Corrigendum: Can the natural diversity of quorum-sensing advance synthetic biology?
- 1Ira A. Fulton School of Biological and Health Systems Engineering, Arizona State University, Tempe, AZ, USA
- 2Biological Design Graduate Program, Arizona State University, Tempe, AZ, USA
- 3Department of Chemistry and Biochemistry, Arizona State University, Tempe, AZ, USA
- 4School of Life Sciences, Arizona State University, Tempe, AZ, USA
A corrigendum on
The gene WP_023917333 was incorrectly used to generate the GtaR protein motif map in Figure 5 of the original publication, which led us to publish erroneous conclusions about GtaR structure and function (Davis et al., 2015). At the time this manuscript was published, the gene WP_023917333 was incorrectly titled “LuxR family transcriptional regulator Rhodobacter capsulatus” in the NCBI database. Analysis of the correct GtaR protein sequence (WP_013066073) does not show “sequence conservation with TatD family of deoxyribonuclease proteins” nor does it lead us to conclude that GtaR “might represent a unique class of HSL-responsive regulator proteins” (Davis et al., 2015).
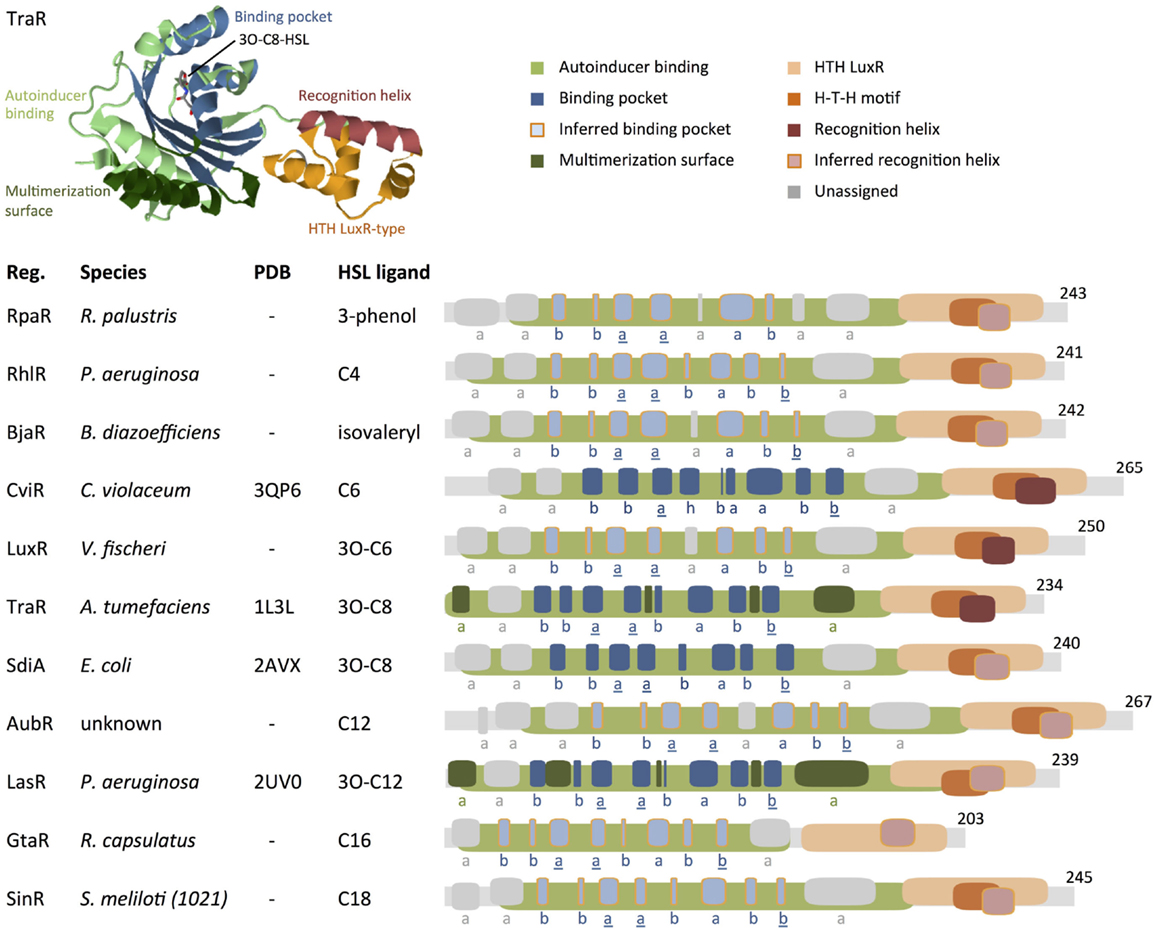
Figure 5. Comparison of protein motifs in select regulators. The three-dimensional (3D) structure of TraR is shown as an example of how domains and the homoserine lactone (HSL) ligands are typically positioned in space. The underlined letters in the b–b–a–a–b–a–b–b secondary structure motif indicate the location of highly conserved amino acids that form hydrogen bonds with the homoserine lactone head of HSLs. Published 3D structure data (Protein Data Bank) are listed where available (“–” = not available). Abbreviations used are: Reg. = regulator protein, H–T–H = helix–turn–helix, a = alpha helix, b = beta strand, h = 3/10 helix. Database entries for conserved motifs are: autoinducer binding = IPR005143 (Mitchell et al., 2015), HTH LuxR = SM00421 (Schultz et al., 1998; Letunic et al., 2015). Inferred binding pockets are patterns of secondary structures that are similar to the TraR-binding pocket. Inferred recognition helices are the second alpha helix from the C-terminus. Secondary structures for proteins with no available 3D structure data were mapped using the Jpred prediction tool (Cole et al., 2008). Maps were generated using DomainDraw (Fink and Hamilton, 2007). Figure modified from Davis et al. (2015).
Analysis of the correct protein sequence (WP_013066073) shows that GtaR contains the same b–b–a–a–b–a–b–b motif as RhlR, LasR, and SinR. All four regulator proteins, RhlR, LasR, GtaR, and SinR, respond to different ligands: C4-HSL, 3O-C12-HSL, C16-HSL, and C18-HSL, respectively (Llamas et al., 2004; Kumari et al., 2006; Geske et al., 2008; Leung et al., 2012). We surmise that variations in specific residues may account for the regulator proteins’ preferences for different ligands.
The GtaR C-terminus does not match the HTH LuxR-type motif (Prosite PS50043) originally annotated in Figure 5 but does match an “HTH_LuxR” DNA-binding motif designated as SMART motif SM00421 (Schultz et al., 1998; Letunic et al., 2015) at amino acids 140–197. This motif is present in all the regulators analyzed. Figure 5 now illustrates HTH LuxR regions (SMART SM00421) instead of PS50043. Furthermore, in the original publication, the protein motif maps were switched between LasR and AubR, and the SidA map was scaled incorrectly. We have corrected these errors in a new version of Figure 5.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors are grateful to Professor Tom Beatty for bringing these errors to our attention.
References
Cole, C., Barber, J., and Barton, G. (2008). The Jpred 3 secondary structure prediction server. Nucleic Acids Res. 1, W197–W201. doi:10.1093/nar/gkn238
Davis, R., Muller, R., and Haynes, K. (2015). Can the natural diversity of quorum-sensing advance synthetic biology? Front. Bioeng. Biotechnol. 3:30. doi:10.3389/fbioe.2015.00030
Fink, J., and Hamilton, N. (2007). Domaindraw: a macromolecular feature drawing program. In silico Biol. 7, 145–150.
Geske, G., Mattmann, M., and Blackwell, H. (2008). Evaluation of a focused library of N-aryl l-homoserine lactones reveals a new set of potent quorum sensing modulators. Bioorg. Med. Chem. Lett. 18, 5978–5981. doi:10.1016/j.bmcl.2008.07.089
Kumari, A., Pasini, P., Deo, S. K., Flomenhoft, D., Shashidhar, H., and Daunert, S. (2006). Biosensing systems for the detection of bacterial quorum signaling molecules. Anal. Chem. 78, 7603–7609. doi:10.1021/ac061421n
Letunic, I., Doerks, T., and Bork, P. (2015). SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 43, D257–D260. doi:10.1093/nar/gku949
Leung, M., Brimacombe, C., Spiegelman, G., and Beatty, J. (2012). The GtaR protein negatively regulates transcription of the gtaRI operon and modulates gene transfer agent (RcGTA) expression in Rhodobacter capsulatus. Mol. Microbiol. 83, 759–774. doi:10.1111/j.1365-2958.2011.07963.x
Llamas, I., Keshavan, N., and González, J. (2004). Use of Sinorhizobium meliloti as an indicator for specific detection of long-chain N-acyl homoserine lactones. Appl. Environ. Microbiol. 70, 3715–3723. doi:10.1128/AEM.70.6.3715-3723.2004
Mitchell, A., Chang, H., Daugherty, L., Fraser, M., Hunter, S., Lopez, R., et al. (2015). The Interpro protein families database: the classification resource after 15 years. Nucleic Acids Res. 43, D213–D221. doi:10.1093/nar/gku1243
Keywords: quorum sensing, homoserine lactone, crosstalk, orthogonal, genetic wire, synthetic gene circuit
Citation: Davis RM, Muller RY and Haynes KA (2015) Corrigendum: Can the natural diversity of quorum-sensing advance synthetic biology? Front. Bioeng. Biotechnol. 3:99. doi: 10.3389/fbioe.2015.00099
Received: 04 May 2015; Accepted: 22 June 2015;
Published: 07 July 2015
Edited by:
Zhanglin Lin, Tsinghua University, ChinaReviewed by:
Baojun Wang, University of Edinburgh, UKJesus Picó, Universitat Politecnica de Valencia, Spain
Copyright: © 2015 Davis, Muller and Haynes. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: René Michele Davis, rene.davis@asu.edu
 René Michele Davis
René Michele Davis Ryan Yue Muller
Ryan Yue Muller Karmella Ann Haynes
Karmella Ann Haynes