- 1Department of Basic Sciences and Aquatic medicine, Norwegian University of Life Sciences, Oslo, Norway
- 2Institute for Experimental Medical Research, Oslo University Hospital, Oslo, Norway
- 3Department of Biosciences, University of Oslo, Oslo, Norway
- 4Department of Aquatic Bioscience, Graduate School of Agricultural and Life Sciences, The University of Tokyo, Bunkyo, Japan
RFamides (RFa) are neuropeptides involved in many different physiological processes in vertebrates, such as reproductive behavior, pubertal activation of the reproductive endocrine axis, control of feeding behavior, and pain modulation. As research has focused mostly on their role in adult vertebrates, the possible roles of these peptides during development are poorly understood. However, the few studies that exist show that RFa are expressed early in development in different vertebrate classes, perhaps mostly associated with the central nervous system. Interestingly, the related peptide family of FMRFa has been shown to be important for brain development in invertebrates. In a teleost, the Japanese medaka, knockdown of genes in the Kiss system indicates that Kiss ligands and receptors are vital for brain development, but few other functional studies exist. Here, we review the literature of RFa in early vertebrate development, including the possible functional roles these peptides may play.
Introduction
Neuropeptides with an arginine (R) and an amidated phenylalanine (F)-motif at its C-end (called RFamides or RFa) were first described in mollusks in the 70s [FMRFamide (FMRFa)] (1). Soon after, an RFa was also found in a vertebrate (2). Since then, many RFa have been identified in different vertebrate species, with the most recent group of vertebrate RFa found as late as 2002 (3). Vertebrate RFa are currently divided into five groups: (i) neuropeptide FF (NPFF) group or PQRFa group, consisting of NPFF, neuropeptide AF (NPAF), and neuropeptide SF (NPSF); (ii) prolactin-releasing peptide (PrRP) group, consisting of PrRP20 and PrRP31, crucian carp RFamide (C-RFa), and salmon RFa; (iii) gonadotropin-inhibitory hormone (GnIH) group, including mammalian RFa-related peptides (RFRP-1 and RFRP-3), frog growth hormone-releasing peptide (fGRP), and goldfish LPXRFamide peptide; (iv) kisspeptin group also known as metastin; and finally (v) 26RFa/QRFP, including the peptides 26RFa and 43RFa (QRFP) (4, 5). These peptides have been studied in several different physiological contexts, and are found to have a role in a wide range of processes in vertebrates, as in reproductive behavior and in control of the reproductive axis (6), in pain modulation (7), and in control of feeding (8). However, much more research is needed to fully comprehend the function of RFa in different processes.
Interestingly, FMRFa are expressed in the nervous system at very early developmental stages in several phyla of metazoans, as mollusks (cephalopods and gastropods) (9, 10), cnidarians (11), and annelids (a polychete) (12). This may indicate an evolutionary ancient role of FMRFa in the development of the nervous system. It has also been shown that regeneration of flatworm anterior body fragments are stimulated by RFa (13), further supporting a role for RFa in development of tissues in lower metazoans. Also in vertebrates, the developmental studies of RFa show exciting results. Common for all the peptides is that they seem to be expressed at a very early stage in most vertebrate species studied, ranging from fishes to mammals. It seems that RFa could have important roles in development not yet discovered. This review aims to sum up what is known about the temporal and spatial expression pattern, as well as potential functional roles of the different RFa in vertebrate development, using a comparative approach.
NPFF Group
The first RFa to be identified in vertebrates was NPFF (also known as F8F-amide) and NPAF (2). Also belonging to this group is the peptide NPSF, and all three peptides are transcribed from the same gene in mammals (5). Genes encoding for NPFF have been identified in many different vertebrate classes, from hagfish to mammals, and members of this group bind to the receptor NPFFR2 (also called GPR74, NPFF2) (5).
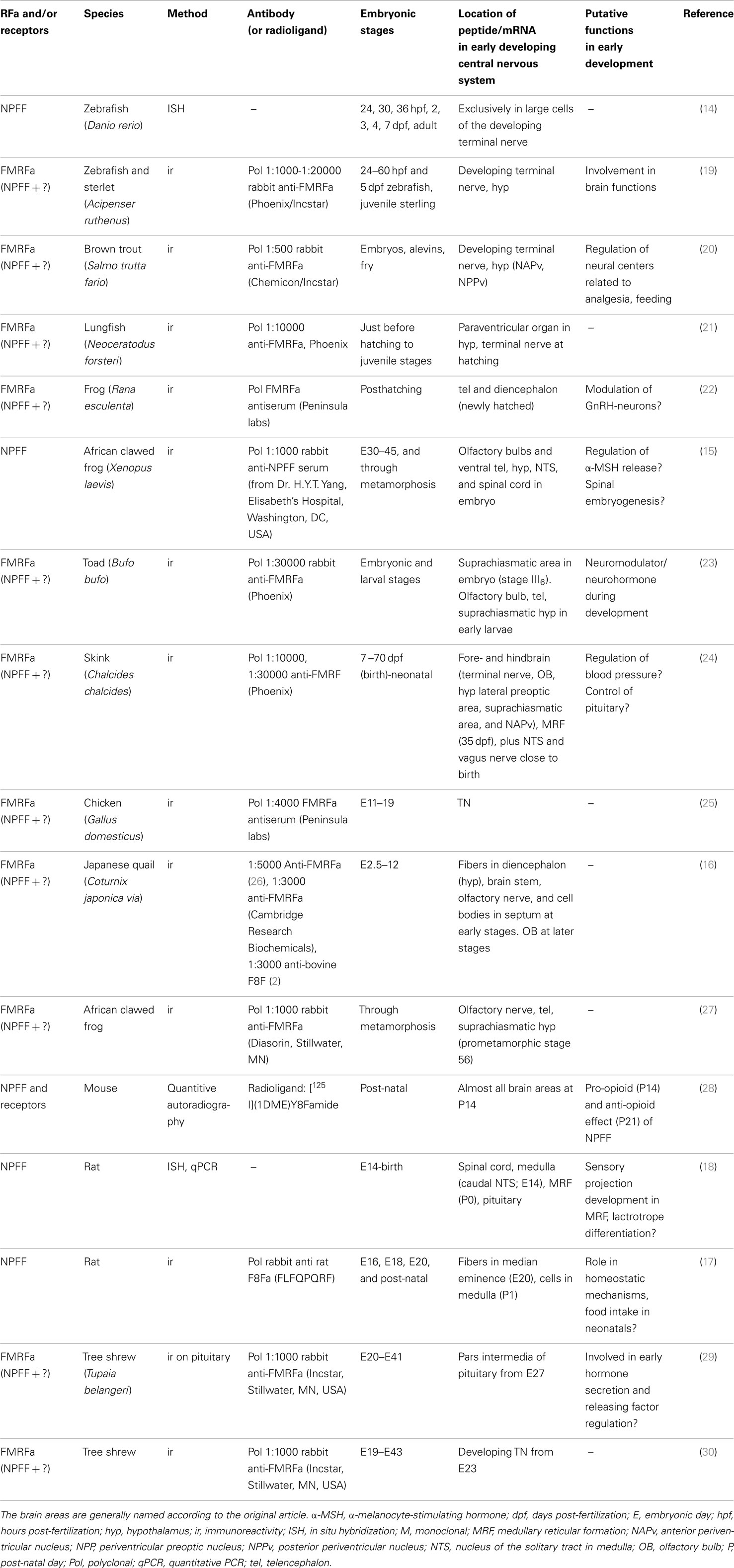
Few studies have investigated the expression, location, or function of NPFF during development. However, one study in teleosts (14), one in amphibians (15), one in birds (16), and two studies in mammals (17, 18) show that this peptide is expressed early in embryonic life in different vertebrate classes. In addition, many studies have used polyclonal antibodies against FMRFa that also seem to label the NPFF peptide (see later, and Table 1), providing additional information regarding NPFF expression patterns. Below, a brief description of where NPFF is expressed in adult vertebrates is included, followed by a description of the studies performed during development.
NPFF in Adult Vertebrates
In adult agnathans, NPFF RFa has been found expressed in the hypothalamus. Furthermore, it has been shown that NPFF stimulates the expression of the gonadotropin-β gene in the pituitary of hagfish, which suggests that NPFF can have a role in control of reproduction in lower vertebrates (31). In adult teleost fishes, NPFF seems to be exclusively expressed in gonadotropin releasing hormone 3 (GnRH3) neurons of the terminal nerve (TN) that stretches parallel to the olfactory nerve from the olfactory organ to the nucleus olfactoretinalis in the telencephalon (14). The TN, also known as the cranial nerve 0 (or N), was first described in sharks over 100 years ago, and later it has become clear that most vertebrates possess this nerve, from teleosts to primates, although the function is still not fully understood (32). One characteristic of these cells is that they express one variant of GnRH (32), which, as mentioned above, is GnRH3 in fish. The TN cell bodies are located parallel to the olfactory nerve, through the olfactory bulb to the telencephalon, but their axons project throughout the brain, affecting many different behaviors, especially reproduction-related behavior (33–35). Since it has been shown that NPFF can inhibit pacemaker activity of TN GnRH-cells, NPFF is believed to be involved in the regulation of reproductive behavior in fishes (36). In amphibians, NPFF-immunoreactive (NPFFir) cells have been identified in the preoptic area and hypothalamus (primarily the suprachiasmatic region), and extensive networks of NPFFir fibers are found throughout the brain, such as in the telencephalon, hypothalamus, medulla, and dorsal spinal cord (15, 37–39). In addition, many amphibians show NPFFir of the TN cells and fibers, similar to the findings in fishes. However, there seem to be species-specific variations regarding expression of NPFF in this site in amphibians (40). To our knowledge, the roles of NPFF in amphibians, reptiles, and birds are unknown. In rodents, NPFF cell bodies have not been identified in anterior brain regions, but are instead found in the hypothalamus, medulla, and spinal cord (41, 42). However, a dense network of NPFF fibers extends throughout most of the brain, and also to the pituitary, suggesting that NPFF can be involved in a range of different processes in mammals. In addition, there seem to be species-specific differences in the location of NPFF in mammals, since bovine cortex and hippocampus contains NPFFir (43). The known effects of NPFF are very diverse in mammals; most importantly NPFF has been found to act as a neuromodulator in the opioid system, but has also been found to increase arterial blood pressure, reduce water intake, inhibit the release of vasopressin from the neurohypophysis, and influence food intake (6, 44).
NPFF in Fish Development
In zebrafish (Danio rerio), NPFF expression first appears already at 30 h post-fertilization (hpf; faringula period) in a small cell cluster just ventral to the olfactory placode (in situ hybridization) (14). The NPFF-positive cells co-express GnRH3, a marker of ganglion cells of the TN. These cells are also marked with FMRFa polyclonal antibodies, both in 30 hpf zebrafish embryos and juvenile sturgeon (sterlet, Acipenser ruthenus) (19), as well as in early embryos of trout (Salmo trutta fario) (20) and lungfish (Neoceratodus forsteri) (21). However, in the latter three studies, at least one additional cluster of cell bodies was marked with FMRFa immunoreactivity (FMRFir), situated in the diencephalon (more specifically, the periventricular hypothalamus in zebrafish and trout, circumventricular regions of hypothalamus in sterlet, and the paraventricular organ in lungfish). These cells were not labeled with in situ hybridization in zebrafish. Thus it seems that the FMRFa polyclonal antibodies bind one or several other RFa in addition to NPFF, making it challenging to interpret immunohistochemical data for FMRFa (14, 19). At 2 dpf, NPFF-expressing cells are breaking off from the small cell cluster near the olfactory placode and start caudal migration, forming a chain along the TN trajectory in zebrafish (in situ hybridization) (14). This pattern is also seen with FMRFa-immunohistochemistry in zebrafish, sturgeon, and lungfish (19, 21), and is proposed to visualize a migration route for cells of the TN from the olfactory placode to the nucleus olfactoretinalis in the telencephalon (19). The origin of the TN cells was believed to be the olfactory placode, but more recent studies done in zebrafish have shown that the cells of the TN originate from the neural crest and then invade the olfactory placode and later migrate from the olfactory placode area to anterior brain regions (45, 46). The hypothalamus and the spinal cord of the zebrafish did not show any NPFF-labeled cells with in situ hybridization, neither in the embryo nor the adult (14). This pattern seems to be in contrast to the pattern in other vertebrates, where NPFF-expressing cell bodies can be found in other brain areas during development (see NPFF in Amphibian, Reptile, and Avian Development and NPFF in Mammalian Development).
NPFF in Amphibian, Reptile, and Avian Development
Similar to teleosts and lungfish, FMRFa-immunohistochemistry labels neurons of the TN of African clawed frog (Xenopus laevis), also during development. Using an anti-NPFF serum produced in rabbit, Lopez et al. (15) observed NPFFir cells in the embryonic olfactory placode, which is attached to the developing telencephalon at stage 40 in Xenopus. Later (stage 43), NPFFir cells could also be seen in the ventral part of olfactory bulb, and the developing telencephalon, rostral to the anterior commissure. Similar labeling of the developing TN is also found with less specific antibodies for FMRFa in Xenopus (27) and also in other amphibians, such as European green frog (Rana esculenta) (22) and toad (Bufo bufo) (23), in the reptile skink (Chalcides chalcides) (24), and in the birds Japanese quail (Coturnix japonicavia) (16) and chicken (25). Similar to zebrafish, the cells of the TN in these studies migrate from the olfactory placode, along the olfactory bulbs, to the telencephalon. Destruction of the olfactory placode in the toad embryo leads to elimination of FMRFa cells in the olfactory bulbs, ventral telencephalon, and anterior preoptic area, but not the cells in the hypothalamus (see next paragraph), showing that as in fish, the NPFF neurons of the TN migrate from the olfactory placode (23). Also in Japanese quail, the migration of FMRFa (F8Fa) neurons of the TN is similar to fish and amphibians (16), showing that this is an evolutionary conserved feature of the TN.
In contrast to the apparent situation in teleosts, the developing and the adult brains of amphibians and birds show presence of NPFF mRNA and protein also in brain areas other than the TN (15, 16). The main population of NPFFir cells in Xenopus embryos is found in the suprachiasmatic region in the hypothalamus, and these cells appear earlier than the neurons of the TN (15). This area projects to the intermediate lobe of the pituitary and is involved in the control of body color in Xenopus through the control of α-melanocyte-stimulating hormone (α-MSH) that stimulates the melanophores in the skin (47). The suprachiasmatic cells were immunoreactive very early in development (stage 30), before production of α-MSH starts, suggesting that NPFF may be involved in the control of melanotrope cell development (15). The hypothalamic neurons seem to innervate the tectum, torus semicirculris, and tegmentum in the mesencephalon, and the innervation increases during development in Xenopus (34). The suprachiasmatic region is also labeled with FMRFa antibodies early in embryonic Xenopus, toad, European green frog, and skink (22–24, 27, 39). Placodectomy studies in both birds and amphibians have shown that this population of cells has a different developmental origin than the FMRFir TN cells (16, 23, 39, 48).
Interestingly, neurons in the nucleus of the solitary tract in the medulla show NPFFir at an early stage in Xenopus (20). This area projects to the parabranchial region and the innervation is involved in the control of feeding in mammals (49). The cell bodies in the nucleus of the solitary tract seem to decrease in their immunoreactivity during development, but the projection to the parabranchial area persists and this area is heavily innervated with NPFFir fibers when the tadpoles start feeding (15). Also in developing mammals, NPFF cells are found in the nucleus of the solitary tract (see next section). At the climax of metamorphosis, NPFFir cells are detected in the reticular formation of the brain stem (15), also similar to findings in mammals. In the spinal cord, NPFFir elements were detected early in development, first in rostral spinal segments and later in the thoracic and upper lumbar segments. Interestingly, the NPFFir intensity was higher in the Xenopus spinal cord than in the adult, suggesting that NPFF has a developmental role in spinal cord embryogenesis. At the end of metamorphosis, the adult pattern of NPFFir structures in Xenopus is established (15). The brain stem and the spinal cord is not labeled with FMRFa antibodies in frog or toad (22, 23), but the skink shows FMRFir in these locations during development (24).
In the Japanese quail, the first NPFFir (F8F) fibers appear in the diencephalon (later hypothalamus) and the brain stem at embryonic stage (E) 6 (16). Later, also fibers in the olfactory nerve and the septum are labeled. In later developmental stages (E12), fibers and cell bodies are seen in the already mentioned areas in addition to the olfactory bulb. The location of NPFFir cells is similar to GnRHir cells.
NPFF in Mammalian Development
Using an antibody against rat NPFF (F8F-amide), Kivipelto et al. showed the presence of fibers and terminal-like structures as early as E20 in the rat (Rattus norvegicus) (17). The labeling was seen in the internal layer of the median eminence in hypothalamus, an area important for control of pituitary hormone secretion. However, no labeled cell bodies were detected anywhere in the brain at this stage. At birth, NPFFir cells were found in the caudal part of the medial nucleus of the solitary tract in medulla, parallel to findings in Xenopus. As mentioned, this area is involved in control of feeding through its innervation to the lateral parabranchial nucleus (an area associated with feeding control) (50). In accordance with the findings in Xenopus, a relatively dense area of NPFFir terminals and fibers were found in the parabranchial nucleus in the rat (17). In addition, fibers and terminals were found in numerous other parts of the brain; for instance paraventricular hypothalamic area, supraoptic nucleus, optic decussation, and the periventricular hypothalamic area. Dense networks of fibers and terminals were seen in the internal layer of median eminence and infundibular stem. Further caudally, scattered fibers were found in the central gray and the inferior colliculus. At post-natal day (P) 3, also some cell bodies were found in the caudal spinal nucleus of the trigeminal nerve (which is the place all the pain and temperature fibers from the face terminate) and the dorsal horn of the spinal cord. By the age of 4 weeks, the distribution of immunoreactivity was similar to adults, where cell bodies could be seen in the periventricular area of the medial hypothalamus in addition to more intense labeling of the previously mentioned areas (17, 51).
In an in situ hybridization study on developing rat, Nieminen et al. (18) found expression of NPFF at E14, much earlier than the presence of NPFF protein seen with immunohistochemistry [at E20; (17)]. E14 embryos showed NPFF expression in the medulla and spinal cord (18). Later (E17), distinct neurons expressed NPFF in the spinal cord, and at birth NPFF expression was seen in neurons in the rostral nucleus of the solitary tract in the medulla, similar to what was found with immunohistochemistry. In addition, reticular nucleus (corresponding to lateral medullary reticular nucleus in adult rats) was found to express NPFF at birth, parallel to findings in developing Xenopus (15). This expression was transient in the rat, indicating that NPFF may be involved in development of the sensory trajectories passing through this nucleus (18). Expression of NPFF was also seen in the pituitary of the embryonic rat, but no NPFF-expressing cells were observed here. Finally, NPFF expression was found in the developing lung and spleen.
Using an NPFF radioligand, Desprat et al. showed the presence of receptors for NPFF in regions of the developing mouse brain and spinal cord involved in the analgesic effects of opiates (28). During post-natal development, they found that NPFF affected the morphine-induced analgesia in different ways in the neonatal, but in adults NPFF had only anti-opioid effect. From birth, they could detect binding of NPFF in the olfactory bulb, and from P7 in the ventral pallidum and nucleus ventral endopiriform in the telencephalon. Also in the diencephalon in the nucleus reuniens NPFF binding appeared at P7, and in the mesencephalon a few binding sites was visible at birth. This study shows that the interplay between NPFF receptors and opioid receptors is established at early stages in mice. However, embryonic stages were not studied, so the pattern of NPFF binding in the early developing mouse is not known.
Interestingly, the TN of the mammal tree shrew (Tupaia belangeri) shows FMRFir (30). This mammal is closely related to primates. The FMRFir pattern was similar to the pattern in fish, amphibians, reptiles, and birds, with FMRFa cells appearing early in embryogenesis (E20) near the olfactory epithelium, and later along the migrating route for the TN. Thus, it seems that NPFF expression in the TN is a feature conserved from fishes to mammals.
In summary, NPFF is detected early in embryonic development in all vertebrates studied, see overview in Table 1. In fishes, it seems to be exclusively expressed in the TN, also during development. In amphibians, reptiles, birds, and mammals NPFF can in addition be found in the suprachiasmatic region in hypothalamus at very early developmental stages. In Xenopus and mammals, medullary reticular formation also shows NPFF labeling in embryos. No function has been demonstrated for NPFF during development, but its expression pattern in the brain may suggest it could be involved in development of neurons of the TN, and nerve circuits involved in control of feeding.
PrRP Group
The PrRP group includes the peptides PrRP31 and PrRP20. A new member of this family, C-RFa is found in Japanese crucian carp (Carassius cuvieri), zebrafish, Xenopus, and chicken (Gallus gallus), but this variant is not found in mammals (52). PrRP peptides bind the receptor GPR10 (prolactin-releasing hormone receptor; PRLHR; also named GR3), but they also bind NPFFR2. Three different receptors for PrRP peptides exist in some vertebrates, while only one is found in mammals [see receptor synteny in Wang et al. (52)].
PrRP in Adult Vertebrates
It is believed that PrRP is involved in the control of pituitary function in fishes. Firstly, in many species of adult fishes, PrRP fibers project to and terminate on prolactin-producing cells of the pituitary. Secondly, C-RFa injections in rainbow trout and tilapia cause release of prolactin and somatostatin (53–56). Furthermore, PrRP cell bodies are found in the nucleus lateralis tuberis pars posterioris in guppy (Poecilia reticulata), rainbow trout (Oncorhynchus mykiss), and goldfish (Carassius auratus) (52, 56, 57), an area suggested to be important for control of pituitary function in teleosts (58). There seems to be some variation in PrRP expression between species. Some fish species have PrRP cell bodies also in other brain areas, while others do not seem to have PrRP fibers projecting to the pituitary (57).
In mammals, PrRP was thought to act on the pituitary, because of the high expression of its receptor GPR10 in the anterior pituitary (59). Preliminary studies in rats showed that the peptide could promote prolactin release from pituitary cells and from these studies the peptide got its name (60). However, later studies have shown that this pathway may not be physiologically relevant in mammals. Instead, it has been shown that PrRP is involved in control of food intake and energy balance in rats and mice, and that it can affect the stress response by elevating circulating levels adrenocorticotropic hormone (ACTH) (59). Further, it has been shown that PrRP has an effect on the cardiovascular system and on circadian rhythms in mammals. Interestingly, it has been suggested that endogenously produced PrRP peptide has an autocrine role in cell-cycle progression and growth (61), processes that are closely linked to development. PrRP is expressed in the nucleus of the solitary tract, ventrolateral medulla, and in the caudal portion of the dorsomedial hypothalamic nucleus in adult mammals (59). PrRP fibers are found in many areas of the forebrain, as in preoptic area, periventricular nucleus of the thalamus, and in periventricular nucleus and paraventricular nucleus of hypothalamus (62). In contrast to the situation in fish and amphibians, no PrRPir can be detected in the median eminence or in the hypophysiotropic cells of the hypothalamus (63, 64). In peripheral tissues, PrRP mRNA is found in the adrenal gland, pancreas, placenta, and testis (59).
PrRP in Vertebrate Development
Very few studies have looked at the possible role of PrRP in development. However, the few that exist show that this peptide is expressed at an early stage in Xenopus (65), chicken (52), and rat (18, 62).
In the teleost guppy, PrRPir cells were detected in the nucleus lateralis tuberis pars posterioris in the hypothalamus already at the day of birth (57). However, innervation of PrRP fibers to the prolactin cells of the pituitary was not seen at birth day, but appeared later. Earlier developmental stages were not investigated, so the role of this peptide in teleost development is not clear.
In Xenopus, the expression of PrRP mRNA is detected at an early stage (stage 54; measured with qPCR) (65). PrRP mRNA levels were highest in early premetamorphic stages (stage 57), and decreased during prometamorphosis. This coincides with prolactin starting to appear in the pituitary. In the chicken pituitaries, the PrRP receptor PrRPR1 was expressed at E8 (measured with qPCR) (52), and the expression increased at later developmental stages (E12, E16, and E20). However, earlier stages were not studied.
In rat, PrRP mRNA and PrRPir cells are found in the nucleus of the solitary tract at E18, and in the ventral and lateral reticular nucleus of the caudal medulla oblongata at E20 (62). The hypothalamus first showed PrRP expression and PrRPir at P13. Similar to sexually mature rats, P6 animals had PrRPir fibers in paraventricular hypothalamic nucleus, periventricular hypothalamic nucleus, medial preoptic area, basolateral amygdaloid nucleus, dorsomedial hypothalamus, ventromedial hypothalamus, periventricular nucleus of the thalamus, and bed nucleus of the stria terminalis. However, also areas not showing PrRPir in the adult had PrRP in the developing rat brain at P6 and P9, like optic chiasm, dorsal endopiriform nucleus, cingulum, intermediate reticular nucleus, and caudal ventrolateral reticular nucleus. This transient expression could indicate a role in development of these brain areas.
The presence of mRNA of PrRP and its receptor GPR10 has also been investigated with in situ hybridization in rat embryos in a different study. Nieminen et al. (18) found a similar pattern as shown with immunohistochemistry. However, they found expression of PrRP in the reticular formation at a much earlier stage [E17 compared to P6 in (62)]. The receptor GPR10 was expressed at very early stages; at E15 in the pallium, at E16 in the hippocampus, and at E19 in the reticular formation. Interestingly, this is before any PrRP mRNA can be detected in these locations (18, 62), which may suggest that the receptor has an alternative ligand. In the pituitary, the expression of both PrRP and GPR10 starts at E18, at the same time as the lactotrops first appear (18). This is the opposite of the situation in Xenopus, where PrRP expression decreases when prolactin starts to appear in the pituitary (65). In the periphery, PrRP expression was seen in the developing liver, and to some extent in the spleen and kidney (18).
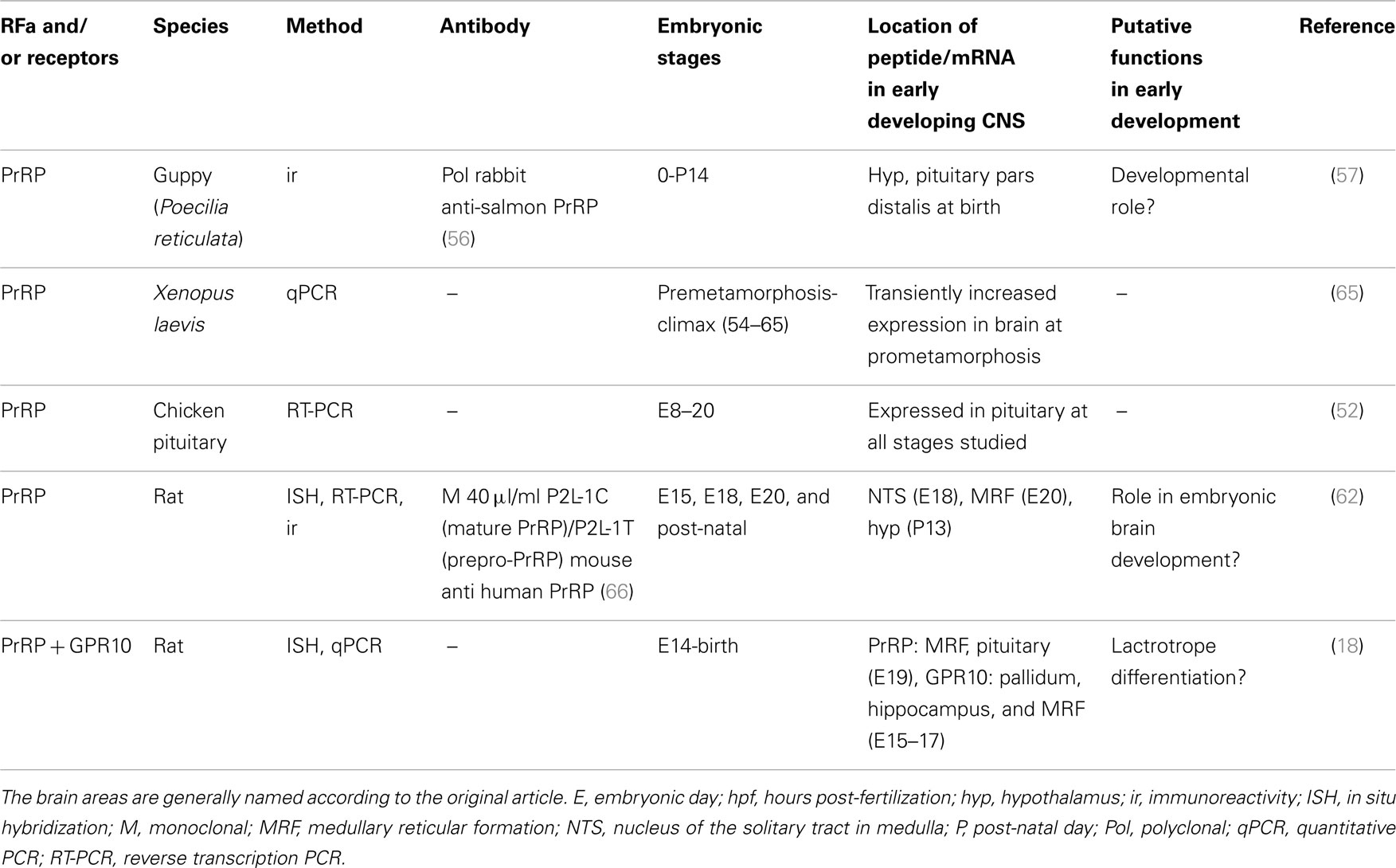
Studies of PrRP in vertebrate development are summarized in Table 2.
GnIH Group
GnIH was first described by Tsutsui et al. (67). This RFa was found to inhibit gonadotropin release in the quail through binding to the G-protein-coupled receptor GPR147. It has since been found that in birds, GnIH can act directly on gonadotrope cells to inhibit both synthesis and release of gonadotropins, e.g., Ref. (68–70). Moreover, it may also act on GnRH-neurons to inhibit GnRH release and thereby indirectly inhibit gonadotrope cells [e.g., Ref. (71)]. Since its first discovery in birds, GnIH orthologs have been found in most vertebrate classes [see review by Tsutsui and Ubuka (72) and references therein]. Alternative names are sometimes used in different vertebrate classes – the mammalian ortholog being named RFa-related peptide (RFRP) with the RFRP gene encoding two bioactive peptides, RFRP-1 (or NPSV) and RFRP-3 (or NPVF) (73). The mammalian receptor is sometimes referred to as NPFFR1 or NPFF1. In amphibians, GnIH is sometimes referred to as GRP, GRP-RPs, or R-RFa [e.g., Ref. (74)], while in teleosts the term LPXRFa may be used [e.g., Ref. (75)]. In the following, we will use the common name GnIH regardless of vertebrate class.
GnIH in Adult Vertebrates
In adult vertebrates, GnIH positive cells are found in different regions of the brain, notably in hypothalamic regions like the avian paraventricular nucleus, from where they send their projections to GnRH1 neurons in the preoptic region or to gonadotrope cells in the pituitary. GnIH terminals and GnIH receptors have also been identified on GnRH2 neurons in birds and mammals, e.g., Ref. (71, 76). A recent paper shows that GnIH inhibits socio-sexual behavior of male quail through a direct activation of aromatase and thereby increased neuroestrogen synthesis in the preoptic area (77). Both GnIH and its receptor are also expressed in the pituitary in different vertebrate classes, indicating auto- or paracrine regulation [e.g., Ref. (78)]. In addition, various studies have revealed GnIH positive cells in the gonads (both testis and ovary), while PCR experiments have revealed gene expression in peripheral tissues like muscle, spleen, eye, and kidney [e.g., Ref. (79)]. The expression of both ligand and receptor in the avian gonads (80) again points to an auto- or paracrine role during gametogenesis.
The spatial expression pattern should indicate potential functions, although much remains to be discovered when it comes to GnIH functions in general and during development in particular. Similar to in birds, GnIH in mammals have been shown to inhibit gonadotropin synthesis and release, either directly in the pituitary or via inhibition of hypothalamic GnRH-neurons. The situation seems different in frogs and teleost fish where GnIH can either inhibit or stimulate gonadotropin (and also growth hormone and prolactin) release, depending on reproductive stage, species, and sex, e.g., Ref. (74, 79, 81–84). Also in an agnathan species (sea lamprey; Petromyzon marinus), GnIH stimulates expression of GnRH and gonadotropin β-subunit (85), indicating that this neuropeptide may have experienced a shift in function during vertebrate evolution.
As most interest has focused on its role as an inhibitor of GnRH and gonadotropin release during reproduction, very little is known about GnIH during vertebrate development. Apart from some studies looking at pre-pubertal stages, the information we have is mostly limited to studies on the spatio-temporal expression pattern in mammalian and avian (post-natal) development, and some very few in teleost early development.
GnIH in Vertebrate Development
A recent article from Biswas and colleagues (78) provided some interesting information regarding the spatial expression pattern of GnIH peptides in Indian major carp (Labeo rohita), although detailed origin of their antibodies are missing from the paper. For instance, they found GnIH expression in the olfactory system (epithelium and bulb) in newly hatched larvae, indicating a non-reproductive function. Expression in hypothalamic nuclei such as the periventricular preoptic nucleus and the posterior periventricular nucleus, usually related to gonadotropin regulation in the adult, were also found in newly hatched larvae of the Indian major carp. Moreover, GnIH was found expressed in the anterior part of the carp pituitary [rostral pars distalis (RPD) and proximal pars distalis (PPD)] already from hatching, although no staining could be found at the adult stage, again pointing to an autocrine or paracrine function. Another recent study provides information of the temporal gene expression pattern of GnIH and its receptors during zebrafish early development (79). Whereas each of the three GnIH receptor paralogs could be found expressed (RT-PCR) already at the blastula stage and all the way through to the adult stage, although with differential expression patterns, the GnIH ligand was found only from a later embryonic stage; at 24 hpf (early pharyngula period). Nevertheless, these results indicate a functional GnIH ligand/receptor system active already from early embryonic stages in zebrafish. This is supported by recent results from our own lab, where we find expression of medaka (Oryzias latipes) gnih and gnihr1 already from 1 hpf and throughout the larval period, indicating maternal transfer in medaka (own unpublished data). The other two receptor paralogs in medaka, gnihr2 and gnihr3 were also expressed from early stages and throughout the larval period, although not until after the mid-blastula transition. The expression profile of medaka gnih resembled that seen in GnIH neurons in post-natal mice (86) with an initial increase followed by a steady decrease in expression levels. There is no existing data on the spatial expression of GnIH ligand or receptor during early development in fish.
In birds, where GnIH was first characterized, the existing literature focuses on the function of the GnIH system during sexual development, especially during the pre-pubertal period. For instance, circulating gonadotropin levels have been found to be negatively correlated with hypothalamic GnIH content (87). In immature male quail, chronic injections of GnIH suppressed normal testicular development, including reduced plasma testosterone levels and suppression of germ cell proliferation and seminiferous tubule development (70).
As mammalian model systems are less suited for studies of early embryogenesis, the few papers dealing with GnIH in mammalian development starts from late gestational stages. Yano et al. (88) found both GnIHs (RFRP-1 and -3) expressed in rat fetal hypothalamus from E15–E16 (mRNA) and embryonic day 16–17 (protein), showing first evidence of the existence of a functional system at the fetal stage also in mammals. Using a combination of GnIH in situ hybridization and BrdU immunohistochemistry, Legagneux et al. (89) identified GnIH producing neurons in male and female rat exclusively in the tuberal hypothalamus. These neurons to a large extent developed early, around E13–E14. In male and female rats, both Quenell et al. (90) and Iwasa et al. (73) found progressive gene expression of both GnIH and its receptor from early post-natal stages (from 4 days) all the way through puberty to the adult stage. Iwasa et al. also measured GnIH peptide levels and found a profile similar to that of GnIH gene expression, suggesting that the GnIH system indeed is active and plays a role during sexual development in rats (73). In accordance with these results, Poling et al. (86) also detected early gene expression of GnIH. mRNA levels increased in both sexes during post-natal and pre-pubertal development, before a decline was seen between post-natal day 20 and adulthood. During development in male rats, sustained knockdown of GnIH led to increased plasma levels of LH and increased testicular growth (91). These data indicate that GnIH act as inhibitor of gonadal maturation and puberty, similar to the situation in birds. However, Iwasa et al. (73) found increased GnRH gene expression levels concomitant with increased GnIH ligand and receptor expression during development in rat. This suggests that GnIH alone is not sufficient as inhibitor, but that regulation of sexual development is more complex, probably including additional factors such as other RFa, in addition to energy-related factors like ghrelin or leptin, or other.
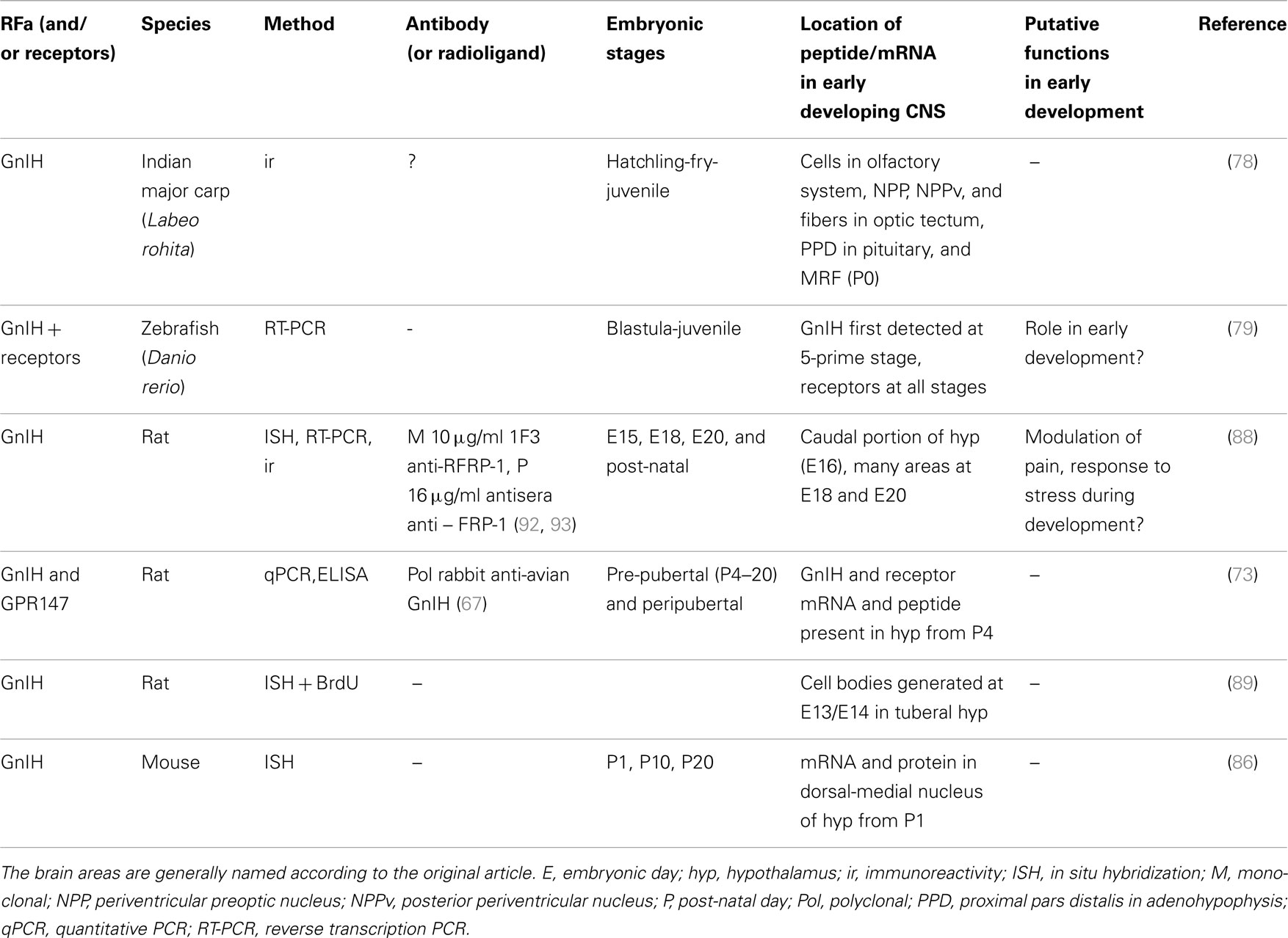
In summary, the expression of a seemingly functional GnIH system in fish, birds, and mammals already from early development suggests important developmental function(s) of this RFa in vertebrates. If these include more than the above mentioned regulatory (inhibitory/modulatory) effects on sexual development, remains to be seen. See Table 3 for an overview over developmental studies of GnIH.
Kisspeptin Group
Kisspeptins are RFa encoded by the Kiss gene. The resulting protein is further processed into bioactive peptides of variable lengths (94), while their receptors belong to the rhodopsin family of G-protein-coupled receptors (95–98).
Kisspeptin in Adult Vertebrates
The product of the Kiss gene was first discovered as a metastasis suppressor and therefore termed metastin (99). However, kisspeptins and their putative receptors (Kissr or Gpr54) have during the last decade emerged as major gatekeepers of reproduction because of their central role in regulating the brain–pituitary–gonadal (BPG) axis [reviewed by Ref. (100)]. The importance of the Kiss system as a regulator of the BPG-axis came after observations that mutations in the Gpr54-1 lead to idiopathic hypogonadotropic hypogonadism (101, 102). Besides its role as a tumor suppressor and regulator of the BPG-axis, several studies report additional roles of the Kiss system, including vasoconstriction (103, 104), neuronal migration, and increased synaptic transmission (105, 106). For example, Fiorini et al. (107) showed that stimulation with KISS increased neurite growth in GnRH-positive neurons in vitro. Although detailed mechanisms of action are still lacking, these seemingly pleiotropic roles may reflect the diversity of intracellular signaling pathways that can be triggered by Kiss receptor activation (100, 108). The current understanding in mammalian systems suggests that major endogenous and environmental signals act through Kiss neurons, which then directly or indirectly provide an integrated signal to the hypophysiotropic GnRH neurons.
Kisspeptins during Development
Despite the accumulating data of the role of kisspeptins in adult vertebrates, less is known about kisspeptins during post-natal/pre-pubertal development, and very little is known regarding the potential expression and function of the kiss system during embryogenesis/early development. This could at least partly be due to the lack of a suitable model system. Because Kiss or Gpr54 KO mice are infertile, homozygous offspring need to be established from heterozygous parents. This means that the possibility of maternal transfer of transcripts, including those of Kiss and Gpr54, cannot be excluded in this system. Furthermore, and common to all RFa families discussed here, studying embryonic development in mammals in vivo is difficult due to their viviparity.
The few existing data on kisspeptins during early development come from studies in medaka and zebrafish. We recently performed a study of kisspeptin ligand and receptor expression pattern and function during early development in medaka, exploiting the advantages of the teleost model system (109). qPCR gene expression profiles (Figure 1) revealed maternally provided Kiss systems involving the two kiss ligands (kiss1 and kiss2) and one of the receptors (gpr54-1), indicating the possibility of functional Kiss receptor-ligand systems at very early stages. gpr54-2, on the other hand, was not detected until after the zygotic phase, at stage 15, with a significant increase in expression levels between stage 30 and stage 36. In zebrafish, kiss1 and kiss2 gene expression was reported in 24 hpf (30 somite stage) embryos (110), but earlier stages were not investigated. In another teleost, the cobia (Rachycentron canadum) gpr54-1 expression was detected at 1 day post hatching (111). The early expression of kiss and gpr54 also coincides with the early expression of gonadotropins in fish (112), indicating a potentially functional BPG-axis already during early embryogenesis.
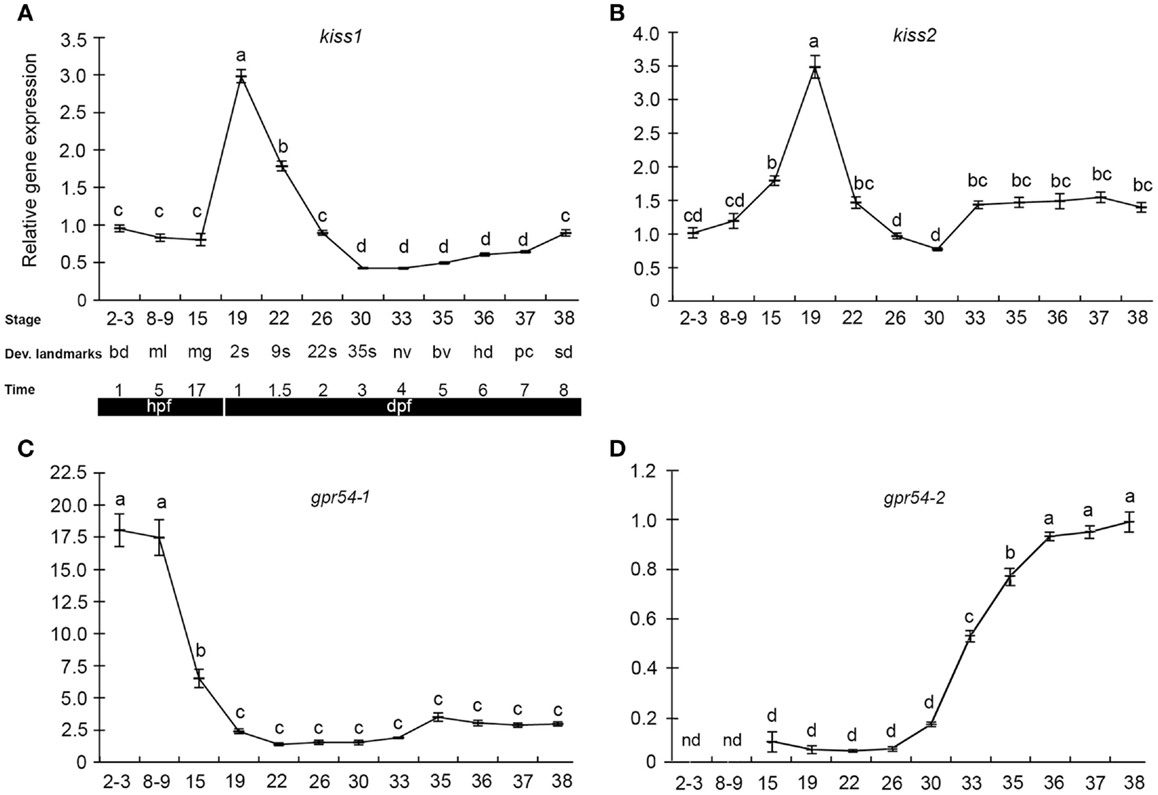
Figure 1. Kiss and kiss receptors are expressed at very early stages in medaka embryos. Relative gene expression of kiss1 (A), kiss2 (B), gpr54-1 (C), and gpr54-2 (D) was analyzed at different developmental stages (mean ± SEM; n = 7). Key developmental stages are given above the age of the hours (h) or days (d) post-fertilization (hpf). The gene expression levels are given relative to a reference gene (β-actin). Different letters indicate significant differences (P < 0.05). Figure from Ref. (109).
In Hodne et al., we performed a series of knockdown experiments that indicated several independent kiss systems during medaka embryonic development (109). Both maternally and zygotically expressed kiss1 and gpr54-1 seemed critical for proper development (Figure 2). However, the apparent functions of the maternally and zygotically expressed transcripts were quite distinct, as explained below.
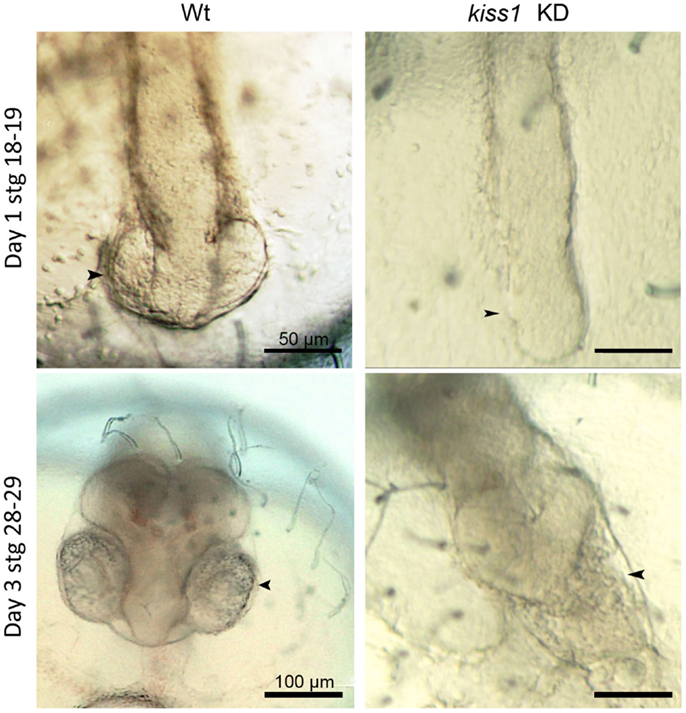
Figure 2. Low dose antisense knockdown (KD) lead to impaired brain and eye development. kiss1 knockdown embryos showed impaired eye development at 1 dpf (top right). At 3 dpf, there was no development of the forebrain after kiss1 knockdown (bottom right). Scale bars correspond to 50 μm (top images) or 100 μm (bottom images). Arrowheads point to the area of the developing eyes. Figure from Ref. (109).
Knockdown of maternal kiss1 and gpr54-1 led to developmental arrest and subsequent death around the blastula stage (stage 10–11), suggesting that this early expressed system could be involved either in regulation of early asynchronous cell division or early cell migration. The downstream factors controlled by kiss1/gpr54-1 signaling are not known. However, cell migration during blastulation and gastrulation is dependent on sdf1/cxcr4 chemotaxis. This signaling pathway is also known to be important during bone-directed migration of GPR54-positive breast cancer cells (113) and kisspeptin can indirectly regulate sdf1/cxcr4 through desensitization of cxcr4 by preventing rise in intracellular Ca2+ levels after sdf1 stimulation (114, 115). In zebrafish, knockdown of sdf1/cxcr4 inhibits migration of endodermal cells during gastrulation (115). Moreover, sdf1 signaling is crucial for survival in mice, and individuals lacking either receptor or ligand have defective hematopoiesis, developmental lymphoid tissue, vascularization of gastrointestinal tract, migration of neuronal cells, and patterning in the central nervous system, and they die prenatally (116, 117).
Zygotic knockdown of kiss1 and gpr54-1, on the other hand, allowed the embryos to survive gastrulation and a seemingly normal development continued until completion of neurulation (stage 18). At this point, early eye development is normally observed. However, after zygotic knockdown with either morpholino or low dose of peptide nucleic acid (PNA), eye development was interrupted, and further brain development was severely disrupted (Figure 2). Surprisingly, knockdown of kiss2, which was expressed at similar levels as kiss1, did not produce any increased mortality or malformed embryos. Unless kiss1 takes over the role of kiss2 following kiss2 knockdown, kiss2 does not seem to be critical for proper development. If kiss2 is translated and active before zygotic activation, our results indicate that it does not work through gpr54-1 (which is not expressed at this point), but possibly through gpr54-2 (although knockdown of this receptor leads to developmental arrest and 100% mortality), or through other, currently unknown, RFa receptors. The different effects observed following kiss1 and kiss2 knockdown suggest the possibility of a very early separation of two functional systems during embryonic development. One system, comprised of Kiss1 and Gpr54-1, has a functional role important for survival during the maternal stage of development. This system continues to function throughout embryonic development, although it seems more important for regulating brain development at later embryonic stages. A second system seems to be comprised of Kiss2 binding to either Gpr54-2 or other unknown RFa receptors. The possible function of this second system remains to be clarified.
Contrary to the observed phenotypes following zygotic knockdown of kiss1 and gpr54-1 outlined above, a moderate knockdown of medaka gpr54-2 arrested further development at stage 16 (late gastrula). This phenotype resembled that of maternal kiss1 and gpr54-1 knockdowns. However, as gpr54-2 is first detected after transition to zygotic gene expression, new questions arise as to why a similar phenotype was not also observed after zygotic kiss1 and gpr54-1 knockdown. One explanation could be that the two receptors are functionally separated. If they are involved in similar functions, our results indicate that the actions of Gpr54-1 may be partly compensated for by Gpr54-2, whereas Gpr54-2 cannot be functionally replaced by Gpr54-1.
A recent work by Zhao et al. has investigated the role of kiss on GnRH neuron development in zebrafish (118). In line with Kitahashi et al. (110), kiss1 and kiss2 mRNA could be detected by qPCR from 24 hpf. Furthermore, it was shown that both kiss ligands stimulated GnRH3 neuron proliferation peripherally, while only kiss1 stimulated proliferation and synaptic contact points of GnRH3 neurons in the TN and hypothalamic regions.
The existence of a functional kisspeptin system in birds is not clarified [see in Ref. (119)], and consequently, there are no data on this during bird development.
In mammals, the kisspeptin system has been intensively investigated during the last decade. Whereas, most literature covers the key role of Kiss in regulating GnRH neuron around and after puberty, pre- and early neonatal stages have been looked into more closely during recent years [see reviews in Ref. (120, 121) and references therein]. For instance, Kiss1 gene expression has been detected in hypothalamic areas during the late fetal period in mice [stage E13.5; (122)] and both gene expression and peptide have been detected in rats [stage E14.5, (123)]. There are two hypothalamic areas expressing Kiss; the arcuate nucleus (ARC), and the preoptic AVPV (anteroventral periventricular nucleus) and PeN (rostral periventricular nucleus). Whereas, Kiss neurons appear in the ARC prenatally, Kiss expression in the AVPV/PeN is not seen until neonatal stages. There seems to be clear sex differences in the expression of Kiss in rodents, with females expressing higher levels than males in both the ARC and the AVPV/PeN. Gpr54-1 gene expression has also been detected in stage E13.5 prenatal mice (122), indicating the possibility of a functional ligand–receptor system from this stage on. Also in second trimester human fetuses, KISS and GPR54 immunoreactivity was detected in the hypothalamus. Maternally provided Kiss ligands or receptors have, however, not been reported.
It seems that the early kisspeptin systems are functional in rodents in that Kiss neurons already are in close contact with GnRH-neurons prenatally, and that GnRH-neurons are able to respond to kisspeptins by enhanced GnRH secretion during prenatal life (120, 122, 124, 125). Based on these and several other studies, a more generalized picture is starting to appear with a seemingly functional kisspeptin system in place during the last part of gestation. The Kiss neurons increase in number and activity and reach a peak prenatally, before a decreased activity around birth, and then a new increase again during early neonatal life before the activity decreases to low levels until the pre-pubertal stage. The prenatal and early neonatal peaks in Kiss neuron activity seemingly coincide with similar peaks in GnRH and pituitary gonadotropin secretion [see in Ref. (120, 121, 126–132)]. However, the function of this early expression of Kiss remains elusive.
In line with the more severe phenotypes observed in medaka following receptor knockdown (109), Lapatto et al. (133) also described a more severe phenotype following Gpr54-1 knockout- compared with Kiss1 knockout mice. As mice possess only the one Gpr54 paralog, one of several suggestions was a possible weak activation of Gpr54-1 by other ligands. The results of Hodne et al. (109) and those of Lapatto et al. (133) suggest that Kiss and possibly other RFa may promiscuously bind to different RFa receptors [see also in Ref. (134–136)]. Interestingly, Mayer and Boehm (137) found that female mice with genetically ablated kisspeptin neurons underwent puberty and became fertile. In contrast, acute ablation in adult mice inhibited fertility. These results clearly indicate compensatory mechanisms for early loss of kisspeptins. Whether maternally transferred kiss is crucial for mouse development has not been investigated.
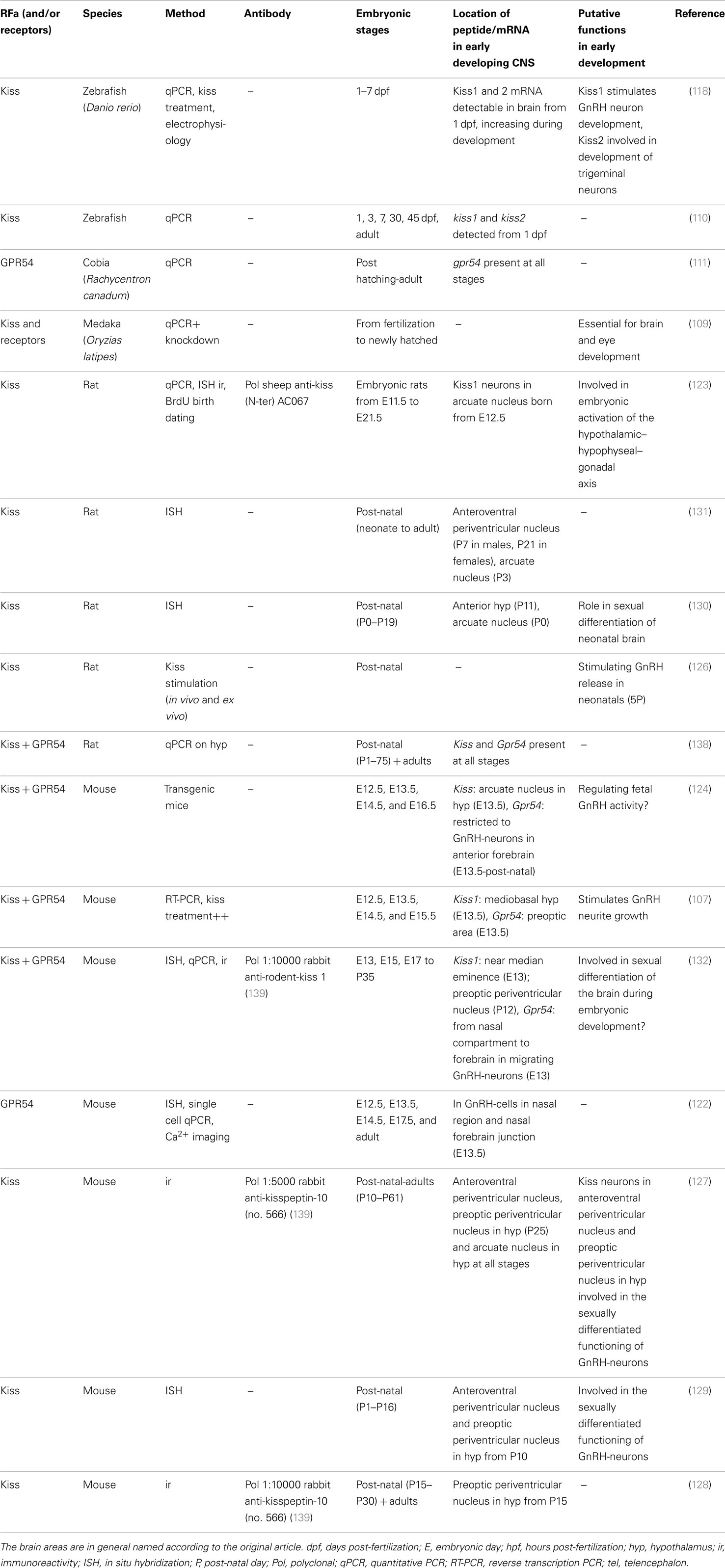
Although more data are available regarding the role of kisspeptins during vertebrate development compared to the role of other RFa, there are still much work to be done. One important aspect probably will be to elucidate their role in neuronal migration/development, where they seemingly play a major role, at least in fish. See Table 4 for an overview over developmental studies of kisspeptins.
26RFa/QRFP Group
The 26RF/QRFP group is the newest member of the RFa family, first described in 2003 in the brain of European green frog (140). The gene for 26RFa/QRFP is found in genomes of many species, from teleost fish to human, with preserved synteny in human, mouse, chicken, and Xenopus (4). Mature peptides generated from this gene are 26RFa, 43RFa in rat, mouse, human, and frog, 9RFa in human and frog, and 26RFa and 7RFa in fish (4, 141). 26RFa/QRFP binds the receptor GPR103/26RFaR (142, 143). In addition, the peptide has affinity for NPFFR2 (144).
26RFa/QRFP in Adult Vertebrates
In adult goldfish, 26RFa/qrfp mRNA is found in the hypothalamus, optic tectum-thalamus, and testis (141). Because the expression in the hypothalamus was significantly reduced after 4 days of starvation, and intraperitoneal injections of the 26RFa/QRFP peptide increased LH levels, 26RFa/QRFP has been suggested to play a role in the regulation of energy homeostasis and regulation of the BPG-axis in fish (141). Also in birds, 26RFa/QRFP is expressed in the diencephalon; in the anterior hypothalamic nucleus in chicken, and in the anterior-medial area, the ventromedial nucleus and the lateral hypothalamic area in zebra finch (Taeniopygia guttata), areas involved in hypothalamic control over feeding behavior (145, 146). The findings are similar in mammals, where the presence of 26RFa/QRFP has been shown in regions that are important for regulation of food intake and energy homeostasis. In rodents, high levels of 26RFa/QRFP are found in dorsolateral and mediobasal hypothalamic areas, and in humans 26RFa/QRFP-cells are found in the paraventricular and ventromedial nuclei of the hypothalamus (140, 144, 147). It has been found that injections of 26RFa/QRFP have an orexigenic effect (increased food intake) in rodents (148). 26RFa/QRFP has also been found to affect aldosterone secretion, insulin secretion, adipogenesis, bone formation, nociceptive transmission, blood pressure and, as in fish, to stimulate pituitary hormone secretion in mammals (4).
26RFa/QRFP in Development
The developmental expression of 26RFa/QRFP and GPR103 has only been studied in human adrenal gland and rat adrenals (149). Both the receptor and ligand were present from early stages of adrenal development, mostly expressed in the adrenal cortex, but also in the medulla in human fetus. Some clues for the potential role of 26RFa/QRFP in development are also obtained from knockout studies of the receptor for 26RFa/QRFP in mice (150). Homozygous mice was viable, but they suffered from osteopenia (reduced bone density), and the investigations suggested that the bone formation was arrested at an early stage. It remains unclear if this effect is due to lack of hypothalamic signaling of 26RFa/QRFP, because mRNA of the receptor is found in bone and in osteoblast cell lines in addition to the hypothalamic expression (150). The knockout mice seemed to behave normally and were fertile. However, studies of the brain and the behavior of the animals was not included in the paper, thus it is unknown if knockout of the 26RFa/QRFP receptor can affect specific behaviors.
Conclusion
The RFa form a complex family with many different members acting in various physiological processes, with one peptide seemingly having several functions in the same animal in some cases. Common to all the peptides seems to be that they could have a role in appetite regulation, pain modulation, and reproduction. With the newest member of the RFa family found only a little over 10 years ago, the field of RFa is relatively new and requires much more research. Especially, how these peptides can influence development is poorly understood. However, the few studies of RFa in developing vertebrates show interesting results that may indicate that many of the RFa could have a separate role in development. Interestingly, it seems that all RFa are expressed early in development in many different groups of vertebrates. However, most of our knowledge of RFa comes from in situ hybridization and immunohistochemistry experiments. Very few functional studies have been conducted, so it is difficult to assess the role of this early expression. One knockdown study on medaka performed in our laboratory shows that kiss1, gpr54-1, and gpr54-2 play vital roles in early development in this fish species, and that these genes are probably important for proper brain development. It will be very interesting to see if loss or gain of function could reveal novel functions of the other RFa.
The cellular pathways of RFa are poorly understood, and more research is required to find out how RFa can act on developmental processes. However, some RFa (26RFa and Kiss) have been shown to affect migration in cancer cells (114, 151). RFa may also be important for the proper migration of neurons in the developing brain. However, more research is needed to clarify the role of RFa in neuronal migration. Another interesting aspect of RFa is the fact that many of them can affect apoptosis and cell-cycle progression, possibly through affecting opioid receptors (152). PrRP is also found to influence human lymphocyte proliferation (153). In invertebrates, FMRFa has been found to inhibit apoptosis in a snail, indicating that the link between RFa and apoptosis is an evolutionary conserved mechanism (154). Interestingly, NPFF gene expression in mammals is regulated by transcription factors also involved in cell-cycle regulation and apoptosis (155).
The field of RFa in vertebrates is exiting and rapidly expanding. The few developmental studies that have been done show promising and important results. Taken together, these studies indicate that RFa may have a role in development of the nervous system not yet identified. More research is needed, especially functional studies that can give insight into the role these peptides play in development.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the Research Council of Norway, Grants Nos. 184851 (to Finn-Arne Weltzien) and 231767/F20 (to Guro Katrine Sandvik).
References
1. Price DA, Greenberg MJ. Structure of a molluscan cardioexcitatory neuropeptide. Science (1977) 197:670–1. doi: 10.1126/science.877582
2. Yang HYT, Fratta W, Majane EA, Costa E. Isolation, sequencing, synthesis, and pharmacological characterization of two brain neuropeptides that modulate the action of morphine. Proc Natl Acad Sci U S A (1985) 82:7757–61. doi:10.1073/pnas.82.22.7757
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
3. Chartrel N, Dujardin C, Leprince J, Desrues L, Tonon MC, Cellier E, et al. Isolation, characterization, and distribution of a novel neuropeptide, Rana RFamide (R-RFa), in the brain of the European green frog Rana esculenta. J Comp Neurol (2002) 448:111–27. doi:10.1002/cne.10253
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
4. Ukena K, Osugi T, Leprince J, Vaudry H, Tsutsui K. Molecular evolution of GPCRs: 26Rfa/GPR103. J Mol Endocrinol (2014) 52:T119–31. doi:10.1530/JME-13-0207
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
5. Findeisen M, Rathmann D, Beck-Sickinger AG. RFamide peptides: structure, function, mechanisms and pharmaceutical potential. Pharmaceuticals (2011) 4:1248–80. doi:10.3390/ph4091248
6. Parhar I, Ogawa S, Kitahashi T. RFamide peptides as mediators in environmental control of GnRH neurons. Prog Neurobiol (2012) 98:176–96. doi:10.1016/j.pneurobio.2012.05.011
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
7. Elhabazi K, Humbert JP, Bertin I, Schmitt M, Bihel F, Bourguignon JJ, et al. Endogenous mammalian RF-amide peptides, including PrRP, kisspeptin and 26RFa, modulate nociception and morphine analgesia via NPFF receptors. Neuropharmacology (2013) 75:164–71. doi:10.1016/j.neuropharm.2013.07.012
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
8. Bechtold DA, Luckman SM. The role of RFamide peptides in feeding. J Endocrinol (2007) 192:3–15. doi:10.1677/JOE-06-0069
9. Croll RP. Development of embryonic and larval cells containing serotonin, catecholamines, and FMRFamide-related peptides in the gastropod mollusc Phestilla sibogae. Biol Bull (2006) 211:232–47. doi:10.2307/4134546
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
10. Aroua S, Andouche A, Martin M, Baratte S, Bonnaud L. FaRP cell distribution in the developing CNS suggests the involvement of FaRPs in all parts of the chromatophore control pathway in Sepia officinalis (Cephalopoda). Zoology (2011) 114:113–22. doi:10.1016/j.zool.2010.11.002
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
11. Nakanishi N, Yuan D, Jacobs DK, Hartenstein V. Early development, pattern, and reorganization of the planula nervous system in Aurelia (Cnidaria, Scyphozoa). Dev Genes Evol (2008) 218:511–24. doi:10.1007/s00427-008-0239-7
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
12. Franchini A. Distribution of cells containing FMRFamide-related molecules in the embryonic development of Ophryotrocha labronica (Polychaeta: Dorvilleidae). Invertebr Biol (2008) 127:327–35. doi:10.1111/j.1744-7410.2008.00133.x
13. Kreshchenko ND. Functions of flatworm neuropeptides NPF, GYIRF and FMRF in course of pharyngeal regeneration of anterior body fragments of planarian, Girardia tigrina. Acta Biol Hung (2008) 59:199–207. doi:10.1556/ABiol.59.2008.Suppl.29
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
14. Oehlmann VD, Korte H, Sterner C, Korsching SI. A neuropeptide FF-related gene is expressed selectively in neurons of the terminal nerve in Danio rerio. Mech Dev (2002) 117:357–61. doi:10.1016/S0925-4773(02)00210-1
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
15. Lopez JM, Moreno N, Morona R, Munoz M, Gonzalez A. Spatiotemporal sequence of appearance of NPFF-immunoreactive structures in the developing central nervous system of Xenopus laevis. Peptides (2006) 27:1036–53. doi:10.1016/j.peptides.2005.07.028
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
16. Yamamoto N, Uchiyama H, OhkiHamazaki H, Tanaka H, Ito H. Migration of GnRH-immunoreactive neurons from the olfactory placode to the brain: a study using avian embryonic chimeras. Dev Brain Res (1996) 95:234–44. doi:10.1016/0165-3806(96)00078-8
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
17. Kivipelto L, Rubenstein J, Yang HYT, Panula P. Ontogeny of the F8Famide-like (morphine-modulating) peptides in the central nervous system of rats. J Comp Neurol (1991) 304:14–33. doi:10.1002/cne.903040103
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
18. Nieminen ML, Nystedt J, Panula P. Expression of neuropeptide FF, prolactin-releasing peptide, and the receptor UHR1/GPR10 genes during embryogenesis in the rat. Dev Dyn (2003) 226:561–9. doi:10.1002/dvdy.10261
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
19. Pinelli C, D’Aniello B, Sordino P, Meyer DL, Fiorentino M, Rastogi RK. Comparative immunocytochemical study of FMRFamide neuronal system in the brain of Danio rerio and Acipenser ruthenus during development. Dev Brain Res (2000) 119:195–208. doi:10.1016/S0165-3806(99)00172-8
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
20. Castro A, Becerra M, Anadon R, Manso MJ. Distribution and development of FMRFamide-like immunoreactive neuronal systems in the brain of the brown trout, Salmo trutta fario. J Comp Neurol (2001) 440:43–64. doi:10.1002/cne.1369
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
21. Fiorentino M, D’Aniello B, Joss J, Polese G, Rastogi RK. Ontogenetic organization of the FMRFamide immunoreactivity in the nervus terminalis of the lungfish, Neoceratodus forsteri. J Comp Neurol (2002) 450:115–21. doi:10.1002/cne.10309
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
22. D’Aniello B, Fiorentino M, Pinelli C, di Meglio M, Vallarino M, Rastogi RK. Distribution of FMRFamide-like immunoreactivity in the brain and pituitary of Rana esculenta during development. Brain Res Dev Brain Res (1996) 95:194–204. doi:10.1016/S0165-3806(96)00088-0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
23. Fiorentino M, Pinelli C, D’Aniello B, Iela L, di Meglio M, Rastogi RK. Development and distribution of FMRFamide-like immunoreactivity in the toad (Bufo bufo) brain. J Chem Neuroanat (2001) 21:201–13. doi:10.1016/S0891-0618(01)00110-7
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
24. D’Aniello B, Fiorentino M, Pinelli C, Guarino FM, Angelini F, Rastogi RK. Localization of FMRFamide-like immunoreactivity in the brain of the viviparous skink (Chalcides chalcides). Brain Behav Evol (2001) 57:18–32. doi:10.1159/000047223
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
25. Wirsig-Wiechmann CR. The nervus terminalis in the chick: a FMRFamide-immunoreactive and AChE-positive nerve. Brain Res (1990) 523:175–9. doi:10.1016/0006-8993(90)91655-Z
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
26. O’Donohue TL, Bishop JF, Chronwall BM, Groome J, Watson WH III. Characterization and distribution of FMRFamide immunoreactivity in the rat central nervous system. Peptides (1984) 5:563–8. doi:10.1016/0196-9781(84)90087-1
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
27. Pinelli C, D’Aniello B, Polese G, Rastogi RK. Extrabulbar olfactory system and nervus terminalis FMRFamide immunoreactive components in Xenopus laevis ontogenesis. J Chem Neuroanat (2004) 28:37–46. doi:10.1016/j.jchemneu.2004.06.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
28. Desprat C, Zajac JM. Ontogeny of neuropeptide FF pharmacology and receptors in mouse brain. Dev Brain Res (1994) 82:118–26. doi:10.1016/0165-3806(94)90154-6
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
29. Malz CR, Kuhn H-J. FMRFamide immunoreactivity and the invasion of adenohypophyseal cells into the neural lobe in the developing pituitary of the tree shrew Tupaia belangeri. Brain Res (1999) 834:83–8. doi:10.1016/S0006-8993(99)01555-3
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
30. Malz CR, Kuhn H-J. Calretinin and FMRFamide immunoreactivity in the nervus terminalis of prenatal tree shrews (Tupaia belangeri). Dev Brain Res (2002) 135:39–44. doi:10.1016/S0165-3806(02)00299-7
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
31. Osugi T, Uchida K, Nozaki M, Tsutsui K. Characterization of novel RFamide peptides in the central nervous system of the brown hagfish: isolation, localization, and functional analysis. Endocrinology (2011) 152:4252–64. doi:10.1210/en.2011-1375
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
32. Vilensky JA. The neglected cranial nerve: nervus terminalis (cranial nerve N). Clin Anat (2014) 27:46–53. doi:10.1002/ca.22130
33. Okubo K, Nagahama Y. Structural and functional evolution of gonadotropin-releasing hormone in vertebrates. Acta Physiol (2008) 193:3–15. doi:10.1111/j.1748-1716.2008.01832.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
34. Abe H, Oka Y. Mechanisms of neuromodulation by a nonhypophysiotropic GnRH system controlling motivation of reproductive behavior in the teleost brain. J Reprod Dev (2011) 57:665–74. doi:10.1262/jrd.11-055E
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
35. Okuyama T, Yokoi S, Abe H, Isoe Y, Suehiro Y, Imada H, et al. A neural mechanism underlying mating preferences for familiar individuals in medaka fish. Science (2014) 343:91–4. doi:10.1126/science.1244724
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
36. Saito TH, Nakane R, Akazome Y, Abe H, Oka Y. Electrophysiological analysis of the inhibitory effects of FMRFamide-like peptides on the pacemaker activity of gonadotropin-releasing hormone neurons. J Neurophysiol (2010) 104:3518–29. doi:10.1152/jn.01027.2009
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
37. Crespo M, Moreno N, Lopez JM, Gonzalez A. Comparative analysis of neuropeptide FF-like immunoreactivity in the brain of anuran (Rana perezi, Xenopus laevis) and urodele (Pleurodeles waltl) amphibians. J Chem Neuroanat (2003) 25:53–71. doi:10.1016/S0891-0618(02)00097-2
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
38. Smeets W, Lopez JM, Gonzalez A. Distribution of neuropeptide FF-like immunoreactivity in the brain of the lizard Gekko gecko and its relation to catecholaminergic structures. J Comp Neurol (2006) 498:31–45. doi:10.1002/cne.21035
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
39. Rastogi RK, D’Aniello B, Pinelli C, Fiorentino M, Di Fiore MM, Di Meglio M, et al. FMRFamide in the amphibian brain: a comprehensive survey. Microsc Res Tech (2001) 54:158–72. doi:10.1002/jemt.1130
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
40. Lopez JM, Moreno N, Morona R, Gonzalez A. Distribution of neuropeptide FF-Like immunoreactivity in the brain of Dermophis mexicanus (Amphibia; Gymnophiona): comparison with FMRFamide immunoreactivity. Brain Behav Evol (2006) 67:150–64. doi:10.1159/000090979
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
41. Vilim FS, Aarnisalo AA, Nieminen ML, Lintunen M, Karlstedt K, Kontinen VK, et al. Gene for pain modulatory neuropeptide NPFF: induction in spinal cord by noxious stimuli. Mol Pharmacol (1999) 55:804–11.
42. Panula P, Aarnisalo AA, Wasowicz K. Neuropeptide FF, a mammalian neuropeptide with multiple functions. Prog Neurobiol (1996) 48:461–87. doi:10.1016/0301-0082(96)00001-9
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
43. Majane EA, Yang HYT. Distribution and characterization of two putative endogenous opioid antagonist peptides in bovine brain. Peptides (1987) 8:657–62. doi:10.1016/0196-9781(87)90041-6
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
44. Dockray GJ. The expanding family of -RFamide peptides and their effects on feeding behaviour. Exp Physiol (2004) 89:229–35. doi:10.1113/expphysiol.2004.027169
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
45. Whitlock KE. Development of the nervus terminalis: origin and migration. Microsc Res Tech (2004) 65:2–12. doi:10.1002/jemt.20094
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
46. Whitlock KE, Wolf CD, Boyce ML. Gonadotropin-releasing hormone (GnRH) cells arise from cranial neural crest and adenohypophy seal regions of the neural plate in the zebrafish, Danio rerio. Dev Biol (2003) 257:140–52. doi:10.1016/S0012-1606(03)00039-3
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
47. Tuinhof R, Artero C, Fasolo A, Franzoni MF, Donkelaar HJT, Wismans PGP, et al. Involvement of the retinohypothalamic input, suprachiasmatic nucleus, magnocellular nucleus and locus-coeruleus in control of melanotrope cells of Xenopus laevis – a retrograde and anterograde tracing study. Neuroscience (1994) 61:411–20. doi:10.1016/0306-4522(94)90241-0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
48. Northcutt RG, Muske LE. Multiple embryonic origins of gonadotropin-releasing hormone (GnRH) immunoreactive neurons. Dev Brain Res (1994) 78:279–90. doi:10.1016/0165-3806(94)90037-X
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
49. Nicklous DM, Simansky KJ. Neuropeptide FF exerts pro- and anti-opioid actions in the parabrachial nucleus to modulate food intake. Am J Physiol Regul Integr Comp Physiol (2003) 285:R1046–54. doi:10.1152/ajpregu.00107.2003
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
50. Hyde TM, Miselis RR. Effects of area postrema caudal medial nucleus of solitary tract lesions on food-intake and body weight. Am J Physiol (1983) 244:R577–87.
51. Kivipelto L, Majane EA, Yang HYT, Panula P. Immunohistochemical distribution and partial characterization of FLFQPQRFamide-like peptides in the central nervous system of rats. J Comp Neurol (1989) 286:269–87. doi:10.1002/cne.902860211
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
52. Wang Y, Wang CY, Wu Y, Huang G, Li J, Leung FC. Identification of the receptors for prolactin-releasing peptide (PrRP) and Carassius RFamide peptide (C-RFa) in chickens. Endocrinology (2012) 153:1861–74. doi:10.1210/en.2011-1719
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
53. Wang XY, Morishita F, Matsushima O, Fujimoto M. Immunohistochemical localization of C-RFamide, a FMRF-related peptide, in the brain of the goldfish, Carassius auratus. Zoolog Sci (2000) 17:1067–74. doi:10.2108/zsj.17.1067
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
54. Seale AP, Itoh T, Moriyama S, Takahashi A, Kawauchi H, Sakamoto T, et al. Isolation and characterization of a homologue of mammalian prolactin-releasing peptide from the tilapia brain and its effect on prolactin release from the tilapia pituitary. Gen Comp Endocrinol (2002) 125:328–39. doi:10.1006/gcen.2001.7727
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
55. Montefusco-Siegmund RA, Romero A, Kausel G, Muller M, Fujimoto M, Figueroa J. Cloning of the prepro C-RFa gene and brain localization of the active peptide in Salmo salar. Cell Tissue Res (2006) 325:277–85. doi:10.1007/s00441-006-0168-6
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
56. Moriyama S, Ito T, Takahashi A, Amano M, Sower SA, Hirano T, et al. A homolog of mammalian PRL-releasing peptide (fish arginyl-phenylalanyl-amide peptide) is a major hypothalamic peptide of PRL release in teleost fish. Endocrinology (2002) 143:2071–9. doi:10.1210/endo.143.6.8744
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
57. Amano M, Oka Y, Amiya N, Yamamori K. Immunohistochemical localization and ontogenic development of prolactin-releasing peptide in the brain of the ovoviviparous fish species Poecilia reticulata (guppy). Neurosci Lett (2007) 413:206–9. doi:10.1016/j.neulet.2006.10.011
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
58. Ball JN. Hypothalamic control of the pars distalis in fishes, amphibians, and reptiles. Gen Comp Endocrinol (1981) 44:135–70. doi:10.1016/0016-6480(81)90243-4
59. Dodd GT, Luckman SM. Physiological roles of GPR10 and PrRP signaling. Front Endocrinol (2013) 4:20. doi:10.3389/fendo.2013.00020
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
60. Hinuma S, Habata Y, Fujii R, Kawamata Y, Hosoya N, Fukusumi S, et al. A prolactin-releasing peptide in the brain. Nature (1998) 393:272–6. doi:10.1038/30515
61. Samson WK, Taylor MM. Prolactin releasing peptide (PrRP): an endogenous regulator of cell growth. Peptides (2006) 27:1099–103. doi:10.1016/j.peptides.2006.01.008
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
62. Yano T, Iijima N, Kataoka Y, Hinuma S, Tanaka M, Ibata Y. Developmental expression of prolactin releasing peptide in the rat brain: localization of messenger ribonucleic acid and immunoreactive neurons. Dev Brain Res (2001) 128:101–11. doi:10.1016/S0165-3806(01)00148-1
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
63. Maruyama M, Matsumoto H, Fujiwara K, Noguchi J, Kitada C, Fujino M, et al. Prolactin-releasing peptide as a novel stress mediator in the central nervous system. Endocrinology (2001) 142:2032–8. doi:10.1210/en.142.5.2032
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
64. Matsumoto H, Murakami Y, Horikoshi Y, Noguchi J, Habata Y, Kitada C, et al. Distribution and characterization of immunoreactive prolactin-releasing peptide (PrRP) in rat tissue and plasma. Biochem Biophys Res Commun (1999) 257:264–8. doi:10.1006/bbrc.1999.0463
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
65. Sakamoto T, Oda A, Yamamoto K, Kaneko M, Kikuyama S, Nishikawa A, et al. Molecular cloning and functional characterization of a prolactin-releasing peptide homol from Xenopus laevis. Peptides (2006) 27:3347–51. doi:10.1016/j.peptides.2006.08.003
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
66. Maruyama M, Matsumoto H, Fujiwara K, Kitada C, Hinuma S, Onda H, et al. Immunocytochemical localization of prolactin-releasing peptide in the rat brain. Endocrinology (1999) 140:2326–33. doi:10.1210/endo.140.5.6685
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
67. Tsutsui K, Saigoh E, Ukena K, Teranishi H, Fujisawa Y, Kikuchi M, et al. A novel avian hypothalamic peptide inhibiting gonadotropin release. Biochem Biophys Res Commun (2000) 275:661–7. doi:10.1006/bbrc.2000.3350
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
68. Bentley GE, Perfito N, Ukena K, Tsutsui K, Wingfield JC. Gonadotropin-inhibitory peptide in song sparrows (Melospiza melodia) in different reproductive conditions, and in house sparrows (Passer domesticus) relative to chicken-gonadotropin-releasing hormone. J Neuroendocrinol (2003) 15:794–802. doi:10.1046/j.1365-2826.2003.01062.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
69. Ciccone NA, Dunn IC, Boswell T, Tsutsui K, Ubuka T, Ukena K, et al. Gonadotrophin inhibitory hormone depresses gonadotrophin alpha and follicle-stimulating hormone beta subunit expression in the pituitary of the domestic chicken. J Neuroendocrinol (2004) 16:999–1006. doi:10.1111/j.1365-2826.2005.01260.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
70. Ubuka T, Ukena K, Sharp PJ, Bentley GE, Tsutsui K. Gonadotropin-inhibitory hormone inhibits gonadal development and maintenance by decreasing gonadotropin synthesis and release in male quail. Endocrinology (2006) 147:1187–94. doi:10.1210/en.2005-1178
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
71. Ubuka T, Kim S, Huang Y-C, Reid J, Jiang J, Osugi T, et al. Gonadotropin-inhibitory hormone neurons interact directly with gonadotropin-releasing hormone-I and -II neurons in European starling brain. Endocrinology (2008) 149:268–78. doi:10.1210/en.2007-0983
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
72. Tsutsui K, Ubuka T. Breakthrough in neuroendocrinology by discovering novel neuropeptides and neurosteroids: 1. Discovery of gonadotropin-inhibitory hormone (GnIH) across vertebrates. Gen Comp Endocrinol (2014) 205:4–10. doi:10.1016/j.ygcen.2014.03.006
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
73. Iwasa T, Matsuzaki T, Murakami M, Kinouchi R, Osugi T, Gereltsetseg G, et al. Developmental changes in the mammalian gonadotropin-inhibitory hormone (GnIH) ortholog RFamide-related peptide (RFRP) and its cognate receptor GPR147 in the rat hypothalamus. Int J Dev Neurosci (2012) 30:31–7. doi:10.1016/j.ijdevneu.2011.10.003
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
74. Koda A, Ukena K, Teranishi H, Ohta S, Yamamoto K, Kikuyama S, et al. A novel amphibian hypothalamic neuropeptide: isolation, localization, and biological activity. Endocrinology (2002) 143:411–9. doi:10.1210/endo.143.2.8630
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
75. Sawada K, Ukena K, Satake H, Iwakoshi E, Minakata H, Tsutsui K. Novel fish hypothalamic neuropeptide – cloning of a cDNA encoding the precursor polypeptide and identification and localization of the mature peptide. Eur J Biochem (2002) 269:6000–8. doi:10.1046/j.1432-1033.2002.03351.x
76. Johnson MA, Tsutsui K, Fraley GS. Rat RFamide-related peptide-3 stimulates GH secretion, inhibits LH secretion, and has variable effects on sex behavior in the adult male rat. Horm Behav (2007) 51:171–80. doi:10.1016/j.yhbeh.2006.09.009
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
77. Ubuka T, Haraguchi S, Tobari Y, Narihiro M, Ishikawa K, Hayashi T, et al. Hypothalamic inhibition of socio-sexual behaviour by increasing neuroestrogen synthesis. Nat Commun (2014) 5:3061. doi:10.1038/ncomms4061
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
78. Biswas S, Jadhao AG, Pinelli C, Palande NV, Tsutsui K. GnIH and GnRH expressions in the central nervous system and pituitary of Indian major carp, Labeo rohita during ontogeny: an immunocytochemical study. Gen Comp Endocrinol (2014). doi:10.1016/j.ygcen.2014.06.005
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
79. Zhang Y, Li S, Liu Y, Lu D, Chen H, Huang X, et al. Structural diversity of the gnih/gnih receptor system in teleost: its involvement in early development and the negative control of LH release. Peptides (2010) 31:1034–43. doi:10.1016/j.peptides.2010.03.003
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
80. Maddineni SR, Ocon-Grove OM, Krzysik-Walker SM, Hendricks GL III, Ramachandran R. Gonadotropin-inhibitory hormone (GnIH) receptor gene is expressed in the chicken ovary: potential role of GnIH in follicular maturation. Reproduction (2008) 135:267–74. doi:10.1530/REP-07-0369
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
81. Amano M, Moriyama S, Iigo M, Kitamura S, Amiya N, Yamamori K, et al. Novel fish hypothalamic neuropeptides stimulate the release of gonadotrophins and growth hormone from the pituitary of sockeye salmon. J Endocrinol (2006) 188:417–23. doi:10.1677/joe.1.06494
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
82. Moussavi M, Wlasichuk M, Chang JP, Habibi HR. Seasonal effect of gonadotrophin inhibitory hormone on gonadotrophin-releasing hormone-induced gonadotroph functions in the goldfish pituitary. J Neuroendocrinol (2013) 25:506–16. doi:10.1111/jne.12024
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
83. Qi X, Zhou W, Li S, Lu D, Yi S, Xie R, et al. Evidences for the regulation of GnRH and GTH expression by GnIH in the goldfish, Carassius auratus. Mol Cell Endocrinol (2013) 366:9–20. doi:10.1016/j.mce.2012.11.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
84. Ukena K, Koda A, Yamamoto K, Kobayashi T, Iwakoshi-Ukena E, Minakata H, et al. Novel neuropeptides related to frog growth hormone-releasing peptide: isolation, sequence, and functional analysis. Endocrinology (2003) 144:3879–84. doi:10.1210/en.2003-0359
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
85. Osugi T, Daukss D, Gazda K, Ubuka T, Kosugi T, Nozaki M, et al. Evolutionary origin of the structure and function of gonadotropin-inhibitory hormone: insights from lampreys. Endocrinology (2012) 153:2362–74. doi:10.1210/en.2011-2046
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
86. Poling MC, Kim J, Dhamija S, Kauffman AS. Development, sex steroid regulation, and phenotypic characterization of RFamide-related peptide (Rfrp) gene expression and RFamide receptors in the mouse hypothalamus. Endocrinology (2012) 153:1827–40. doi:10.1210/en.2011-2049
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
87. Ubuka T, Ueno M, Ukena K, Tsutsui K. Developmental changes in gonadotropin-inhibitory hormone in the Japanese quail (Coturnix japonica) hypothalamo-hypophysial system. J Endocrinol (2003) 178:311–8. doi:10.1677/joe.0.1780311
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
88. Yano T, Iijima N, Hinuma S, Tanaka M, Ibata Y. Developmental expression of RFamide-related peptides in the rat central nervous system. Dev Brain Res (2004) 152:109–20. doi:10.1016/j.devbrainres.2004.06.008
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
89. Legagneux K, Bernard-Franchi G, Poncet F, La Roche A, Colard C, Fellmann D, et al. Distribution and genesis of the RFRP-producing neurons in the rat brain: comparison with melanin-concentrating hormone- and hypocretin-containing neurons. Neuropeptides (2009) 43:13–9. doi:10.1016/j.npep.2008.11.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
90. Quennell JH, Rizwan MZ, Relf HL, Anderson GM. Developmental and steroidogenic effects on the gene expression of RFamide related peptides and their receptor in the rat brain and pituitary gland. J Neuroendocrinol (2010) 22:309–16. doi:10.1111/j.1365-2826.2010.01963.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
91. Johnson MA, Fraley GS. Rat RFRP-3 alters hypothalamic GHRH expression and growth hormone secretion but does not affect KiSS-1 gene expression or the onset of puberty in male rats. Neuroendocrinology (2008) 88:305–15. doi:10.1159/000145718
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
92. Yano T, Iijima N, Kakihara K, Hinuma S, Tanaka M, Ibata Y. Localization and neuronal response of RFamide related peptides in the rat central nervous system. Brain Res (2003) 982:156–67. doi:10.1016/S0006-8993(03)02877-4
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
93. Fukusumi S, Habata Y, Yoshida H, Iijima N, Kawamata Y, Hosoya M, et al. Characteristics and distribution of endogenous RFamide-related peptide-1. Biochim Biophys Acta (2001) 1540:221–32. doi:10.1016/S0167-4889(01)00135-5
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
94. West A, Vojta PJ, Welch DR, Weissman BE. Chromosome localization and genomic structure of the KiSS-1 metastasis suppressor gene (KISS1). Genomics (1998) 54:145–8. doi:10.1006/geno.1998.5566
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
95. Clements MK, McDonald TP, Wang RP, Xie GC, O’Dowd BF, George SR, et al. FMRFamide-related neuropeptides are agonists of the orphan G-protein-coupled receptor GPR54. Biochem Biophys Res Commun (2001) 284:1189–93. doi:10.1006/bbrc.2001.5098
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
96. Kotani M, Detheux M, Vandenbbogaerde A, Communi D, Vanderwinden JM, Le Poul E, et al. The metastasis suppressor gene KiSS-1 encodes kisspeptins, the natural ligands of the orphan G protein-coupled receptor GPR54. J Biol Chem (2001) 276:34631–6. doi:10.1074/jbc.M104847200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
97. Muir AI, Chamberlain L, Elshourbagy NA, Michalovich D, Moore DJ, Calamari A, et al. AXOR12, a novel human G protein-coupled receptor, activated by the peptide KiSS-1. J Biol Chem (2001) 276:28969–75. doi:10.1074/jbc.M102743200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
98. Ohtaki T, Shintani Y, Honda S, Matsumoto H, Hori A, Kanehashi K, et al. Metastasis suppressor gene KiSS-1 encodes peptide ligand of a G-protein-coupled receptor. Nature (2001) 411:613–7. doi:10.1038/35079135
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
99. Lee JH, Welch DR. Suppression of metastasis in human breast carcinoma MDA-MB-435 cells after transfection with the metastasis suppressor gene, KiSS-1. Cancer Res (1997) 57:2384–7.
100. Oakley AE, Clifton DK, Steiner RA. Kisspeptin signaling in the brain. Endocr Rev (2009) 30:713–43. doi:10.1210/er.2009-0005
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
101. de Roux N, Genin E, Carel JC, Matsuda F, Chaussain JL, Milgrom E. Hypogonadotropic hypogonadism due to loss of function of the KiSS1-derived peptide receptor GPR54. Proc Natl Acad Sci U S A (2003) 100:10972–6. doi:10.1073/pnas.1834399100
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
102. Seminara SB, Messager S, Chatzidaki EE, Thresher RR, Acierno JS, Shagoury JK, et al. The GPR54 gene as a regulator of puberty. N Engl J Med (2003) 349:1614–27. doi:10.1056/NEJMoa035322
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
103. Mead EJ, Maguire JJ, Kuc RE, Davenport AP. Kisspeptins: a multifunctional peptide system with a role in reproduction, cancer and the cardiovascular system. Br J Pharmacol (2007) 151:1143–53. doi:10.1038/sj.bjp.0707295
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
104. Sawyer I, Smillie S-J, Bodkin JV, Fernandes E, O’Byrne KT, Brain SD. The vasoactive potential of Kisspeptin-10 in the peripheral vasculature. PLoS One (2011) 6:e14671. doi:10.1371/journal.pone.0014671
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
105. Arai AC, Orwig N. Factors that regulate KiSS1 gene expression in the hippocampus. Brain Res (2008) 1243:10–8. doi:10.1016/j.brainres.2008.09.031
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
106. Arai AC. The role of kisspeptin and GPR54 in the hippocampus. Peptides (2009) 30:16–25. doi:10.1016/j.peptides.2008.07.023
107. Fiorini Z, Jasoni CL. A novel developmental role for kisspeptin in the growth of gonadotrophin-releasing hormone neurites to the median eminence in the mouse. J Neuroendocrinol (2010) 22:1113–25. doi:10.1111/j.1365-2826.2010.02059.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
108. Castano JP, Martinez-Fuentes AJ, Gutierrez-Pascual E, Vaudry H, Tena-Sempere M, Malagon MM. Intracellular signaling pathways activated by kisspeptins through GPR54: do multiple signals underlie function diversity? Peptides (2009) 30:10–5. doi:10.1016/j.peptides.2008.07.025
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
109. Hodne K, Weltzien FA, Oka Y, Okubo K. Expression and putative function of kisspeptins and their receptors during early development in medaka. Endocrinology (2013) 154:3437–46. doi:10.1210/en.2013-1065
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
110. Kitahashi T, Ogawa S, Parhar IS. Cloning and expression of kiss2 in the Zebrafish and Medaka. Endocrinology (2009) 150:821–31. doi:10.1210/en.2008-0940
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
111. Mohamed JS, Benninghoff AD, Holt GJ, Khan IA. Developmental expression of the G protein-coupled receptor 54 and three GnRH mRNAs in the teleost fish cobia. J Mol Endocrinol (2007) 38:235–44. doi:10.1677/jme.1.02182
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
112. Weltzien FA, Hildahl J, Hodne K, Okubo K, Haug TM. Embryonic development of gonadotrope cells and gonadotropic hormones – lessons from model fish. Mol Cell Endocrinol (2014) 385:18–27. doi:10.1016/j.mce.2013.10.016
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
113. Olbrich T, Ziegler E, Tuerk G, Schubert A, Emons G, Gruendker C. Kisspeptin-10 inhibits bone-directed migration of GPR54-positive breast cancer cells: evidence for a dose-window effect. Gynecol Oncol (2010) 119:571–8. doi:10.1016/j.ygyno.2010.08.018
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
114. Navenot JM, Wang ZX, Chopin N, Fujii N, Peiper SC. Kisspeptin-10-induced signaling of GPR54 negatively regulates chemotactic responses mediated by CXCR4: a potential mechanism for the metastasis suppressor activity of kisspeptins. Cancer Res (2005) 65:10450–6. doi:10.1158/0008-5472.CAN-05-1757
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
115. Mizoguchi T, Verkade H, Heath JK, Kuroiwa A, Kikuchi Y. Sdf1/Cxcr4 signaling controls the dorsal migration of endodermal cells during zebrafish gastrulation. Development (2008) 135:2521–9. doi:10.1242/dev.020107
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
116. Ma Q, Jones D, Borghesani PR, Segal RA, Nagasawa T, Kishimoto T, et al. Impaired B-lymphopoiesis, myelopoiesis, and derailed cerebellar neuron migration in CXCR4- and SDF-1-deficient mice. Proc Natl Acad Sci U S A (1998) 95:9448–53. doi:10.1073/pnas.95.16.9448
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
117. Nagasawa T, Tachibana K, Kishimoto T. A novel CXC chemokine PBSF/SDF-1 and its receptor CXCR4: their functions in development, hematopoiesis and HIV infection. Semin Immunol (1998) 10:179–85. doi:10.1006/smim.1998.0128
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
118. Zhao Y, Lin M-CA, Mock A, Yang M, Wayne NL. Kisspeptins modulate the biology of multiple populations of gonadotropin-releasing hormone neurons during embryogenesis and adulthood in zebrafish (Danio rerio). PLoS One (2014) 9:e104330. doi:10.1371/journal.pone.0104330
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
119. Pasquier J, Lafont AG, Rousseau K, Querat B, Chemineau P, Dufour S. Looking for the bird Kiss: evolutionary scenario in sauropsids. BMC Evol Biol (2014) 14:30. doi:10.1186/1471-2148-14-30
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
120. Franceschini I, Desroziers E. Development and aging of the Kisspeptin-GPR54 system in the mammalian brain: what are the impacts on female reproductive function? Front Endocrinol (2013) 4:22. doi:10.3389/fendo.2013.00022
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
121. Poling MC, Kauffman AS. Organizational and activational effects of sex steroids on kisspeptin neuron development. Front Neuroendocrinol (2013) 34:3–17. doi:10.1016/j.yfrne.2012.06.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
122. Constantin S, Caligioni CS, Stojilkovic S, Wray S. Kisspeptin-10 facilitates a plasma membrane-driven calcium oscillator in gonadotropin-releasing hormone-1 neurons. Endocrinology (2009) 150:1400–12. doi:10.1210/en.2008-0979
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
123. Desroziers E, Droguerre M, Bentsen AH, Robert V, Mikkelsen JD, Caraty A, et al. Embryonic development of kisspeptin neurones in rat. J Neuroendocrinol (2012) 24:1284–95. doi:10.1111/j.1365-2826.2012.02333.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
124. Kumar D, Freese M, Drexler D, Hermans-Borgmeyer I, Marquardt A, Boehm U. Murine arcuate nucleus kisspeptin neurons communicate with GnRH neurons in utero. J Neurosci (2014) 34:3756–66. doi:10.1523/JNEUROSCI.5123-13.2014
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
125. Constantin S, Caraty A, Wray S, Duittoz AH. Development of gonadotropin-releasing hormone-1 secretion in mouse nasal explants. Endocrinology (2009) 150:3221–7. doi:10.1210/en.2008-1711
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
126. Castellano JM, Navarro VM, Fernández-Fernández R, Castaño JP, Malagón MM, Aguilar E, et al. Ontogeny and mechanisms of action for the stimulatory effect of kisspeptin on gonadotropin-releasing hormone system of the rat. Mol Cell Endocrinol (2006) 257–258:75–83. doi:10.1016/j.mce.2006.07.002
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
127. Clarkson J, Herbison AE. Postnatal development of kisspeptin neurons in mouse hypothalamus; sexual dimorphism and projections to gonadotropin-releasing hormone neurons. Endocrinology (2006) 147:5817–25. doi:10.1210/en.2006-0787
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
128. Clarkson J, Boon WC, Simpson ER, Herbison AE. Postnatal development of an estradiol-kisspeptin positive feedback mechanism implicated in puberty onset. Endocrinology (2009) 150:3214–20. doi:10.1210/en.2008-1733
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
129. Semaan SJ, Murray EK, Poling MC, Dhamija S, Forger NG, Kauffman AS. BAX-dependent and BAX-independent regulation of Kiss1 neuron development in mice. Endocrinology (2010) 151:5807–17. doi:10.1210/en.2010-0783
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
130. Cao J, Patisaul HB. Sexually dimorphic expression of hypothalamic estrogen receptors α and β and kiss1 in neonatal male and female rats. J Comp Neurol (2011) 519:2954–77. doi:10.1002/cne.22648
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
131. Takumi K, Iijima N, Ozawa H. Developmental changes in the expression of kisspeptin mRNA in rat hypothalamus. J Mol Neurosci (2011) 43:138–45. doi:10.1007/s12031-010-9430-1
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
132. Knoll JG, Clay C, Bouma G, Henion T, Schwarting G, Millar R, et al. Developmental profile and sexually dimorphic expression of Kiss1 and Kiss1r in the fetal mouse brain. Front Endocrinol (2013) 4:140. doi:10.3389/fendo.2013.00140
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
133. Lapatto R, Pallais JC, Zhang D, Chan Y-M, Mahan A, Cerrato F, et al. Kiss1(-/-) mice exhibit more variable hypogonadism than Gpr54(-/-) mice. Endocrinology (2007) 148:4927–36. doi:10.1210/en.2007-0078
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
134. Kanda S, Oka Y. Evolutionary insights into the steroid sensitive kiss1 and kiss2 neurons in the vertebrate brain. Front Endocrinol (2012) 3:28. doi:10.3389/fendo.2012.00028
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
135. Lyubimov Y, Engstrom M, Wurster S, Savola JM, Korpi ER, Panula P. Human kisspeptins activate neuropeptide FF2 receptor. Neuroscience (2010) 170:117–22. doi:10.1016/j.neuroscience.2010.06.058
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
136. Oishi S, Misu R, Tomita K, Setsuda S, Masuda R, Ohno H, et al. Activation of neuropeptide FF receptors by kisspeptin receptor ligands Acs. Med Chem Lett (2011) 2:53–7. doi:10.1021/ml1002053
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
137. Mayer C, Boehm U. Female reproductive maturation in the absence of kisspeptin/GPR54 signaling. Nat Neurosci (2011) 14:704–10. doi:10.1038/nn.2818
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
138. Navarro VM, Castellano JM, Fernández-Fernández R, Barreiro ML, Roa J, Sanchez-Criado JE, et al. Developmental and hormonally regulated messenger ribonucleic acid expression of KiSS-1 and its putative receptor, GPR54, in rat hypothalamus and potent luteinizing hormone-releasing activity of KiSS-1 peptide. Endocrinology (2004) 145:4565–74. doi:10.1210/en.2004-0413
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
139. Franceschini I, Lomet D, Cateau M, Delsol G, Tillet Y, Caraty A. Kisspeptin immunoreactive cells of the ovine preoptic area and arcuate nucleus co-express estrogen receptor alpha. Neurosci Lett (2006) 401:225–30. doi:10.1016/j.neulet.2006.03.039
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
140. Chartrel N, Dujardin C, Anouar Y, Leprince J, Decker A, Clerens S, et al. Identification of 26RFa, a hypothalamic neuropeptide of the RFamide peptide family with orexigenic activity. Proc Natl Acad Sci U S A (2003) 100:15247–52. doi:10.1073/pnas.2434676100
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
141. Liu Y, Zhang Y, Li S, Huang W, Liu X, Lu D, et al. Molecular cloning and functional characterization of the first non-mammalian 26RFa/QRFP orthologue in Goldfish, Carassius auratus. Mol Cell Endocrinol (2009) 303:82–90. doi:10.1016/j.mce.2009.01.009
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
142. Fukusumi S, Yoshida H, Fujii R, Maruyama M, Komatsu H, Habata Y, et al. A new peptidic ligand and its receptor regulating adrenal function in rats. J Biol Chem (2003) 278:46387–95. doi:10.1074/jbc.M305270200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
143. Jiang Y, Luo L, Gustafson EL, Yadav D, Laverty M, Murgolo N, et al. Identification and characterization of a novel RF-amide peptide ligand for orphan G-protein-coupled receptor SP9155. J Biol Chem (2003) 278:27652–7. doi:10.1074/jbc.M302945200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
144. Bruzzone F, Lectez B, Alexandre D, Jegou S, Mounien L, Tollemer H, et al. Distribution of 26RFa binding sites and GPR103 mRNA in the central nervous system of the rat. J Comp Neurol (2007) 503:573–91. doi:10.1002/cne.21400
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
145. Ukena K, Tachibana T, Iwakoshi-Ukena E, Saito Y, Minakata H, Kawaguchi R, et al. Identification, localization, and function of a novel avian hypothalamic neuropeptide, 26RFa, and its cognate receptor, G protein-coupled receptor-103. Endocrinology (2010) 151:2255–64. doi:10.1210/en.2009-1478
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
146. Tobari Y, Iijima N, Tsunekawa K, Osugi T, Haraguchi S, Ubuka T, et al. Identification, localisation and functional implication of 26RFa orthologue peptide in the brain of Zebra finch (Taeniopygia guttata). J Neuroendocrinol (2011) 23:791–803. doi:10.1111/j.1365-2826.2011.02179.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
147. Takayasu S, Sakurai T, Iwasaki S, Teranishi H, Yamanaka A, Williams SC, et al. A neuropeptide ligand of the G protein-coupled receptor GPR103 regulates feeding, behavioral arousal, and blood pressure in mice. Proc Natl Acad Sci U S A (2006) 103:7438–43. doi:10.1073/pnas.0602371103
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
148. Chartrel N, Bruzzone F, Leprince J, Tollemer H, Anouar Y, Do-Rego JC, et al. Structure and functions of the novel hypothalamic RFamide neuropeptides R-RFa and 26RFa in vertebrates. Peptides (2006) 27:1110–20. doi:10.1016/j.peptides.2005.06.035
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
149. Ramanjaneya M, Karteris E, Chen J, Rucinski M, Ziolkowska A, Ahmed N, et al. QRFP induces aldosterone production via PKC and T-type calcium channel-mediated pathways in human adrenocortical cells: evidence for a novel role of GPR103. Am J Physiol Endocrinol Metab (2013) 305:E1049–58. doi:10.1152/ajpendo.00191.2013
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
150. Baribault H, Danao J, Gupte J, Yang L, Sun BH, Richards W, et al. The G-protein-coupled receptor GPR103 regulates bone formation. Mol Cell Biol (2006) 26:709–17. doi:10.1128/MCB.26.2.709-717.2006
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
151. Alonzeau J, Alexandre D, Jeandel L, Courel M, Hautot C, El Yamani FZ, et al. The neuropeptide 26RFa is expressed in human prostate cancer and stimulates the neuroendocrine differentiation and the migration of androgeno-independent prostate cancer cells. Eur J Cancer (2013) 49:511–9. doi:10.1016/j.ejca.2012.05.028
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
152. Roszer T, Banfalvi G. FMRFamide-related peptides: anti-opiate transmitters acting in apoptosis. Peptides (2012) 34:177–85. doi:10.1016/j.peptides.2011.04.011
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
153. Lecron JC, Minault M, Allard M, Delaforest PG, Gombert J, Simonnet G. Modulation of human lymphocyte-proliferation by FLFQPQRFamide, a FMRFamide-like peptide with anti-opiate properties. J Neuroimmunol (1992) 38:1–8. doi:10.1016/0165-5728(92)90084-X
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
154. Roszer T, Kappelmayer J, Nagy GG, Szentmiklosi A, Basnakian AG, Banfalvi G. The neuropeptide FMRFamide can protect cells against apoptosis in the snail digestive gland. Apoptosis (2006) 11:173–82. doi:10.1007/s10495-006-3391-4
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
155. Nystedt JM, Brandt A, Vilim FS, Ziff EB, Panula P. Identification of transcriptional regulators of neuropeptide FF gene expression. Peptides (2006) 27:1020–35. doi:10.1016/j.peptides.2005.07.029
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Keywords: RFa, NPFF, PrRP, Kiss, GnIH, 26RFa/QRFP, early development, brain development
Citation: Sandvik GK, Hodne K, Haug TM, Okubo K and Weltzien F-A (2014) RFamide peptides in early vertebrate development. Front. Endocrinol. 5:203. doi: 10.3389/fendo.2014.00203
Received: 18 July 2014; Accepted: 16 November 2014;
Published online: 04 December 2014.
Edited by:
Sylvie Dufour, Muséum National d’Histoire Naturelle, FranceReviewed by:
Gregoy Y. Bedecarrats, University of Guelph, CanadaAlexander S. Kauffman, University of California San Diego, USA
Copyright: © 2014 Sandvik, Hodne, Haug, Okubo and Weltzien. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Finn-Arne Weltzien, Department of Basic Sciences and Aquatic medicine, Norwegian University of Life Sciences – Campus Adamstuen, PO Box 8146 Dep, Oslo 0033, Norway e-mail: finn-arne.weltzien@nmbu.no
 Guro Katrine Sandvik
Guro Katrine Sandvik Kjetil Hodne
Kjetil Hodne Trude Marie Haug
Trude Marie Haug Kataaki Okubo
Kataaki Okubo Finn-Arne Weltzien
Finn-Arne Weltzien