-University Magdeburg, Magdeburg, Germany
- 8Division of Informatics, Department of Psychiatry, University of Massachusetts Medical School, Worcester, MA, USA
- 9Massachusetts General Hospital and Department of Radiology, Athinoula A Martinos Center for Biomedical Imaging, Harvard Medical School, Boston, MA, USA
- 10Department of Psychiatry and Human Behavior, and Department of Computer Science, University of California at Irvine, CA, USA
- 11Department of Radiology, Washington University School of Medicine, St. Louis, MO, USA
- 12Imaging Research Center and Departments of Psychology and Neurobiology, University of Texas at Austin, Austin, TX, USA
- 13Wellcome Trust Centre for Neuroimaging, London, UK
Significant resources around the world have been invested in neuroimaging studies of brain function and disease. Easier access to this large body of work should have profound impact on research in cognitive neuroscience and psychiatry, leading to advances in the diagnosis and treatment of psychiatric and neurological disease. A trend toward increased sharing of neuroimaging data has emerged in recent years. Nevertheless, a number of barriers continue to impede momentum. Many researchers and institutions remain uncertain about how to share data or lack the tools and expertise to participate in data sharing. The use of electronic data capture (EDC) methods for neuroimaging greatly simplifies the task of data collection and has the potential to help standardize many aspects of data sharing. We review here the motivations for sharing neuroimaging data, the current data sharing landscape, and the sociological or technical barriers that still need to be addressed. The INCF Task Force on Neuroimaging Datasharing, in conjunction with several collaborative groups around the world, has started work on several tools to ease and eventually automate the practice of data sharing. It is hoped that such tools will allow researchers to easily share raw, processed, and derived neuroimaging data, with appropriate metadata and provenance records, and will improve the reproducibility of neuroimaging studies. By providing seamless integration of data sharing and analysis tools within a commodity research environment, the Task Force seeks to identify and minimize barriers to data sharing in the field of neuroimaging.
Introduction
The practice of data sharing is growing in society, particularly in the scientific community, as vast amounts of data continue to be acquired (Gantz and Reinsel, 2011; National Science Foundation, 2011). It mirrors an increasing demand for transparency, openness, and efficiency, and complements trends like open-source software development and open access publications. However, most of the data is not generally accessible. This review article summarizes the current state of data sharing in the field of neuroimaging, and provides recommendations for future directions.
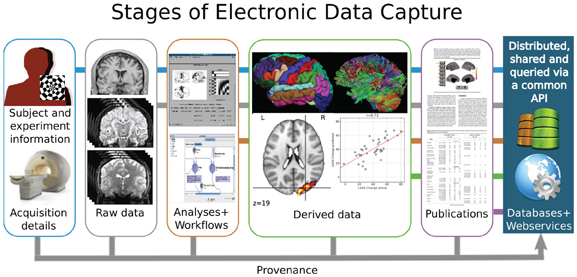
The premise that data sharing is of value to the scientific community requires that the shared data have adequate description to be of utility to those interested in its reuse. Figure 1 presents a stylized vision of neuroimaging data sharing that spans the original acquisition of images from an individual subject to the aggregation and comparison of data from groups of subjects to derive inferences into the underlying biophysical properties that correspond to observed subject characteristics. Apart from the subjects themselves, this imaging process is intrinsically a digital electronic enterprise: image acquisition, storage, processing, databasing, and sharing are all accomplished in the digital domain. Each step of this process, therefore, affords the opportunity to capture all the pertinent information that characterizes the step. Despite the seeming ease of electronic data capture (EDC) for processes that occur in the electronic domain, the neuroimaging data sharing effort has, nonetheless, often been hampered by missing and inaccurately aggregated descriptive information (metadata), which in turn has led to reduced compliance, trust, and value to the community, despite the arguably compelling philosophical or sociological rationale for data sharing. What is lacking is not the technology, but the standards, agreement, and specifications for what, how, and when to capture specific types of information in the natural course of the neuroimaging data lifecycle. In concert with other challenges to data sharing, the International Neuroinformatics Coordinating Facility (INCF) Neuroimaging Datasharing initiative is a timely and coordinated effort to propose the necessary community standards in this area.
The Section entitled “Why Should Data Be Shared?” of this review outlines a number of the benefits and rationales for greater data sharing in neuroimaging. The Section “Some Data ARE Shared” reviews ongoing neuroimaging data sharing efforts, with an emphasis on MRI data, and discusses how they can be augmented. The Section “Why Most Data AREN'T Shared” describes the barriers to data sharing and presents possible solutions to these challenges, and Section “How to Reduce Barriers to Data Sharing?” includes recommendations for future efforts to advance data sharing. Finally, the Section “The Potential Benefits of Neuroimaging Data Sharing” concludes with examples of neuroimaging initiatives that would benefit from a broader data sharing policy.
Why should Data be Shared?
To Accelerate Progress in Our Fundamental Understanding of the Brain
Several researchers have argued that more rapid scientific discoveries are possible within a culture of shared data (Poldrack, 2011; Milham, 2012), and that some questions can only be answered with large datasets or meta-analysis. Databases such as Brainmap1, SumsDB2, Neurosynth3 aggregate coordinate-based structural and functional MRI results derived from the scientific literature, and several publications have validated the use of such resources to perform meta-analytic studies. For example, Smith and colleagues (Toro et al., 2008; Smith et al., 2009) used BrainMap to perform an independent component analysis of thousands of activation maps and compared the resulting components to those extracted from resting-state MRI data. They reported substantial consistency between networks obtained using these very different datasets. More recently, Yarkoni and colleagues (Yarkoni et al., 2011) combined a similar strategy with text mining to, among other applications, accurately “decode” many cognitive states from brain activity stored in the Neurosynth database. However, such studies also point to the need for intelligent and planned data sharing, as Brainmap and Neurosynth store only (x, y, z) activation peak coordinates. Salimi-Khorshidi and colleagues showed that the reliability between a study using the original functional contrast maps and those derived from the coordinates alone was poor (Salimi-Khorshidi et al., 2009), providing an argument for the need to share original and derived images, not only the Talairach coordinates as are often published in journal articles.
The Function Biomedical Informatics Research Network (FBIRN) has accelerated progress in understanding schizophrenia using shared neuroimaging data (Glover et al., 2012). In FBIRN, each site maintains their own database and storage resources for datasets collected locally. The consortium benefits from shared access to the data which is ultimately made public after data collection is complete. Kim et al. (Kim et al., 2009) used multi-site FBIRN data to identify lateralized DLPFC dysfunction in schizophrenia using a working memory task and resting state data collected across six institutions. Potkin et al. (Potkin and Ford, 2009; Potkin et al., 2009) identified cortical disfunction in memory retrieval and decreased accuracy and reaction times by memory loads in schizophrenia using data from collected and shared across 10 institutions.
To Improve Publication and Data Quality
Above all, open data sharing allows more meaningful review of studies to be published, and fosters careful scientific enquiry (Birney et al., 2009). Greater appreciation of the fact that datasets will always have problems (missing data, noise, errors, etc.) should also be an incentive: sharing data helps uncover these errors and improves the quality of the data. For example, the 1000 Functional Connectomes team, representing a massive data release from about 30 sites and over 1000 subjects, publicly rectified occasional errors with data entry or scoring, without damage to the effort's credibility (Milham, 2012). The burden of detecting and fixing errors can encourage the use of better methods for data collection and storage, and promote EDC methods, e.g., web-based forms to capture responses directly, fMRI-compatible touchscreen tablets to record subject responses during a scan (Tam et al., 2011). Specifically, EDC holds key advantages over paper-based source documents to ensure data quality: it permits real-time validation schemes and integrity checks, as well as mechanisms to reconcile data acquired with blinded-data entry or double-data entry. It also enables additional features such as bulk import of data with automatic validation, and export functions to common analysis packages. REDCap4 (Harris et al., 2009) is a freely available software solution to deploy EDC tools for the environment.
To Reduce the Cost of Research and Increase the Return on Current Research Investments
Neuroimaging research is costly both in terms of the data acquisition costs and the significant time spent in data curation and documentation. As many funding institutions are trying to improve the cost-benefit ratio of biomedical research, the research community must find ways to do the most with what is already there. A significant amount of money could be saved from redundant data acquisition if data were shared with appropriate metadata descriptions. This savings could be redirected toward analysis and interpretation of extant data. In particular, many clinical studies acquire new data from healthy control subjects that almost certainly exist already. As data sharing becomes more prevalent, researchers who delay or choose not to release data, or who share it in a limited form (e.g., without metadata or provenance information) may find their grant applications or paper submissions criticized by their peers for neglecting data sharing.
To Foster Neuroimaging Research and Advances in Clinical Practice
One of the major challenges for the field of neuroimaging research is to generate insights that will have direct clinical relevance to the treatment of psychiatric illness (Malhi and Lagopoulos, 2008; Insel, 2009). Clinical benefits in the diagnosis and treatment of psychiatric disorders from neuroimaging research (e.g., fMRI, diffusion-weighted imaging, EEG, or MEG) may emerge from the ability to detect biomarkers in individual subjects that inform treatment decisions and predict outcome. However, in the high-dimensional space of neuroimaging studies, establishing validated image-based biomarkers of psychiatric disorders will require large numbers of subjects, specific data components, and sophisticated data processing methods. Retrospective aggregation of data from many small studies could be a useful precursor to larger, well-controlled prospective studies. For example, anatomical models fitted to large databases of subjects could be of practical use in establishing estimates of normal human brain variability with respect to age, gender, or other characteristics. In this context, paradigm independent neuroimaging data such as anatomical, diffusion-weighted, and resting-state functional data are easier to share and are gaining momentum in terms of public availability.
“Grand challenges” and competitions are a beneficial way to leverage existing data (where clinical ground-truth is known) for the development of better assessment tools and resources. An example is the recent ADHD-200 Global Competition5 challenge promoted the availability of shared ADHD resting-state data with a competition to develop the best performing classifier for identification of subject diagnosis from resting-state data. While several teams achieved significant above-chance performance, no team achieved high sensitivity and high specificity (e.g., both greater than 0.8). This includes the model that was based purely on non-imaging data. The ADHD200 competition is one demonstration of the need for large amounts of data to generate a clinically useful prediction model; another is the Biomag6 competition for MEG data.
A Requirement for Reproducible Science
An even more fundamental issue at stake in the discussion of data sharing is scientific replication. Reproducible research, or the ability to repeat another scientist's experiment and obtain consistent results, has long been considered a fundamental requirement of good scientific practice (Perneger, 2011). While computational results are essential to published experiments, only a small number will be reproduced. Despite the fundamental questions regarding the meaning of reproducibility (reproducible by whom? to what extent? etc.), the issue is attracting increased attention from funding agencies, journals, and research institutions, and has sparked a growing interest in the use of electronic lab notebooks. Scientists have long been educated in the importance of a laboratory notebook as the primary tool to record all experimental data and procedures, but its role has been complicated in the digital age, as the amount of acquired data and the number and type of analyses exceeds that which an individual researcher can readily record in detail. In order to reproduce a colleague's result, one needs to understand both how the data was acquired and what was done to the data in the processing and analysis phases of the experiment.
A growing number of workshops (e.g., “Reproducible Research: Tools and Strategies for Scientific Computing”7) are now organized around the development of electronic laboratory notebook systems, and many labs have developed open-source electronic laboratory notebooks for neuroscience researchers. In addition, projects such as Sweave8 are developing frameworks for the integration of statistical analysis and manuscript preparation. We believe that in the future, reviewers and the community should be able to access both the data and the scripts used for analyses (see the recent launch of the new journal Open Research Computation9).
Other Scientific Fields have Demonstrated the Benefits of Data Sharing
It is likely that the neuroimaging community would learn a great deal about the merits of data sharing from other scientific fields, like astronomy (see the Sloan Digital Sky Survey10), natural history (Guralnick et al., 2007; Constable et al., 2010), and genetics. The GenBank and Hapmap archives have been essential for major scientific discovery (Manolio et al., 2008), and have led to new research disciplines aimed at integrating and modeling data repositories. The field of genomics is a very clear example of how successful data sharing or data publication policies can foster scientific progress (Kaye et al., 2009).
While shared data might certainly be re-used by neuroscientists or clinical researchers, the size and complexity of neuroimaging datasets and their associated challenges have increasingly attracted communities of applied mathematicians, statisticians, image processors, data miners, and bioinformaticians who wish to apply their techniques on neuroimaging data. While their work may seem tangential to many neuroscientists, the history of science has shown that cross-disciplinary work may lead to major advances or even domain shifts of paradigm. Neuroimaging will benefit tremendously from more interactions with computer scientists, mathematicians, statisticians, etc., and a crucial first step in these collaborations will be for data to be available to those who work outside traditional neuroscience fields.
Required by Funding Agencies
Nearly 15 years ago, the US National Research Council published Bits of Power, a report on the state of data sharing and stated that “the value of data lies in their use. Full and open access to scientific data should be adopted as the international norm for the exchange of scientific data derived from publicly funded research” (National Research Council, 1997). Despite this recommendation, sharing has not become normative practice in many research disciplines, prompting several funding agencies to formalize a data sharing policy for grant recipients. In the UK, for example, “The Wellcome Trust expects all of its funded researchers to maximize the availability of research data with as few restrictions as possible,”11 “the Medical Research Council expects valuable data arising from MRC-funded research to be made available to the scientific community with as few restrictions as possible. Such data must be shared in a timely and responsible manner”12. In the United States, the National Institutes of Health (NIH) has noted “Sharing data reinforces open scientific inquiry, encourages diversity of analysis and opinion, promotes new research, makes possible the testing of new or alternative hypotheses and methods of analysis, supports studies on data collection methods and measurement, facilitates the education of new researchers, enables the exploration of topics not envisioned by the initial investigators, and permits the creation of new datasets when data from multiple sources are combined.” Further, NIH recommends “Data should be made as widely and freely available as possible while safeguarding the privacy of participants, and protecting confidential and proprietary data13. In 2007, the Organisation for Economic Co-operation and Development (OECD) released a report on the importance of sharing data obtained from publicly-funded mechanisms: “One of the central goals of promoting data access and sharing is to improve the overall efficiency of publicly funded scientific research to avoid the expensive and unnecessary duplication of data collection efforts,” and the “rapidly expanding body of research data represents both a massive investment of public funds and a potential source of the knowledge needed to address the myriad challenges facing humanity.” (Organisation for Economic Co-operation and Development, 2007).
Some Data are Shared
Much more interest in data sharing is evident in the neuroimaging community compared to just a few years ago, as a new generation of researchers recognizes its importance and utility. In many respects, the neuroimaging community has been one of the most progressive in data sharing compared to other fields of neuroscience (see, for instance, Van Horn et al., 2004; Van Horn and Ishai, 2007; Van Horn and Toga, 2009). Several major initiatives currently provide publicly available datasets, including OpenfMRI, XNAT Central, 1000 Functional Connectomes/International Neuroimaging Datasharing Initiative (Biswal et al., 2010; Milham, 2012), OASIS (Marcus et al., 2007a, 2010), and, eventually, the Human Connectome Project (Marcus et al., 2011). Still more data are available to researchers willing to do some administrative legwork to obtain access [ADNI14 (Mueller et al., 2005), NIH MRI Study of Normal Brain Development15 (Evans, 2006), NDAR16), the FBIRN consortium (Keator et al., 2008; Glover et al., 2012)]. There are also a number of organizations that are helping to foster and promote neuroimaging data sharing [e.g., the Neuroimaging Data Access Group17 (NIDAG), INCF and its Neuroimaging Data Sharing initiative, and the Biomedical Informatics Research Network (Helmer et al., 2011, BIRN18)]. The most prominent initiative with respect to EEG data sharing appears to be the Australian EEG Database, “a web-based de-identified searchable database of 18,500 EEG records recorded […] over an 11-year period” (Hunter et al., 2005). At the time of writing, there were no neuroimaging data among Amazon's public datasets19.
While few might argue with the benefits of data sharing with respect to scientific progress and the public good, the technical hurdles associated with data sharing are very real and many researchers struggle with the challenges of capturing, preparing, and releasing their data. Fortunately, a growing number of sophisticated tools that support neuroimaging data sharing have emerged during the last decade. The development of many was necessary for projects that included multiple data sites. Groups such as BIRN (Keator et al., 2008; Helmer et al., 2011) and the Alzheimer's Disease Neuroimaging Initiative (ADNI) have produced infrastructure or websites to help groups share data. For example, BIRN supplies open-source data-transfer infrastructure as well as tools to allow queries across multiple extant data sources. In addition, data management tools such as the eXtensible Neuroimaging Archive Toolkit (XNAT) (Marcus et al., 2007b) and the Human Imaging Database [HID20; (Ozyurt et al., 2010)] are freely available and work with current data movement infrastructure. Some projects better known for other purposes also have data repository components. For instance, LONI, well-known for its pipeline environment also hosts an Image Data Archive [IDA; (Dinov et al., 2010)] to help with neuroimaging data handling (such as de-identification, anonymization, upload and download, curation, search, open and user-level sharing). The Neuroimaging Informatics Tools and Resources Clearinghouse21 (NITRC), while not a data management tool per se, also hosts data, including the widely accessed 1000 Functional Connectomes/International Neuroimaging Datasharing Initiative22 (INDI). The amount of work and money put into these projects is considerable, and the technical advances of cross-platform software are opening the doors to some exciting new possibilities in various directions such as web-distributed file systems for sharing, cloud computing for pipelining, semantic web technologies for ontologies, etc.
Although some resources have emerged to help find shared neuroimaging data through federation or mediation (e.g., NIF, NITRC), most cognitive researchers cannot rely on existing shared data to pursue their projects or analyses. Federation systems typically involve multiple sources under a common schema; whereas mediation systems support variable schema, as long as the schema can be retrospectively unified or aligned. Both approaches ultimately require the concept of the overarching unifying schema or framework that, frankly, has yet to fully emerge from within the community. NIF allows researchers to query and identify sources of neuroscience data that go beyond neuroimaging data and in fact link multiple disciplines such as genetics, animal studies, pharmacology, etc., which previously were difficult to search simultaneously. That said, the broad and powerful scope of NIF's query engine perhaps make it unreasonable to expect this initiative to solve the myriad of challenges related to aggregating and deploying neuroimaging data from the many resources that have registered with it.
To date, most re-used data generally derive from large projects, such as ADNI, that have been specifically financed to make data available, and have done so by streamlining the workflow for their specific acquisition and description needs. Such projects are generally very costly; the first ADNI grant was about $60 million USD, and ADNI II is likely to be at least as expensive. Small laboratories or even individuals should be able to share their acquired data within more reasonable budgets. Despite the success of some neuroimaging data sharing initiatives, the community should turn to a more generalized and sustainable model in which small research groups could easily make their data available for re-analysis, in a distributed and lightweight manner. In the following section, we review why such data sharing is often difficult.
Why Most Data aren't Shared
The Current State
A recent PubMed search found over 22,000 fMRI-related publications between the early 1990s and October 2011. A conservative estimate of the data this represents amounts to 12,000 datasets with 12 subjects each and hour-long scans per subject, at a cost of about $300 USD/hour. This corresponds to 144,000 scan hours (around 144 TB of raw data and up to 1.5 petabytes of processed data) at a cost of about $43 million USD. However, the proportion of such data currently available in public repositories (e.g., the fMRI Data Center, 1000 Functional Connectomes, Open fMRI, OASIS, ADNI, FBIRN) is less than a few percent. Even when available publicly, the authorization required to access the data may hinder their re-distribution and use.
Why isn't More Happening?
There are many reasons why more data aren't being shared and they can be divided roughly into three categories: motivation (why should I share my data? why would I use someone else 's data?), ethical and legal issues (do I have the right to share? do I have the right to download shared data?), and technical issues (what should I share? how do I do it?). These questions may get answered differently depending on which data are being shared, when, with whom and for how long. While the lack of lightweight tools and simple procedures is an obvious barrier, it is likely not the largest. The greatest challenge may be the reconciliation of the individual researcher's desire for recognition of their work to obtain further funding and academic advancement with the larger community's expectation of greater data sharing to accelerate scientific and medical discoveries, when these scientific discoveries may indeed be made by someone other than the data collector.
Individual Motivation to Share Data
An individual's motivation (or lack thereof) to share data is a fundamental barrier. Once this is resolved, the scientific community is likely to find solutions to even the most challenging technical problems. The acquisition of imaging data is costly in both money and time, and requires training and expertise. It must be noted that the process of sharing data and then maintaining the infrastructure are costly, and in most cases, budgeted for only the duration of the grant. In a competitive environment where funding is scarce, there must be motivation to release hard-earned data. When sharing occurs soon after acquisition, many researchers fear being “scooped” by a peer, or if data are released with a publication, there is a greater risk that someone will challenge the findings or interpretations by carrying out their own data analyses. Finally, some researchers may be concerned that their research methods are not of the highest quality and that they might be viewed as incompetent if their data were exposed. Releasing data early, by definition, results in less time for a researcher to review the technical quality of their data collection and/or analytic methods and thus is a further impediment.
Ethical and Legal Issues
Even when there is the individual will and means to share data, legal, or ethical constraints may prevent researchers from doing so. As described by Kaye (Kaye et al., 2009), data are usually acquired in the context of a specific study created to address a specific research question. Research studies involve a degree of trust between subjects and researchers, the former giving their consent because they feel that the research question is worthy of their involvement and they trust that the latter will protect their privacy. Kaye noted that the obligation to share data because of funding stipulations “may be perceived as an imposition on the relationships that have been built up between researchers and participants.” While sharing of human genetic data may be more controversial than that of standard magnetic resonance images, it may not be long before sulcal and gyral “fingerprints” or functional activation patterns are sensitive enough to permit individual identification. However, technical solutions to these types of privacy concerns exist (e.g., data enclaves in which only aggregated, rather than individual, subject data are shared), and these techniques will certainly evolve and improve.
Different countries have various regulations and ethical procedures regarding the protection of human subject data. Generally, subjects have the right to withdraw their data from a study at any time, with the provision that it may not be possible to remove data that has already been shared. Informed consent documents must usually describe how data obtained from subjects will be used, and it is easier to get approval from Institutional/Ethical Review Boards (IRB/ERB) with specific research questions. Many informed consent documents do not mention the possibility of broad data sharing, thereby posing a major barrier, as it is uncommon for IRBs to grant the researcher the right to publicly distribute the data if written consent for such release wasn't requested in the original submission. In such cases, researchers wishing to share retrospective data might be permitted to do so if they were able to obtain new written informed consent for data sharing, a daunting task which for many research labs would be time-consuming and often fruitless. Further, some IRB/ERBs simply have not been willing to approve protocols that request open data sharing (Al Grudzsinksas, personal communication). As many researchers cite the amount of time that they already spend in completing IRB/ERB paperwork and related administrative tasks as a major burden (Kehagia et al., 2011), efforts to compel IRBs to be more receptive to broad data sharing should ideally take place at an institutional level.
Technical Issues
One might think that after many years of work, large and well-funded projects would have emerged with something close to a definitive answer to the technical issues associated with data sharing (BIRN, for instance, was established in 2001). Indeed, data aggregation tools to meet the requirements for large, collaborative studies, like the Human Connectome Project23, are generally available, but these are tailored to the specific project and not always easy to adapt (for instance, there is no easy way of including genome data in an XNAT database). Moreover, straightforward solutions for small- or medium-sized studies (i.e., up to a few hundred scans) like those routinely performed in cognitive neuroscience and imaging centers are still lacking widespread utilization. If one wants to share a study of 20 subjects and link the imaging data with behavioral and demographic data, the simplest solution would likely be to copy and ship the data, or make the files available on an ftp site. But this strategy will not scale with time, with the size or number of the studies, or with more than a few collaborators, nor does it readily allow for “dynamic” sharing of data.
The sharing of data raises questions about which data should (or need) be shared, and whether ethical or legal regulations permit it. With either raw or processed data, choosing the descriptive level of detail to accompany the data varies, and questions of data organization and format arise. Somewhat intertwined with the format issue is the anonymization or de-identification required before sharing can be done, and possibly the choice of a license. Then, one needs to pick a technology to make the data accessible (e.g., ftp/sftp, http/https, choice of infrastructure). This step requires technical expertise that is not always available to all laboratories. In particular, if a server is set up to expose data, the security of this server has to be appropriately handled, putting demands on IT infrastructure and/or requiring strong individual expertise. Technical solutions will also depend on the duration for which data will be hosted, and what kind of service can be provided (such as search the data with metadata, etc.). If a public resource, either commercial or non-commercial, is chosen, one needs to know how data can be pushed toward this repository.
Further, while EDC shows promise for easing metadata collection and storage, EDC solves issues of manual metadata curation at the cost of additional technical issues. For clinical and behavioral data, metadata must be captured by a system that is as easy to use as a notebook and pencil, and then stored together with the data in an appropriate format. For imaging data, few systems use the NIfTI-1 extension field (which would obviate the metadata format issue), so metadata is often stored separately from the image data and is easily lost; DICOM data, on the other hand, provides a well-defined format and space for electronic metadata management, but extensibility is a challenge. XCEDE and CDISC can handle metadata and embed base-64 encoded binary data to combine metadata with images, as well as the MINC format, but none of these solutions has yet been widely adopted.
Most current neuroimaging database systems depend on their technical storage and sharing infrastructure for metadata management and, to some extent, capture. However, capture of metadata is not yet generalized outside of these large systems.
Lack of Local Organization and Standard Descriptions, within and Outside the Laboratory
In many laboratories, data are not always well-organized locally, which makes it more difficult to describe and share data. In addition, there are no widely-adopted standards for describing data in terms of both the lexicon used and the definition of and relationship between the terms, or ontology (but see the work on the XML-Based Clinical Experiment Data Exchange Schema, or XCEDE 2.0) (Gadde et al., 2011). Each researcher who wishes to share data may propose his or her own organization and description; however, even to simply organize and document the data sufficiently so that they would be easily usable requires time and funding. An even more ambitious goal is to link and retrieve data from several sources. This would require a mapping of the terms used in each source, ideally along with a standard and widely-used lexicon and ontology (e.g., for anatomy, tasks, stimuli) or to use databases that share a common schema and natively provide for multi-site query/download such as the HID database.
Pioneering work in this area is available through the Neuroscience Information Framework (NIF) web portal24 and via Neurolex25, as well as in recent work augmenting RadLex26 to annotate neuroimaging data with ontological principles derived from the Foundational Model of Anatomy (Mejino et al., 2010; Turner et al., 2010), and which should be incorporated into neuroimaging research. Significant progress has also been made to formalize some aspects such as cognitive paradigms, e.g., CogPo (Turner and Laird, 2011). At the moment, mediation among different neuroimaging databases, such as an effort involving XNAT Central and FBIRN's HID federation (Keator et al., 2008; Ozyurt et al., 2010), requires significant programming and ad hoc mapping between the instances (Ashish et al., 2010). Both the NeuroLex and RadLex sites depend on continuous editing and updating by experts, as the task of curating this information is simply too great for any one lab or group. Realization of a standard description (data models, ontologies) would be a great step forward and could improve tools for searching relevant data across the entire web, but would require annotation of existing electronic data and metadata with the associated terms.
How to Reduce Barriers to Data Sharing?
The Push for a More Open Data Model
A number of recent examples point to a general trend to make information, particularly governmental or administrative data open to the public, within the limits of privacy protections. Last year, The Economist reported that “Barack Obama issued a presidential memorandum ordering the heads of federal agencies to make available as much information as possible […]”, and that “Providing access to data creates a culture of accountability” (“The Open Society,” 2010). The US government27 and New York City28 websites release a great amount of information and data to the public. Public transportation systems make their data available and private developers use this data to produce transit-tracking applications; the European Union also funds research on this theme (see “The Open Data Open Society Report”29). Closer to the concerns of the neuroimaging community, the British parliament released a report on the importance “of reviewing the underlying data behind research and how those data should be managed”30. Individual researchers' choices as well as institution-wide policies will be influenced by this societal trend for more open data. The current very fast expansion in social networking sites is a good reflection of how quickly people can adopt new habits, and how the society evolves with these profound technological mutations.
Funding Agencies and Journals
It has become clear that cost reduction and maximizing the impact of funding in the community will also shape tool development for sharing data, as exemplified by recent requirements from major funding agencies (NIH, Wellcome Trust, UK Medical Research Council), and more generally their shift in commitment to initiatives that help the community rather than lab-specific projects. As early as 2003, the “Final NIH Statement on Sharing Research Data”31 stated that the “NIH expects and supports the timely release and sharing of final research data from NIH-supported studies for use by other researchers,” and NIH grants above a certain funding level are also required to “to include a plan for data sharing” in their proposals.
Journals will play a key role in the process requiring that data are made available for reviewers or to the article readers. The Journal of Cognitive Neuroscience was a pioneer in this context. This initiative established a neuroimaging data repository (fMRI Data Center, or fMRIDC) that was an unprecedented success in the field, with many works published based on re-analyzed data obtained from the repository. Its success was, however, limited by the lack of standardized formats, paradigm descriptions, and analyses, as well as the limited availability of tools to query and download well-formatted data. The idea that data should be accessible remains in several high-ranked journals, with more and more supplementary material made available for reviewers and for the community [however, see The Journal of Neuroscience's recent decision to not accept supplementary material (Maunsell, 2010)]. In the future, it may be that both data and computational tools will be made available in some new form of data warehouse to help track data provenance (e.g., see the Provenance Interchange Working Group Charter32) and enable reproducibility, not withstanding the associated technical difficulties and costs. Proceedings of the National Academy of Sciences, for instance, now require that data are made available to manuscript reviewers. This again points to the need for software tools able to capture and release data at different stages of their acquisition and analysis.
Increased Citations and Visibility by Releasing Data
Researchers can receive more citations if their data are re-used. Rather than counting the number of publications, h-indices are increasingly used as a metric of output (Ball, 2007). There are now several successful examples of this re-use, such as ADNI. ADNI has yielded many publications since its launch. One specific requirement of ADNI's use agreement is that the ADNI consortium be listed on every related publication's author list. This is a very powerful means for gaining visibility and ADNI has benefited from this—its funding was renewed with ADNI2—but this policy may not meet the standards of authorship of scientific publication (Rohlfing and Poline, 2011), and generally the ADNI requirements are seen as too demanding by the community33.
It has become apparent that papers on data would be a great way to credit researchers who share data. By being cited in the same way that a piece of software or a method is cited, data acquirers could get recognition within the current academic evaluation methods for individuals (e.g., number of citations with h factor). Additionally, the peer review process will help to ensure the quality of the data and accompanying metadata. This, however, requires that journals will accept “data papers” and develop principles for how such papers should be structured and reviewed. It is also necessary that authors and journal editors consider data acquisition and sharing as an important aspect of research, on par with software or methods development. Recently, both Neuroinformatics (Kennedy et al., 2011) and BioMedCentral (Hrynaszkiewicz, 2010) have announced their intention to publish such articles. Datacite34 gathers institutions from around the world and provides them with resources to address the “why” and “how” of data citation. By working with these organizations and participating in data publishing, the neuroimaging community can help ensure appropriate credit is given for shared data.
Provide Guidelines to Help Researchers with the Ethical and Legal Aspects of Data Sharing
There is a need for localized (specific to country or founding body) guidelines on how to prepare ethics applications and anonymize data in order to share them freely, or as freely as possible. It is recommended that research protocols and informed consent documents submitted to ERB/IRBs consider possible further re-use of the data: include provision in consent forms that while subject's data will certainly be used to study the specific research question outlined in the form, limited data may also be shared (anonymously) with other researchers, for different purposes; and that subjects shouldn't participate if they are uncomfortable with such sharing. A recent survey of the UK general public found that while by far the majority of respondents would be willing to take part in neuroimaging studies for the purpose of scientific or medical research, only a small number would be willing to undergo scans for other purposes, like advertising research or insurance coverage (Wardlaw et al., 2011).
Illes and colleagues (Illes et al., 2010) have noted that many researchers feel distrust and confusion when dealing with IRBs despite their “original mission … to ensure the ethical conduct of neuro-research, may be acting as a barrier to science due to time delays, lack of expertise in protocol evaluation by these boards and inconsistent guidelines on the preparation of participant-intended documentation” (Kehagia et al., 2011). In such cases, a few individual researchers cannot single-handedly reform the approach used by their local ERB/IRB; funding agencies, institutions, and the broader scientific community need to work together on providing better information and even outreach materials. Kehalia and colleagues noted that researchers would welcome “the development and dissemination of best practices and standardized ethics review for minimally invasive neuroimaging protocols.” INCF plans to gather and make available such material.
The condition under which data may be licensed varies across countries and often depends how the data were acquired (Uhlir and Schröder, 2007). Creative Commons has done outstanding work in promoting licenses that are compatible with the broader open data movement, and which affect in no way a subject's privacy rights; some examples include all Public Library of Science (PLoS) content is published under a Creative Commons Attribution license, and the MNI's widely-used brain template ICBM-152 is compatible with this license. Note that Creative Commons itself has only one data-oriented license, CCZero, which is a dedication to the public domain, while the other licenses are intended for artistic work. For further reading on open data licensing, see Open Definition35 and Open Data Commons36, as well as Stodden (Stodden, 2009) for a review of copyright and licensing issues in a scientific context.
There is also an important interaction between the technical and ethical aspects of data sharing—what constitutes a data set that is safe to share? The degree of anonymization necessary (removing all metadata? just the name and date of birth of the subject? defacing volumetric scans?) might vary within country, region, and institution. The same concern applies to the way subjects will be informed about how their data might be used in the future. Providing clear guidelines and ready to use document templates will encourage and maximize data sharing. These guidelines and documents could be tagged with respect to their data sharing characteristics (“very open” to “very restricted”).
The need to Share the Tools: the Neurodebian Approach
Even after the legal and technical problems of data capture and sharing are resolved, there are further obstacles to address to make collaborative data analysis efficient. Typically, analysis pipelines for neuroimaging studies vary significantly across labs. They use different data formats, prefer different pre-processing schemes, require different analysis toolkits and favor different visualization techniques. Efficient collaboration in, for example, a multi-center study requires a software platform that can cope with this heterogeneity, allows for uniform deployment of all necessary research tools, and nevertheless remains easy to maintain. However, compatibility differences across software vendors and tedious installation and upgrade procedures often hinder efficiency.
Turning data sharing into efficient collaboration requires sharing of tools (Ince et al., 2012). Ideally, neuroimaging research would be based on a computing platform that can easily be shared as a whole. On one hand this would significantly lower the barrier to explore new tools and to re-use existing analysis workflows developed by other groups. On the other hand it would make sharing of software easier for the respective developers, as consolidation on a standard platform reduces demand for maintenance and support. Today, the NeuroDebian project37 is the most comprehensive effort aimed at creating a common computing platform for neuroimaging research and providing all necessary software from data capture to analysis. NeuroDebian's strategy is to integrate all relevant software into the Debian GNU/Linux operating system which offers some unique advantages in the context of neuroimaging research: it runs virtually on any hardware platform (including mobile devices), it offers the most comprehensive archive of readily usable and integrated software, it is developed as a collaborative effort by experts of their respective fields, and is free to be used, modified, and re-distributed for any purpose. Integration into Debian allows for deploying any software through a uniform and convenient interface, promotes software interoperability by a proven policy, and facilitates software maintenance via (automated) quality assurance efforts covering all integrated software. By means of hardware virtualization the advantages of this platform also benefit many users of commercial systems, such as Windows and Mac OS X (Hanke and Halchenko, 2011). For example, a NeuroDebian virtual appliance with a pre-configured XNAT neuroimaging data management platform38 allows users on any system to start using XNAT within minutes.
The Role of International Coordination
The INCF39 was established through the Global Science Forum of the OECD to develop an international neuroinformatics infrastructure, which promotes the sharing of data and computing resources to the international research community. A larger objective of the INCF is to help develop scalable, portable, and extensible applications that can be used by neuroscience laboratories worldwide. The INCF Task Force on Neuroimaging Datasharing (part of a broader scientific program on data sharing in neuroscience research40) has recently formed to address challenges in databasing and metadata that hinder effective data sharing in the neuroimaging community, and to develop standards for archiving, storing, sharing, and re-using neuroimaging data. The initial focus of this group is MRI data. Representatives from several major efforts around the world are involved.
While the neuroimaging community acknowledges the need for standards in data exchange, the definition of those standards and their acceptance and use is a difficult task involving social engineering and the coordinated efforts of many. What details are necessary to share these data and results, to reproduce the results, and to use the data for other investigations? Through feedback from the neuroimaging community via workshops and informal discussion groups, the Task Force found that while there is enthusiasm for data sharing, the average neuroimaging researcher, particularly in a small lab setting, often experiences several technical barriers that impede effective data sharing. This finding has been noted in other surveys of barriers to biomedical data sharing (Anderson et al., 2007). While certain large research groups have made great strides in establishing federated databases and metadata schemas, these solutions often still involve in-house scripts and specialized software tools, tailored to the particular workflow of a specific lab. As noted previously, a lack of standards, recommendations, and interoperable and easy-to-use tools hinder the degree to which data sharing could be adopted more generally. In an attempt to improve the tools broadly available to the community, the Task Force identified four specific projects to be carried out during 2011 and 2012:
- Creation of a “One-Click Share Tool” to allow researchers to upload MRI data (in DICOM or NIFTI format) to an XNAT database hosted at INCF. Once the data is on the central server, a quality control (QC) check is launched and the report is sent to the researcher. The raw data, metadata, and QC data are stored in the database. The QC will be initially derived from FBIRN recommendations (Glover et al., 2012) and generalized to other methods. “One-click” may only be the idealized operation of the system, but the term does express the principle that the system should be trivial to use: metadata is captured from the uploaded data itself to the extent possible, and the researcher is prompted for missing information. The system encourages best practices of EDC from data acquisition, but fills in the gaps with its own EDC and management.
- Establishment of a neuroimaging data description schema and common API to facilitate communication among databases. A number of efforts have already made progress toward that goal, of which XCEDE is probably the most well-known (Gadde et al., 2011). This standard description would be used to mediate between databases with different data models, or as a recommendation to expose a minimal set of metadata elements and a common vocabulary. Eventually, this will be linked to a set of ontologies to allow for semantic searches and reasoning.
- Introduction of a mechanism to capture related data under a single container. For example, diffusion data requires additional information in the form of diffusion gradient vectors and b-values. Most DICOM conversion utilities will write these out as two separate files. We will attempt to use the Connectome File Format (CFF) Container to store this data (Gerhard et al., 2011). This solution could be applied to other cases such as multi-echo data acquired from a single acquisition that are not handled natively by any major data format (e.g., FreeSurfer's .mgz format captures such information).
- Automatic storage of the metadata and results of processing streams to a database. Using the QC workflows as a starting point, ensure that output of these workflows can be pushed to, initially, an XNAT database. We will augment the processing to use the common application programming interface (API), extended XCEDE schema, and CFF container technology to capture processed data and metadata, in particular provenance. The first trials could be performed with Nipype (Gorgolewski et al., 2011) and PyXNAT (Schwartz et al., submitted). Recently, some first steps have been done to implement the provenance “PROV” data model developed through the World Wide Web Consortium (W3C) into neuroimaging software, to include the provenance information within the XCEDE schema and to create a vocabulary for neuroimaging that can be used to annotate data. This will allow direct and automatic capture of the standardized provenance information that could then be used to maintain appropriate metadata for processed images and in data queries. The generation of machine readable annotations of the metadata (e.g., data, and increase the effectiveness of search engines such as NIF in their ability to aggregate data from disparate sources.
The Potential Benefits of Neuroimaging Data Sharing
If data sharing in neuroimaging were to become widespread, novel and large-scale neuroimaging projects would become possible, for example:
- Meta-analysis at a large-scale using actual data. Meta-analyses can provide greater support for scientific results by aggregating findings from a number of publications that addressed the same scientific question, and a large number of such studies have been conducted in the neuroimaging literature (see, among many, Wager et al., 2003; Laird et al., 2005; Owen et al., 2005). However, none rely on the actual data to establish the results: they use (x, y, z) coordinates, various choices of filtering and preprocessing, and thresholding procedures. A much more reliable and accurate methodology would be to co-analyze the data without thresholding, and homogenize other parameters. This is especially important if the meta-analyses are to define which brain regions are involved in a set of experimental conditions, or for EEG/MEG, which set of evoked potentials are related to these conditions.
- Generalized construction of anatomical and functional atlases. Current MRI analysis procedures use digital brain atlases that are not always appropriate for the particular study. The most commonly used brain templates are distributed with analysis packages such as SPM and FSL, and may not be suitable for a specific clinical population or age group or even certain scanner characteristics. Therefore, researchers often create a new template from the images acquired in their study, normalize the subjects' brain images to this new template, and find the spatial transformation from the constructed template to a more common template, e.g., the MNI-305 T1. This method is far from optimal as the transformation may be poor, and generates one template per study, or even several templates per study. If a large proportion of acquired neuroimaging data were retrievable via the internet, it would be easy to construct a series of brain templates specifically adapted for age, scanner, pathology, gender, etc. Tools to construct specific templates that have spatial warping to the most widely adopted templates suitable for a specific study cohort could also be derived and tested.
- Defining standard brain variations in various populations. In the first instance, characterizing healthy population brain variability is crucial for both basic and clinical research. If a large enough amount of data were available, a number of projects aimed at measuring various brain traits (e.g., amount of cortical matter in a region, thickness of white matter tracts, timing of ERP) for demographic or behavioral characteristics could soon emerge. Comprehensive databases with a large amount of brain feature measurements could be constructed from cohorts acquired on different scanners and populations, thereby avoiding bias associated with acquisition parameters. These resources will soon be incremented with new interesting features and new populations, to become a large distributed neuroscience resource linked to initiatives such as the Neuroscience Information Framework.
Conclusions
It is currently difficult to imagine the full benefit of widespread data sharing. What if, in the future, a researcher interested in development of connectivity in the adolescent brain could launch a search that resembled something like: “get: brain_images = MRI_diffusion_weighted, where: subject_group_type = normal, age >12, age <15, behavioral_assessment = WISC” to find repositories over the world with diffusion weighted images for adolescents having Wisconsin Test data?
The use cases that we describe require that demographic, behavioral, or clinical data are released with neuroimaging data in a standard format and with sufficient documentation. This condition is likely to be only partly fulfilled in most cases, but we hope that the standard practice for data sharing will evolve toward a more automatic and more complete release of all associated data. We believe that the future of neuroimaging research will depend on the integration of many types of data (e.g., multi-modal imaging, imaging genetics, etc.) and the aggregation of previously acquired datasets and results from many sites.
What if in the future, all data analysis tools were able to send annotated and organized results directly to a distributed database, such as that provided by iRODS41 or Chelonia42, or use peer-to-peer distribution (e.g., see Digicode ITS43 for a distributed PACS system), so that contrasts or t-statistic maps could be accessible and easily retrieved with almost no effort by any researcher? This could be linked to a new kind of publication system based on electronic repositories that would connect raw data and their computational analyses to the interpretation of results.
Neuroimaging may then enter an age where research could lean toward knowledge management rather than data management, and the construction of electronic systems that will accumulate results and information over which some reasoning can be done, eventually helping the construction of predictive models useful in neurology, psychiatry, or cognitive neuroscience. As the overarching goal of scientific endeavor is to determine predictive models for the system under study, improvements to existing models are expected as new data are collected. Data availability is necessary for the construction of models based on large numbers of observations, as well as for the improvements or refutations of these models.
For the reasons described above, the neuroimaging community should work to generalize data sharing as well as the capture of associated metadata. This requires software tools to automatically capture, tag, and relate data with metadata. These tools, in turn, will rely on a consistent and standard metadata vocabulary, data model, and ontology. The lack of consistent metadata standards makes it difficult to curate data across research labs and for neuroimaging software to capture metadata and provenance in a systematic manner. Even when the vocabulary exists, there is a lack of digital tools to seamlessly capture and tag these metadata.
At the acquisition level, the DICOM standard allows some formalization of elements stored in the header, but several parameters relevant to brain imaging data are actually stored in private compartments of the DICOM header where no consistent nomenclature exists. At the processing level, workflow-based analysis systems (e.g., LONI pipeline44, Nipype45, CBRAIN46, FisWidgets47, Brainvisa48, Rumba49, PSOM50 etc.) and databases associated with such frameworks (e.g., XNAT, HID, IDA, COINS, LORIS, etc.51) provide us with the ability to capture the provenance of data generated by a workflow. Although we use sophisticated instruments to acquire brain imaging data along with advanced statistical and image processing methods to analyze the data, there is a distinct lack of formal ontologies and vocabularies to capture metadata together with this data, because agreeing to these vocabularies requires a coordinated effort across many countries and laboratories.
The key to achieving these goals is the ability of the community to coordinate its efforts regarding standards in data sharing. This is a sociological challenge, but can build on an already large body of work. We believe organizations like INCF, in conjunction with scientific societies and publishers, share many of these goals and together will open new avenues in brain imaging research. The integration of brain imaging with informatics tools will profoundly modify our current research methods and their impact on advances in the field.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The International Neuroinformatics Coordinating Facility provides financial support for some of the work reported in this article.
Footnotes
- ^http://www.brainmap.org
- ^http://sumsdb.wustl.edu:8081/sums/index.jsp
- ^http://neurosynth.org
- ^http://project-redcap.org/
- ^http://fcon_1000.projects.nitrc.org/indi/adhd200
- ^http://www.biomag2012.org/content/data-analysis-competition: The challenge is to decode word and category specific representations in one dataset, and long-term memory representations in another.
- ^http://www.stodden.net/AMP2011
- ^http://www.stat.uni-muenchen.de/∼leisch/Sweave/
- ^http://www.openresearchcomputation.com/
- ^http://www.sdss.org
- ^http://www.wellcome.ac.uk/About-us/Policy/Spotlight-issues/Data-sharing
- ^http://www.mrc.ac.uk/Ourresearch/Ethicsresearchguidance/Datasharinginitiative/Reports/index.htm
- ^http://grants.nih.gov/grants/policy/data_sharing/data_sharing_guidance.htm
- ^http://www.adni-info.org
- ^https://nihpd.crbs.ucsd.edu/nihpd/info/data_access.html
- ^http://ndar.nih.gov
- ^https://sites.google.com/site/nidaghome/
- ^http://www.birncommunity.org
- ^http://aws.amazon.com/publicdatasets
- ^http://www.nitrc.org/projects/hid
- ^http://www.nitrc.org
- ^http://fcon_1000.projects.nitrc.org
- ^http://www.humanconnectomeproject.org/
- ^http://www.neuinfo.org
- ^http://www.neurolex.org
- ^http://radlex.org
- ^http://www.data.gov
- ^http://www.nyc.gov
- ^http://stop.zona-m.net/2011/01/the-open-data-open-society-report-2/
- ^http://www.publications.parliament.uk/pa/cm201012/cmselect/cmsctech/856/85607.htm
- ^http://grants.nih.gov/grants/guide/notice-files/NOT-OD-03-032.html
- ^http://www.w3.org/2011/01/prov-wg-charter
- ^The ADNI policy is to be in the author line even if ADNI data were used along many other datasets. ADNI asks for a large section of the methods to be dedicated to their data, in which you also have to state who is the PI of the ADNI consortium. ADNI also asks text in the acknowledgment section, in addition to any appropriate citations.
- ^http://datacite.org
- ^http://opendefinition.org/guide/data/
- ^http://opendatacommons.org/
- ^http://neuro.debian.net
- ^http://neuro.debian.net/derivatives.html#xnat
- ^http://www.incf.org
- ^http://datasharing.incf.org
- ^http://www.irods.org
- ^http://www.nordugrid.org
- ^http://www.digicode.com
- ^http://pipeline.loni.ucla.edu/
- ^http://nipy.org
- ^http://cbrain.mcgill.ca/
- ^http://grommit.lrdc.pitt.edu/fiswidgets/
- ^http://brainvisa.info/index.html
- ^http://sites.google.com/site/rumbalab/projects/neuroinformatics/rumba-tools
- ^http://code.google.com/p/psom/, and Bellec et al., 2012.
- ^See Marcus et al., 2007b; Keator et al., 2008; Dinov et al., 2010; Scott et al., 2011; Das et al., 2011.
References
Anderson, N. R., Lee, E. S., Brockenbrough, J. S., Minie, M. E., Fuller, S., Brinkley, J., and Tarczy-Hornoch, P. (2007). Issues in biomedical research data management and analysis: needs and barriers. J. Am. Med. Inform. Assoc. 14, 478–488.
Ashish, N., Ambite, J.-L., Muslea, M., and Turner, J. A. (2010). Neuroscience data integration through mediation: an (F)BIRN case study. Front. Neuroinform. 4:118. doi: 10.3389/fninf.2010.00118
Bellec, P., Courchesne, S. L., Dickinson, P., Lerch, J., Zijdenbos, A., and Evans, A. C. (2012). The pipeline system for Octave and Matlab (PSOM): a lightweight scripting framework and execution engine for scientific workflows. Front. Neuroinform. 6, doi: 10.3389/fninf.2012.00007
Birney, E., Hudson, T. J., Green, E. D., Gunter, C., Eddy, S., Rogers, J., Harris, J. R., Ehrlich, S. D., Apweiler, R., Austin, C. P., Berglund, L., Bobrow, M., Bountra, C., Brookes, A. J., Cambon-Thomsen, A., Carter, N. P., Chisholm, R. L., Contreras, J. L., Cooke, R. M., Crosby, W. L., Dewar, K., Durbin, R., Dyke, S. O., Ecker, J. R., El Emam, K., Feuk, L., Gabriel, S. B., Gallacher, J., Gelbart, W. M., Granell, A., Guarner, F., Hubbard, T., Jackson, S. A., Jennings, J. L., Joly, Y., Jones, S. M., Kaye, J., Kennedy, K. L., Knoppers, B. M., Kyrpides, N. C., Lowrance, W. W., Luo, J., MacKay, J. J., Martín-Rivera, L., McCombie, W. R., McPherson, J. D., Miller, L., Miller, W., Moerman, D., Mooser, V., Morton, C. C., Ostell, J. M., Ouellette, B. F., Parkhill, J., Raina, P. S., Rawlings, C., Scherer, S. E., Scherer, S. W., Schofield, P. N., Sensen, C. W., Stodden, V. C., Sussman, M. R., Tanaka, T., Thornton, J., Tsunoda, T., Valle, D., Vuorio, E. I., Walker, N. M., Wallace, S., Weinstock, G., Whitman, W. B., Worley, K. C., Wu, C., Wu, J., and Yu, J. (2009). Prepublication data sharing. Nature 461, 168–170.
Biswal, B. B., Mennes, M., Zuo, X.-N., Gohel, S., Kelly, C., Smith, S. M., Beckmann, C. F., Adelstein, J. S., Buckner, R. L., Colcombe, S., Dogonowski, A. M., Ernst, M., Fair, D., Hampson, M., Hoptman, M. J., Hyde, J. S., Kiviniemi, V. J., Kötter, R., Li, S. J., Lin, C. P., Lowe, M. J., Mackay, C., Madden, D. J., Madsen, K. H., Margulies, D. S., Mayberg, H. S., McMahon, K., Monk, C. S., Mostofsky, S. H., Nagel, B. J., Pekar, J. J., Peltier, S. J., Petersen, S. E., Riedl, V., Rombouts, S. A., Rypma, B., Schlaggar, B. L., Schmidt, S., Seidler, R. D., Siegle, G. J., Sorg, C., Teng, G. J., Veijola, J., Villringer, A., Walter, M., Wang, L., Weng, X. C., Whitfield-Gabrieli, S., Williamson, P., Windischberger, C., Zang, Y. F., Zhang, H. Y., Castellanos, F. X., and Milham, M. P. (2010). Toward discovery science of human brain function. Proc. Natl. Acad. Sci. U.S.A. 107, 4734–4739.
Committee on Issues in the Transborder Flow of Scientific Data National Research Council. (1997). Bits of Power: Issues in Global Access to Scientific Data. Washington, DC: National Academies Press.
Constable, H., Guralnick, R., Wieczorek, J., Spencer, C., and Peterson, A. T. (2010). VertNet: a new model for biodiversity data sharing. PLoS Biol. 8:e1000309. doi: 10.1371/journal.pbio.1000309
D, S., Zijdenbos, A. P., Harlap, J., Vins, D., and Evans, A. C. (2011). LORIS: a web-based data management system for multi-center studies. Front. Neuroinform. doi: 10.3389/fninf.2011.00037
Dinov, I., Lozev, K., Petrosyan, P., Liu, Z., Eggert, P., Pierce, J., Zamanyan, A., Chakrapani, S., Van Horn, J., Parker, D. S., Magsipoc, R., Leung, K., Gutman, B., Woods, R., and Toga, A. (2010). Neuroimaging study designs, computational analyses and data provenance using the LONI pipeline. PloS One 5:e13070. doi: 10.1371/journal.pone.0013070
Gadde, S., Aucoin, N., Grethe, J. S., Keator, D. B., Marcus, D. S., and Pieper, S. (2011). XCEDE: an extensible schema for biomedical data. Neuroinformatics 10, 19–32.
Gantz, J., and Reinsel, D. (2011). “Extracting Value from Chaos,” June 2011, sponsored by EMC. Available from http://www.emc.com/digital_universe
Gerhard, S., Daducci, A., Lemkaddem, A., Meuli, R., Thiran, J.-P., and Hagmann, P. (2011). The connectome viewer toolkit: an open source framework to manage, analyze and visualize connectomes. Front. Neuroinform. 5:3. doi: 10.3389/fninf.2011.00003
Glover, G. H., Mueller, B. A., Turner, J. A., Van Erp, T. G., Liu, T., Greve, D., Voyvodic, J., Rasmussen, J., Brown, G., Keator, D. B., Calhoun, V. D., Lee, H., Ford, J., Mathalon, D., Diaz, M., O'eary, D., Gadde, S., Preda, A., Lim, K., Wible, C., Stern, H., Belger, A., McCarthy, G., Ozyurt, B., and Potkin, S. G. (2012). FBIRN. Function biomedical informatics research network recommendations for prospective multi-center functional magnetic resonance imaging studies. J. Magn. Reson. Imaging. (in press).
Gorgolewski, K., Burns, C. D., Madison, C., Clark, D., Halchenko, Y. O., Waskom, M. L., and Ghosh, S. S. (2011). Nipype: a flexible, lightweight and extensible neuroimaging data processing framework in Python. Front. Neuroinform. 5:13. doi: 10.3389/fninf.2011.00013
Guralnick, R. P., Hill, A. W., and Lane, M. (2007). Towards a collaborative, global infrastructure for biodiversity assessment. Ecol. Lett. 10, 663–672.
Hanke, M., and Halchenko, Y. O. (2011). Neuroscience runs on GNU/Linux. Front. Neuroinform. 5:8. doi: 10.3389/fninf.2011.00008
Harris, P. A., Taylor, R., Thielke, R., Payne, J., Gonzalez, N., and Conde, J. G. (2009). Research electronic data capture (REDCap)—a metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform. 42, 377–381.
Helmer, K. G., Ambite, J. L., Ames, J., Ananthakrishnan, R., Burns, G., Chervenak, A. L., Foster, I., Liming, L., Keator, D., Macciardi, F., Madduri, R., Navarro, J-P., Potkin, S., Rosen, B., Ruffins, S., Schuler, R., Turner, J. A., Toga, A., Williams, C., and Kesselman, C. (2011). Enabling collaborative research using the Biomedical Informatics Research Network (BIRN). J. Am. Med. Inform. Assoc. 18, 416–422.
Hrynaszkiewicz, I. (2010). A call for BMC Research Notes contributions promoting best practice in data standardization, sharing and publication. BMC Res. Notes 3, 235.
Hunter, M., Smith, R. L. L., Hyslop, W., Rosso, O. A., Gerlach, R., Rostas, J. A. P., Williams, D. B., and Henskens, F. (2005). The Australian EEG database. Clin. EEG Neurosci. 36, 76–81.
Illes, J., Tairyan, K., Federico, C. A., Tabet, A., and Glover, G. H. (2010). Reducing barriers to ethics in neuroscience. Front. Hum. Neurosci. 4:167. 1–5. doi: 10.3389/fnhum.2010.00167
Ince, D. C., Hatton, L., and Graham-Cumming, J. (2012). The case for open computer programs. Nature 482, 485–488.
Insel, T. R. (2009). Translating scientific opportunity into public health impact: a strategic plan for research on mental illness. Arch. Gen. Psychiatry 66, 128–133.
Kaye, J., Heeney, C., Hawkins, N., de Vries, J., and Boddington, P. (2009). Data sharing in genomics re-shaping scientific practice. Nat. Rev. Genet. 10, 331–335.
Keator, D. B., Grethe, J. S., Marcus, D., Ozyurt, B., Gadde, S., Murphy, S., Pieper, S., Greve, D., Notestine, R., Bockholt, H. J., Papadopoulos, P., and BIRN Function, BIRN Morphometry and BIRN Coordinating. (2008). A national human neuroimaging collaboratory enabled by the Biomedical Informatics Research Network (BIRN). IEEE Trans. Inf. Technol. Biomed. 12, 162–172. doi: 10.1109/TITB.2008.917893
Kehagia, A. A., Tairyan, K, Federico, C., Glover, G. H., and Illes, J. (2011). More education, less administration: reflections of neuroimagers' attitudes to ethics through the qualitative looking glass. Sci. Eng. Ethics. (in press).
Kennedy, D. N., Ascoli, G. A., and De Schutter, E. (2011). Next steps in data publishing. Neuroinformatics. 9, 317–320.
Kim, D. I., Manoach, D. S., Mathalon, D. H., Turner, J. A., Mannell, M., Brown, G. G., Ford, J. M., Gollub, R. L., White, T., Wible, C., Belger, A., Bockholt, H. J., Clark, V. P., Lauriello, J., O'Leary, D., Mueller, B. A., Lim, K. O., Andreasen, N., Potkin, S. G., and Calhoun, V. D. (2009). Dysregulation of working memory and default-mode networks in schizophrenia using independent component analysis, an fBIRN and MCIC study. Hum. Brain Mapp. 30, 3795–3811.
Laird, A. R., Fox, P. M., Price, C. P., Glahn, D. C., Uecker, A. M., Lancaster, J. L., Turkeltaub, P. E., Kochunov, P., and Fox, P. T. (2005). ALE meta-analysis: controlling the false discovery rate and performing statistical contrasts. Hum. Brain Mapp. 25, 155–164.
Malhi, G. S., and Lagopoulos, J. (2008). Making sense of neuroimaging in psychiatry. Acta Psychiatr. Scand. 117, 100–117.
Manolio, T. A., Brooks, L. D., and Collins, F. S. (2008). A HapMap harvest of insights into the genetics of common disease. J. Clin. Invest. 118, 1590–1605. doi: 10.1172/JCI34772
Marcus, D. S., Archie, K. A., Olsen, T. R., and Ramaratnam, M. (2007a). The open-source neuroimaging research enterprise. J. Digit. Imaging 20(Suppl. 1), 130–138.
Marcus, D. S., Olsen, T. R., Ramaratnam, M., and Buckner, R. L. (2007b). The extensible neuroimaging archive toolkit. Neuroinformatics 5, 11–33.
Marcus, D. S., Fotenos, A. F., Csernansky, J. G., Morris, J. C., and Buckner, R. L. (2010). Open access series of imaging studies: longitudinal MRI data in nondemented and demented older adults. J. Cogn. Neurosci. 22, 2677–2684.
Marcus, D. S., Harwell, J., Olsen, T., Hodge, M., Glasser, M. F., Prior, F., Jenkinson, M., Laumann, T., Curtiss, S. W., and Van Essen, D. C. (2011). Informatics and data mining tools and strategies for the human connectome project. Front. Neuroinform. 5:4. doi: 10.3389/fninf.2011.00004
Mejino, J. L. V., Detwiler, L. T., Turner, J. A., Martone, M. E., Rubin, D. L., and Brinkley, J. F. (2010). Enabling RadLex with the foundational model of anatomy ontology to organize and integrate neuro-imaging data (Washington, DC). AMIA 2010 Symposium Proceedings. 1171.
Milham, M. P. (2012). Open neuroscience solutions for the connectome-wide association era. Neuron 73, 214–218.
Mueller, S. G., Weiner, M. W., Thal, L. J., Petersen, R. C., Jack, C., Jagust, W., and Trojanowski, J. Q. et al. (2005). The Alzheimer's disease neuroimaging initiative. Neuroimaging Clin. N. Am. 15, 869–877.
National Science Foundation. (2011). Changing the conduct of science in the information age. http://www.nsf.gov/pubs/2011/oise11003/
Organisation for Economic Co-operation and Development. (2007). OECD Principles and Guidelines for Access to Research Data from Public Funding (No. 88180 2007). Paris, France. http://www.oecd.org/dataoecd/9/61/38500813.pdf
Owen, A. M., McMillan, K. M., Laird, A. R., and Bullmore, E. (2005). N-back working memory paradigm: a meta-analysis of normative functional neuroimaging studies. Hum. Brain Mapp. 25, 46–59.
Ozyurt, I. B., Keator, D., Wei, D., Fennema-Notestine, C., Pease, K., Bockholt, B., and Grethe, J. (2010). Federated web-accessible clinical data management within an extensible neuroimaging database. Neuroinformatics 23, 98–106.
Potkin, S. G., Turner, J. A., Brown, G. G., McCarthy, G., Greve, D. N., Glover, G. H., Manoach, D. S., Belger, A., Diaz, M., Wible, C. G., Ford, J. M., Mathalon, D. H., Gollub, R., Lauriello, J., O'Leary, D., van Erp, T. G. M., Toga, A. W., Preda, A., and Lim, K. O. (2009). Working memory and DLPFC inefficiency in schizophrenia: the FBIRN study Schizophrenia bulletin, MPRC 35, 19–31.
Potkin, S., and Ford, J. (2009). Widespread cortical dysfunction in schizophrenia: the FBIRN imaging consortium Schizophrenia bulletin. MPRC 35, 15–18.
Salimi-Khorshidi, G., Smith, S. M., Keltner, J. R., Wager, T. D., and Nichols, T. E. (2009). Meta-analysis of neuroimaging data: a comparison of image-based and coordinate-based pooling of studies. Neuroimage 45, 810–823.
Scott, A., Courtney, W., Wood, D., dela Garza, R., Lane, S., King, M., and Wang, R. et al. (2011). COINS: an innovative informatics and neuroimaging tool suite built for large heterogeneous datasets. Front. Neuroinform. 5, 1–15. doi: 10.3389/fninf.2011.00033
Smith, S. M., Fox, P. T., Miller, K. L., Glahn, D. C., Fox, P. M., Mackay, C. E., Filippini, N., Watkins, K. E., Toro, R., Laird, A. R., and Beckmann, C. F. (2009). Correspondence of the brain's functional architecture during activation and rest. Proc. Natl. Acad. Sci. U.S.A. 106, 13040–13045.
Stodden, V. (2009). Enabling reproducible research: open licensing for scientific innovation. Int. J. Commun. Law Policy 13, 1–25.
Rohlfing, T., and Poline, J.-B. (2011). Why shared data should not be acknowledged on the author byline. Neuroimage 59, 4189–4195.
Tam, F., Churchill, N. W., Strother, S. C., and Graham, S. J. (2011). A new tablet for writing and drawing during functional MRI. Hum. Brain Mapp. 32, 240–248.
The Open Society. (2010, February). Economist. London, UK: The Economist Newspaper Ltd. http://www.economist.com/node/15557477
Toro, R., Fox, P. T., and Paus, T. (2008). Functional coactivation map of the human brain. Cereb. Cortex 18, 2553–2559.
Turner, J. A., Mejino, Jose L. V., Brinkley, J. F., Detwiler, L. T., Lee, H. J., Martone, M. E., and Rubin, D. L. (2010). Application of neuroanatomical ontologies for neuroimaging data annotation. Front. Neuroinform. 4, 1–12. doi: 10.3389/fninf.2010.00010
Turner, J. A., and Laird, A. R. (2011). The cognitive paradigm ontology: design and application. Neuroinformatics 10, 57–66.
Van Horn, J. D., Grafton, S. T., Rockmore, D., and Gazzaniga, M. S. (2004). Sharing neuroimaging studies of human cognition. Nat. Neurosci. 7, 473–481.
Van Horn, J. D., and Ishai, A. (2007). Mapping the human brain: new insights from FMRI data sharing. Neuroinformatics 5, 146–153.
Van Horn, J. D., and Toga, A. W. (2009). Is it time to re-prioritize neuroimaging databases and digital repositories? Neuroimage 47, 1720–1734.
Wager, T. D., Phan, K. L., Liberzon, I., and Taylor, S. F. (2003). Valence, gender, and lateralization of functional brain anatomy in emotion: a meta-analysis of findings from neuroimaging. Neuroimage 19, 513–531.
Wardlaw, J. M., O'Connell, G., Shuler, K., DeWilde, J., Haley, J., Escobar, O., and Murray, S. et al. (2011). “Can it read my mind?” – What do the public and experts think of the current (mis)uses of neuroimaging? PLoS One 6:e25829. doi: 10.1371/journal.pone. 0025829
Keywords: brain imaging, data sharing, standards, magnetic resonance imaging, fMRI, EEG-MEG
Citation: Poline J-B, Breeze JL, Ghosh S, Gorgolewski KF, Halchenko YO, Hanke M, Haselgrove C, Helmer KG, Keator DB, Marcus DS, Poldrack RA, Schwartz Y, Ashburner J and Kennedy DN (2012) Data sharing in neuroimaging research. Front. Neuroinform. 6:9.doi: 10.3389/fninf.2012.00009
Received: 13 October 2011; Accepted: 09 March 2012;
Published online: 05 April 2012.
Edited by:
Jessica A. Turner, Mind Research Network, Albuquerque, USAReviewed by:
Lars Schwabe, University of Rostock, GermanyJohn Van Horn, University of California at Los Angeles, USA
Copyright: © 2012 Poline, Breeze, Ghosh, Gorgolewski, Halchenko, Hanke, Haselgrove, Helmer, Keator, Marcus, Poldrack, Schwartz, Ashburner and Kennedy. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Jean-Baptiste Poline, Neurospin, Bat. 145, CEA, Gif-sur-Yvette, 91191, France.
Henry Wheeler Brain Imaging Center, 10 Giannini Hall, UC Berkeley, CA, USA. e-mail: jbpoline@gmail.com