Success of Wildlife Disease Treatment Depends on Host Immune Response
- 1Ecology, Evolution, and Natural Resources, Rutgers, The State University of New Jersey, New Brunswick, NJ, USA
- 2Rutgers Cooperative Extension, New Jersey Agricultural Experiment Station, Rutgers, The State University of New Jersey, New Brunswick, NJ, US
- 3Ecology and Evolutionary Biology, University of Tennessee, Knoxville, TN, USA
- 4National Institute for Mathematical and Biological Synthesis, University of Tennessee, Knoxville, TN, USA
Infectious diseases significantly threaten many wildlife populations. Proposed treatments often fail to consider resulting impacts on natural host defense mechanisms and ongoing infection risks. We demonstrate the importance of prior knowledge of host defenses by modeling outcomes of various treatment regimes on bat populations infected with white-nose syndrome (WNS), a disease that has caused declines in multiple North American species. Despite differences in species susceptibility to the causative fungus, how hosts respond to and survive infection remains unclear. We explore three potential host-defense mechanisms: upregulation of the responsive innate or adaptive immune systems, and rapid evolution that enables evolutionary rescue. We demonstrate that treatment can accelerate extirpation (relative to inaction) if hosts are defended by responsive immunity; whereas, treatment can cause populations undergoing evolutionary rescue to regain population viability faster. We conclude that successful treatment of wildlife diseases must first consider how individuals respond to and survive infection.
Introduction
Infectious disease historically has not been considered a major cause of species' extinction and, therefore, disease management has thus far remained a low priority relative to other conservation actions (Fisher et al., 2012; Pullin et al., 2013). Mounting evidence suggests, however, that the influence of infectious disease on extinction risk has been largely underestimated and that the coupled effects of anthropogenic forces on hosts and diseases will greatly contribute to our global biodiversity crisis (Smith et al., 2009; McCallum, 2012). Human influences drastically alter the ecological landscape in which pathogens and potential hosts interact (Burge et al., 2014), increasing both host susceptibility to infection and pathogen virulence (Brearley et al., 2013; Becker et al., 2015). Anthropogenic climate change and globalization have expanded the natural range of many pathogens and/or their suite of host species, releasing them from co-evolutionary forces that may have maintained lower-virulence strains (e.g., Brasier and Buck, 2001). Climate change is especially consequential for fungal pathogens, which are becoming increasingly heat-tolerant and thus able to infect previously protected endothermic animals (Robert and Casadevall, 2009; Garcia-Solache and Casadevall, 2010). The unprecedented frequency of disease emergence (Harvell et al., 2002; Fisher et al., 2012) and the strength of impacts on wild populations (Daszak et al., 2000), increasingly point to the importance of successful mitigation of disease in species' conservation.
Advances in antibiotics, fungicides, and probiotics make effective treatment of wildlife diseases more achievable than ever. However, while lab experiments or short-term field trials may produce promising results (e.g., Cornelison et al., 2014), deployment of a given treatment strategy may have unforeseen negative consequences on the host population, especially if host response to the pathogen (or its removal) is not considered in advance. Therefore, it is critical to fully evaluate host-pathogen interactions prior to the deployment of a treatment strategy.
Species broadly mitigate pathogen exposure in two ways: either through an immunological response at the individual level or an evolutionary response at the population level (or both). Immunological responses can be innate or adaptive, and the activation of either can have different consequences on the host. The innate immune system is often a first line of defense, identifying invading pathogens using pattern recognition receptors to set off signaling cascades (i.e., Toll pathway, Lai et al., 2009) and result in general responses such as inflammation and ingestion of pathogens by macrophages (e.g., Rizzetto et al., 2013). The adaptive immune system is more specialized, using antigen-specific cells to recognize individual pathogens and memory cells to mount responses upon repeated exposure (note this adaptation is individual not evolutionary, Hebart et al., 2002; Romani, 2004). In contrast, an evolutionary response acts at the population level, whereby pathogen exposure causes steep demographic decline, inducing an intense selective pressure that favors survival of individuals with a specific genetic signature (i.e., resistance). A population consisting of primarily resistant individuals could revert to positive growth without human intervention, a phenomenon known as evolutionary rescue (Gonzalez et al., 2013), but may benefit from targeted treatment while at low population size (Maslo and Fefferman, 2015). These stark differences in potential disease-defensive mechanisms suggest the possibility for equally significant differences in long-term population effects of treatment. To demonstrate how overlooking these potential differences can compromise conservation efforts, we consider a currently unfolding case study: treatment of white-nose syndrome (WNS) in North American bats.
In the 10 years following WNS emergence, the disease has caused severe declines in several bat species in eastern North America (Langwig et al., 2015a). The conservation community has made significant strides in understanding this previously unknown disease of bats, including identification of the causative agent, Pseudogymnoascus destructans (Lorch et al., 2011). P. destructans is non-native to North America and appears to have been transported by humans from its endemic range in Europe (Puechmaille et al., 2011). The fungus colonizes bats during an immunosuppressed torpor, but while European bats tolerate infection (Puechmaille et al., 2011), naïve North American bats experience a disruption of physiological homeostasis leading to measureable population declines (Willis et al., 2011; Verant et al., 2014). Severity of these declines has created a sense of urgency for conservation intervention in the form of chemical or biological control (e.g., fungistatic volatile organic compounds; Cornelison et al., 2014), probiotics (e.g., Cheng et al., 2016), or synthetic compounds (e.g., Raudabaugh and Miller, 2015) to either prevent pathogen exposure to naïve populations or reduce disease intensity in infected populations.
Population declines due to WNS vary among species (Frick et al., 2015), and drivers of observed survival patterns remain unclear. For example, Myotis lucifugus (little brown bat) survival rebounded markedly in the years following disease emergence (Maslo et al., 2015) suggesting a rapid evolutionary response (Maslo and Fefferman, 2015). However, M. lucifugus exposed to P. destructans also exhibit elevated expression of genes related to the responsive innate immune system (Field et al., 2015). In mammalian fungal infections, innate immunity is thought to be particularly important in regulating infection and suppressing memory T cells to avoid potentially damaging inflammation (Bacher et al., 2014).
To explore treatment impacts on population recovery under each of the possible disease-defense mechanisms, we applied to an existing model set predicting WNS-infected population dynamics (Maslo and Fefferman, 2015), a suite of antifungal treatment regimes informed by the conservation community (WNS workshop, July 2015). Scenarios varied in treatment efficacy and the proportion of remnant populations treated. For simplicity, we consider only the responsive innate immune system, which is upregulated upon pathogen exposure, and not the baseline innate immune system, which protects the host indiscriminately and can be assumed to act equally in all of the considered scenarios. We evaluate persistence time of the population under each scenario, and determine the minimum population size for populations undergoing an evolutionary response. While these results may inform discussions of treatment for WNS in bat populations, our broader goal is to demonstrate the need for consideration of disease-defense mechanisms at play in any infected population being considered for treatment-based conservation actions.
Methods
Host Response-Based Population Viability Models
Maslo and Fefferman (2015) recently modeled the population dynamics of Myotis lucifugus undergoing responsive immunity or evolutionary rescue in the face of WNS (summarized below). The evolutionary rescue model makes the conservative assumption that phenotypic robustness to WNS was based upon a Boolean protective genetic profile. Bats within an uninfected population were either entirely robust, r, or susceptible, wt, to WNS-induced mortality. Using published or estimated vital rates from before ( and ) and after ( and ) P. destructans introduction (Frick et al., 2010b; Maslo et al., 2015), Maslo and Fefferman (2015) parameterized four two-stage Lefkovitch matrices (Morris and Doak, 2002), each representing population growth of one genetic segment of the population in both the uninfected and infected state:
where S represents survival of M. lucifugus; B represents the proportion of females breeding; and the subscripts j and a indicate values for 1st-year or adult bats, respectively. They set 1st-year bat survival as a constant proportion (0.47) of adult survival (Frick et al., 2010b), and held maternity (m) constant at F = 1, as M. lucifugus typically produce a single pup per year (Barclay et al., 2003).
Starting with an initial population vector of 5,000 total individuals, they estimated initial prevalence of the robust genetic profile in the population to be ρ0 and then split the initial demography into two corresponding initial vectors and such that and This approach deviated from traditional PVA, which would apply a single population projection matrix to a single state vector representing the current stage-based demography of the population. Maslo and Fefferman (2015) then projected population growth in the absence of disease by independently applying the appropriate matrix to each vector to obtain the population vector for the next year: and . Maslo and Fefferman (2015) then computed the realized total population vector , and resulting growth rate.
The model assumes environmental contamination for an entire habitat (e.g., hibernaculum) with 100% infection following contamination. They define It to be the percent of the population infected in year t and assume that the rate of exposure/infection in a hibernaculum is independent of genetic robustness. Using these probabilities and matrices, they then computed the subpopulations in year t + 1 within a single population as the sum of individuals that are wild type and uninfected: , robust and uninfected: , wild type and infected: , and robust and infected: .
Additionally, 0.1% of the offspring from same profile pairs spontaneously mutate to the other profile (i.e., wt × wt results in a Robust offspring), and r × wt results in 50% of each profile. Assuming random mating among adults Maslo and Fefferman (2015) “corrected” each of the projected population vectors by subtracting the offspring with different genetic profiles from the projected population vector associated with the matrix relevant for the parents, and adding them to the offspring in the vector associated with the appropriate subpopulation for the resulting juvenile.
Under evolutionary rescue, the resulting total population vector in each year was the sum of the individuals within each subpopulation, and the stabilized growth rate (λ = 1.05) was representative of the shifting proportion of genetic robustness within the population (Maslo and Fefferman, 2015). Under responsive immunity, the model projected a single state vector (N = 5000) representing pre-WNS stage-based demographic structure through a single projection matrix (parameterized with survival estimates generated from population counts of multiple infected hibernacula within the first three years of WNS emergence; Frick et al., 2010a). The model output predicted continued population declines stabilizing at λ = 0.89 after 3 years.
For the current analysis, we separated the acquired immune response into adaptive and innate immunity and used both published and estimated vital rates. Both provide protection against mortality for individuals that survived initial exposure, but adaptive immunity confers life-long individual protection, whereas the innate immune protection is lost in the absence of continued pathogen exposure. Individuals with adaptive immunity were therefore modeled in the same way as (Maslo and Fefferman, 2015). In contrast, treated and re-exposed individuals with innate immunity suffered the same initial mortality risks from re-exposure as though they had never previously been immune (though if they survived, they would again be protected throughout the duration of their continuous exposure). In all other ways, this population was also modeled as in Maslo and Fefferman (2015) (Figure 1). Note that this representation of innate immunity can vary either in dose-response curves, or in physiological state of the individuals. However, we do assume that rapidity of the innate response is not itself consistent for an individual over time. We make this assumption because such a consistent response would likely have a genetic component and be considered a potential mechanism for evolutionary rescue, thus covered by that scenario.
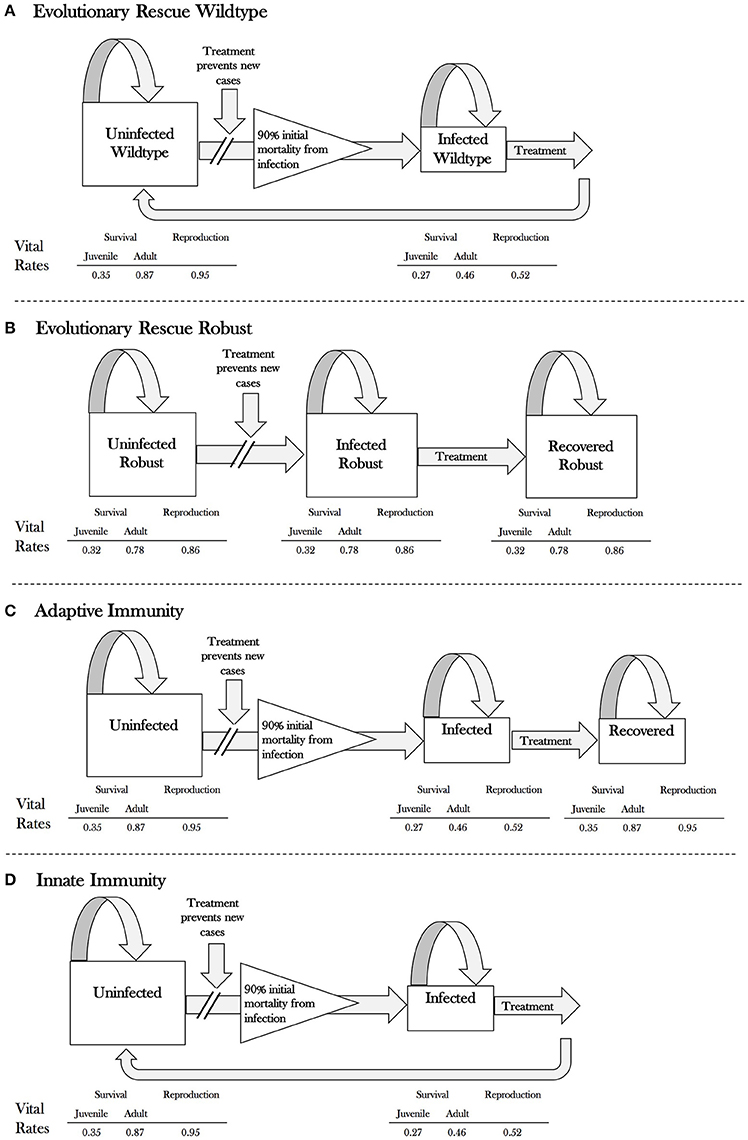
Figure 1. Flow chart of the model under each scenario: (A) evolutionary rescue for wildtype individuals, (B) evolutionary rescue for robust individuals, (C) adaptive immunity for all (wildtype) individuals, and (D) innate immunity for all (wildtype) individuals. Population vital rates (survival and reproduction) for juvenile and adult little brown bats under each scenario are indicated below the flowchart.
Treatment Regimes
We applied a suite of treatment regimes to the three host response scenarios (Figure 2), making the following assumptions based on current treatment research: (1) treatment application occurs just prior to or during the hibernation period; (2) no immigration/emigration occurs after treatment deployment; and (3) treatment suppresses P. destructans growth for a single hibernation season. In the pulsed model (Figure 2), bats are treated in the 1st, 5th, and 10th year following WNS arrival. We modeled constant treatment application regimes, beginning 5 or 15 years following WNS arrival at a site. We also examined the impacts of an interval treatment regime, where bats are treated annually between 5 and 15 years following WNS arrival. We modeled the demographic impacts of each treatment regime for treatments that were 25, 50, 75 or 100% effective and were applied to 25, 50, 75 or 100% of the population. We assumed a quasi-extinction threshold of 150 individuals, below which a population was considered extirpated. If a population reverted to positive growth, we calculated the time required for the abundance to reach its minimum viable population size (assumed to be 2,500 individuals based upon a review of finite rates of increase at WNS-infected hibernacula of various sizes; Maslo and Fefferman, 2015).
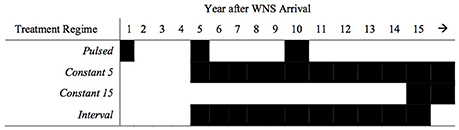
Figure 2. Hypothetical treatment regimes for bat colonies infected with white-nose syndrome. Pulsed treatments occur in the 1st, 5th, and 10th year following disease emergence. Constant 5 and Constant 15 treatments occur annually beginning in Years 5 and 15, respectively. Interval treatments occur annually between the 5th and 15th year following disease emergence.
Results
Model results predict that populations exhibiting a responsive immune reaction after initial pathogen exposure persist for 13 years without treatment application (Figure 3). Regardless of treatment effectiveness or the proportion of the population treated, all populations persist indefinitely when treatment is applied annually beginning in Year 5 of a WNS epizootic. In contrast, beginning treatment in Year 15 results in a persistence time of only 13 years for populations exhibiting responsive immunity.
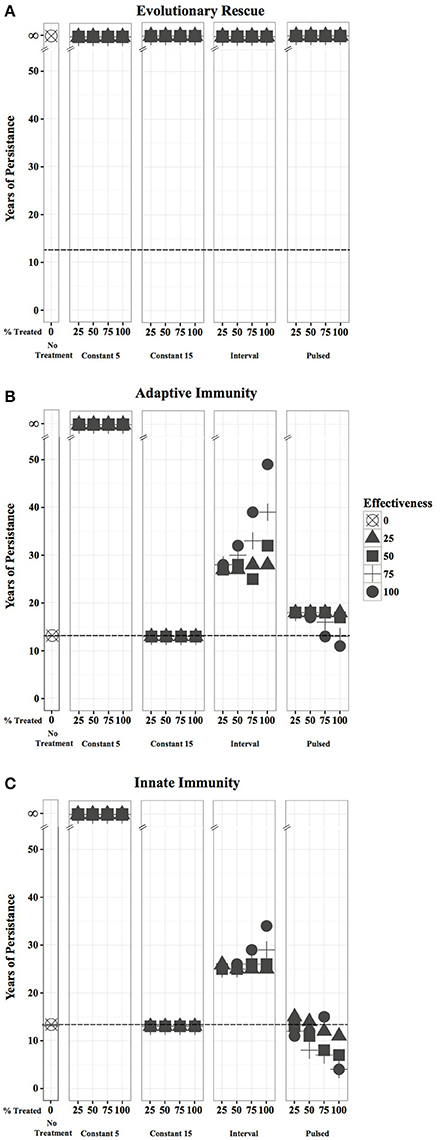
Figure 3. Persistence time for a hibernating bat population if 0, 25, 50, 75 or 100% of the population is treated with an agent that is 0, 25, 50, 75, and 100% effective. Results are shown for bats exhibiting: (A) evolutionary rescue; (B) adaptive immunity; and (C) responsive innate immunity. Dotted lines represent the years to which populations will persist if no treatment is applied.
Pulsed treatments result in expedited extirpation of populations with immunity within 10–21 years, with more effective treatments decreasing years of persistence, a particularly strong trend for innate immunity scenarios. Interval treatments increase the persistence time of populations with adaptive immunity by two to 11 years when >50% of the population is inoculated at treatment efficacies of ≥50%, and persistence time generally increases with increasing treatment effectiveness and proportion of population treated. However, interval treatments of populations with innate immunity only drastically increase persistence time when the entire population is treated with ≥75% efficacy.
At low treatment efficacies (25–50%), the proportion of the population treated minimally impacts the resulting persistence time of each treatment regime across all host responses. As effectiveness increases, treating a large proportion (75–100%) of the population can have appreciable positive or negative impacts on persistence time, extending it for 21 years when an interval treatment is applied to a population with adaptive immunity, or decreasing it to 4 years when a pulsed treatment is applied to a population with innate immunity.
All populations undergoing evolutionary rescue revert to positive growth between 5 and 11 years after disease emergence and persist indefinitely regardless of treatment regime, effectiveness, or the proportion of population treated (Figure 3). With no intervention, a population undergoing evolutionary rescue reaches its lowest size of 626 individuals in 10 years and is predicted to return to an abundance of 2,500 individuals within 39 years. Implementing constant annual treatments beginning in Year 15 reduces that timeframe minimally (1–2 years) across all treatment efficacies, regardless of the proportion of the population treated (Figure 4), and increases the minimum population size only marginally to 686 individuals. At low treatment efficacies (25–50%), the proportion of the population treated does not drastically alter the time required for a population to achieve its minimum viable population size (1–8 years). However, treating at least 75% of the population annually starting in Year 5, once every five years (pulsed), or in Years 5–15 can increase the absolute minimum population size by 35–84%. At 75–100% effectiveness, treating more individuals in a population can significantly reduce the time required for abundance to reach 2,500 individuals, particularly if the treatment is applied annually beginning in Year 5 or between Years 5–15. Treating 100% of the population with a 100% effective treatment under these two scenarios can increase the population to its minimum viable population size in as little as nine years (Figure 4).
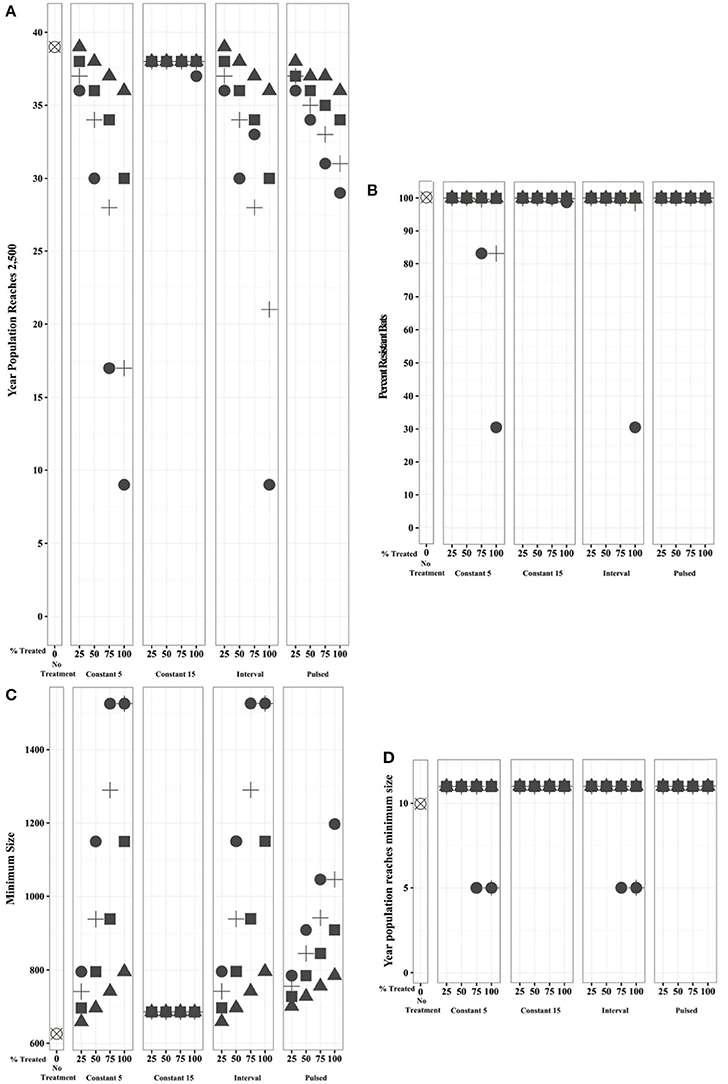
Figure 4. Bats undergoing evolutionary rescue under different treatment regimes display differences in the: (A) time until the population reaches its minimum viable population size of 2,500 individuals; (B) proportion of resistant individuals within the population when it reaches its minimum viable population size; (C) population size at its inflection point (just prior to reversion to positive growth); and (D) year after WNS in which the population reaches this minimum size.
Importantly, the proportion of resistant bats within the population of 2,500 individuals remains above 99% in all but four cases. Annual treatment beginning in Year 5 of either all bats treated at 75% effectiveness or 75% of the bats treated with a 100% effective agent reduces the proportion of resistant bats to 83%. Treating all bats with a 100% effective agent annually beginning in Year 5 or between Years 5 and 10 allows non-resistant bats to persist, thus reducing the proportion of resistant bats to 31% (Figure 4).
Discussion
Implications for Bat Conservation
For bats undergoing either type of immune response, annual treatment beginning early in the disease phase may be extremely effective in reversing demographic decline and creating a positive growth trajectory. However, long-term treatment of multiple bat populations presents several logistical challenges. Treatments applied directly to bats would require significant numbers of individuals to be captured at least once each year (treatments may not last a full hibernation season). Traditional bat capture methods (e.g., harp traps, mist nests) may not guarantee treatment of even 25% of the population in any given year. Actively treating bats during the hibernation period would increase the likelihood that the appropriate threshold number of bats are treated but would disturb hibernating populations unless a passive delivery system is implemented. Hibernaculum treatments (e.g., antifungal treatment of surfaces, VOC treatment of air) may offer a more practicable approach, but treating every infected North American hibernaculum in perpetuity is not sustainable. Identifying the appropriate treatment approach becomes important when monetary and physical feasibility is taken into account. Furthermore, different bat species may be responding to the disease differently, understanding the host response will help to prioritize treatment of species and determine whether one treatment regime is appropriate for all species or if individual treatment plans should be developed.
Implications for All Treatment-Based Conservation
Understanding a host's response to a pathogen prior to intervention is critical to success. While it is tempting to assume that any decrease in pathogen load would be beneficial, the physiology of natural immunity may not always allow such black-and-white solutions. As we have shown, if individuals upregulate physiologically costly innate immune protections only during pathogen exposure and downregulate them once the threat is passed, removal of the trigger for protection can cause a protected population to revert to vulnerability. In these cases, treatment in the absence of reasonable belief that the pathogen will not be reintroduced until after the population has recovered to pre-infection sizes (or beyond) could be catastrophic. Of course, the disease in the absence of treatment may itself be catastrophic. If the ongoing mortality is sufficiently high to compromise the long-term population growth rates, treatment may be the only option. In this case, if innate immunity is providing protection, we must recognize the risks of inconsistent treatment regimes and commit to constant efforts. There is still a point of no return beyond which even constant treatment cannot guarantee the persistence of the population (e.g., small population paradigm, Allee effects, etc.). If the only mechanisms of protection are from immunity and not evolutionary rescue, treatment must begin early enough to preserve a sufficient number of mostly healthy, breeding individuals. Models of this sort will be a critical tool in evaluating when this point-of-no return might happen, what level of efficacy is required and what treatment regime will ensure survival.
The prospects for a population are better under the processes of evolutionary rescue even without intervention. In these cases, treatment becomes a beneficial mechanism for support of the population to mitigate the initial drop in population size as infection decimates individuals without protective genotypes. Increasing the survival of unprotected individuals can prevent the population from growing too small, or being moderately low for too long. However, treatment will also ease the intensity of the selective pressure, slowing the rate of replacement of unprotected individuals by protected individuals. The simulations presented here assumed 50% probability of protection for the offspring of mating between protected and unprotected individuals (i.e., a dominant trait on a single locus that assorts independently). In reality, genetically-derived protection against infection is likely due to a suite of genes that do not assort simply, and dilution of selective pressure must therefore also be considered.
Lastly, evolutionary rescue is not mutually exclusive with either immune protection, meaning that the impact of treatment on real-world population persistence will entail a complicated trade-off between the dilution of selective forces, the mitigation of mortality costs from new infections due to ongoing transmission, and the potential for compromising ongoing innate immune protection in already-exposed individuals. By tailoring models to reflect the genetic and immunological processes of particular systems, we can begin to consider certain treatment strategies, weighing these impacts explicitly, and discovering how best to manage populations under threat.
The implications of these models extend beyond the WNS epizootic. For the increasing numbers of emerging wildlife diseases, such as the recently discovered snake fungal disease (Tetzlaff et al., 2015) or the multiple etiologically complicated coral diseases (Weil and Rogers, 2011; Sutherland et al., 2016), our study underscores the importance of studying the host response concurrently to investigating mechanisms of infection and transmission. The fungus Batrachochytrium dendrobatidi has caused worldwide declines in amphibian populations, leading to coordinated efforts to control pathogen spread, understand mechanisms of transmission and infection, and treat the disease (Whilde et al., 2017). However, efforts to treat infection have been largely unsuccessful and different species have varying degrees of susceptibility (Heard et al., 2011; Woodhams et al., 2012). Recent evidence shows that immunosuppression of the host by the pathogen may play a critical role in its ability to infect hosts (Savage et al., 2016) indicating that a better understanding of the host response(s) will help to elucidate a successful treatment plan for this disease. For more well-characterized diseases, such as Dermo in oysters (Burge et al., 2014) or chronic wasting disease in cervids (Williams et al., 2002), where the host response may be better understood, our study demonstrates the necessity of incorporating what is known of the host response into an efficient and effective treatment plan.
The implications of these models extend beyond the WNS epizootic. For the increasing numbers of emerging wildlife diseases, such as the recently discovered snake fungal disease (Tetzlaff et al., 2015) or the multiple etiologically complicated coral diseases (Weil and Rogers, 2011; Sutherland et al., 2016), our study underscores the importance of studying the host response concurrently to investigating mechanisms of infection and transmission. The fungus Batrachochytrium dendrobatidi has caused worldwide declines in amphibian populations, leading to coordinated efforts to control pathogen spread, understand mechanisms of transmission and infection, and treat the disease (Whilde et al., 2017). However, efforts to treat infection have been largely unsuccessful and different species have varying degrees of susceptibility (Heard et al., 2011; Woodhams et al., 2012). Recent evidence shows that immunosuppression of the host by the pathogen may play a critical role in its ability to infect hosts (Savage et al., 2016) indicating that a better understanding of the host response(s) will help to elucidate a successful treatment plan for this disease. For more well-characterized diseases, such as Dermo in oysters (Burge et al., 2014) or chronic wasting disease in cervids (Williams et al., 2002), where the host response may be better understood, our study demonstrates the necessity of incorporating what is known of the host response into an efficient and effective treatment plan.
Conclusion
Wildlife disease treatments that show promise in the lab still face significant hurdles before treatments can be applied successfully to wild populations (Langwig et al., 2015b). Both treatment efficacy in pathogen removal and the percent of the host population that can be treated given economic and logistic constraints contribute to these hurdles. However, beyond these direct considerations of development and deployment, our results clearly demonstrate the need to consider the impact of treatment on the ongoing dynamics between host defense mechanisms and disease dynamics before treatment-based interventions can truly be added to our arsenal of conservation tools.
Author Contributions
BM and NF conceived of the idea, NF performed the modeling, and BM, NF, and SG interpreted the results and wrote the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We would like to thank several colleagues for discussions relating to treatment and this manuscript: Winifred Frick, Kate Langwig, Julie Lockwood, Christopher Cornelison, Sybill Amelon, and Barrie Overton as well as the reviewers.
References
Bacher, P., Kniemeyer, O., Schönbrunn, A., Sawitzki, B., Assenmacher, M., Rietschel, E., et al. (2014). Antigen-specific expansion of human regulatory T cells as a major tolerance mechanism against mucosal fungi. Mucosal Immunol. 7, 916–928. doi: 10.1038/mi.2013.107
Barclay, R. M., Harder, L. D., Kunz, T., and Fenton, M. (2003). “Life histories of bats: life in the slow lane,” in Bat Ecology, eds T. H. Kunz and M. B. Fenton (Chicago, IL: University of Chicago Press), 209–253.
Becker, D. J., Streicker, D. G., and Altizer, S. (2015). Linking anthropogenic resources to wildlife–pathogen dynamics: a review and meta-analysis. Ecol. Lett. 18, 483–495. doi: 10.1111/ele.12428
Brasier, C. M., and Buck, K. W. (2001). Rapid evolutionary changes in a globally invading fungal pathogen (Dutch elm disease). Biol. Invas. 3, 223–233. doi: 10.1023/A:1015248819864
Brearley, G., Rhodes, J., Bradley, A., Baxter, G., Seabrook, L., Lunney, D., et al. (2013). Wildlife disease prevalence in human-modified landscapes. Biol. Rev. 88, 427–442. doi: 10.1111/brv.12009
Burge, C. A., Mark Eakin, C., Friedman, C. S., Froelich, B., Hershberger, P. K., Hofmann, E. E., et al. (2014). Climate change influences on marine infectious diseases: implications for management and society. Ann. Rev. Mar. Sci. 6, 249–277. doi: 10.1146/annurev-marine-010213-135029
Cheng, T. L., Mayberry, H., McGuire, L. P., Hoyt, J. R., Langwig, K. E., Nguyen, H., et al. (2016). Efficacy of a probiotic bacterium to treat bats affected by the disease white-nose syndrome. J. Appl. Ecol. doi: 10.1111/1365-2664.12757. [Epub ahead of print].
Cornelison, C. T., Gabriel, K. T., Barlament, C., and Crow, S. A. Jr. (2014). Inhibition of Pseudogymnoascus destructans growth from conidia and mycelial extension by bacterially produced volatile organic compounds. Mycopathologia 177, 1–10. doi: 10.1007/s11046-013-9716-2
Daszak, P., Cunningham, A. A., and Hyatt, A. D. (2000). Emerging infectious diseases of wildlife–threats to biodiversity and human health. Science 287, 443–449. doi: 10.1126/science.287.5452.443
Field, K. A., Johnson, J. S., Lilley, T. M., Reeder, S. M., Rogers, E. J., Behr, M. J., et al. (2015). The white-nose syndrome transcriptome: activation of anti-fungal host responses in wing tissue of hibernating little brown myotis. PLoS Pathog. 11:e1005168. doi: 10.1371/journal.ppat.1005168
Fisher, M. C., Henk, D. A., Briggs, C. J., Brownstein, J. S., Madoff, L. C., McCraw, S. L., et al. (2012). Emerging fungal threats to animal, plant and ecosystem health. Nature 484, 186–194. doi: 10.1038/nature10947
Frick, W. F., Pollock, J. F., Hicks, A. C., Langwig, K. E., Reynolds, D. S., Turner, G. G., et al. (2010a). An emerging disease causes regional population collapse of a common North American bat species. Science 329, 679–682. doi: 10.1126/science.1188594
Frick, W. F., Puechmaille, S. J., Hoyt, J. R., Nickel, B. A., Langwig, K. E., Foster, J. T., et al. (2015). Disease alters macroecological patterns of North American bats. Global Ecol. Biogeogr. 24, 741–749. doi: 10.1111/geb.12290
Frick, W. F., Reynolds, D. S., and Kunz, T. H. (2010b). Influence of climate and reproductive timing on demography of little brown myotis Myotis lucifugus. J. Anim. Ecol. 79, 128–136. doi: 10.1111/j.1365-2656.2009.01615.x
Garcia-Solache, M. A., and Casadevall, A. (2010). Global warming will bring new fungal diseases for mammals. mBio 1:e00061-10. doi: 10.1128/mBio.00061-10
Gonzalez, A., Ronce, O., Ferriere, R., and Hochberg, M. E. (2013). Evolutionary rescue: an emerging focus at the intersection between ecology and evolution. Philos. Trans. R. Soc. Lond. B Biol. Sci. 368:20120404. doi: 10.1098/rstb.2012.0404
Harvell, C. D., Mitchell, C. E., Ward, J. R., Altizer, S., Dobson, A. P., Ostfeld, R. S., et al. (2002). Climate warming and disease risks for terrestrial and marine biota. Science 296, 2158–2162. doi: 10.1126/science.1063699
Heard, M. J., Smith, K., and Ripp, K. J. (2011). Examining the evidence for chytridiomycosis in threatened amphibian species. PLoS ONE 6:e23150. doi: 10.1371/journal.pone.0023150
Hebart, H., Bollinger, C., Fisch, P., Sarfati, J., Meisner, C., Baur, M., et al. (2002). Analysis of T-cell responses to Aspergillus fumigatus antigens in healthy individuals and patients with hematologic malignancies. Blood 100, 4521–4528. doi: 10.1182/blood-2002-01-0265
Lai, Y., Di Nardo, A., Nakatsuji, T., Leichtle, A., Yang, Y., Cogen, A. L., et al. (2009). Commensal bacteria regulate Toll-like receptor 3–dependent inflammation after skin injury. Nat. Med. 15, 1377–1382. doi: 10.1038/nm.2062
Langwig, K. E., Frick, W. F., Reynolds, R., Parise, K. L., Drees, K. P., Hoyt, J. R., et al. (2015a). Host and pathogen ecology drive the seasonal dynamics of a fungal disease, white-nose syndrome. Proc. Biol. Sci. 282:20142335. doi: 10.1098/rspb.2014.2335
Langwig, K. E., Voyles, J., Wilber, M. Q., Frick, W. F., Murray, K. A., Bolker, B. M., et al. (2015b). Context-dependent conservation responses to emerging wildlife diseases. Front. Ecol. Environ. 13, 195–202. doi: 10.1890/140241
Lorch, J. M., Meteyer, C. U., Behr, M. J., Boyles, J. G., Cryan, P. M., Hicks, A. C., et al. (2011). Experimental infection of bats with Geomyces destructans causes white-nose syndrome. Nature 480, 376–378. doi: 10.1038/nature10590
Maslo, B., and Fefferman, N. H. (2015). A case study of bats and white-nose syndrome demonstrating how to model population viability with evolutionary effects. Conserv. Biol. 29, 1176–1185. doi: 10.1111/cobi.12485
Maslo, B., Valent, M., Gumbs, J. F., and Frick, W. F. (2015). Conservation implications of ameliorating survival of little brown bats with white-nose syndrome. Ecol. Appl. 25, 1832–1840. doi: 10.1890/14-2472.1
McCallum, H. (2012). Disease and the dynamics of extinction. Philos. Trans. R. Soc. Lond. B Biol. Sci. 367, 2828–2839. doi: 10.1098/rstb.2012.0224
Puechmaille, S. J., Wibbelt, G., Korn, V., Fuller, H., Forget, F., Mühldorfer, K., et al. (2011). Pan-European distribution of white-nose syndrome fungus (Geomyces destructans) not associated with mass mortality. PLoS ONE 6:e19167. doi: 10.1371/journal.pone.0019167
Pullin, A. S., Sutherland, W., Gardner, T., Kapos, V., and Fa, J. E. (2013). “Conservation priorities: identifying need, taking action and evaluating success,” in Key Topics in Conservation Biology 2, ed D. W. Macdonald (Oxdord: John Wiley and Sons), 3–22.
Raudabaugh, D. B., and Miller, A. N. (2015). Effect of Trans, trans-farnesol on Pseudogymnoascus destructans and several closely related species. Mycopathologia 180, 325–332. doi: 10.1007/s11046-015-9921-2
Rizzetto, L., De Filippo, C., Rivero, D., Riccadonna, S., Beltrame, L., and Cavalieri, D. (2013). Systems biology of host–mycobiota interactions: dissecting Dectin-1 and Dectin-2 signalling in immune cells with DC-ATLAS. Immunobiology 218, 1428–1437. doi: 10.1016/j.imbio.2013.07.002
Robert, V. A., and Casadevall, A. (2009). Vertebrate endothermy restricts most fungi as potential pathogens. J. Infect. Dis. 200, 1623–1626. doi: 10.1086/644642
Savage, A. E., Terrell, K. A., Gratwicke, B., Mattheus, N. M., Augustine, L., and Fleischer, R. C. (2016). Reduced immune function predicts disease susceptibility in frogs infected with a deadly fungal pathogen. Conserv. Physiol. 4:cow011. doi: 10.1093/conphys/cow011
Smith, K. F., Acevedo-Whitehouse, K., and Pedersen, A. B. (2009). The role of infectious diseases in biological conservation. Anim. Conserv. 12, 1–12. doi: 10.1111/j.1469-1795.2008.00228.x
Sutherland, K. P., Berry, B., Park, A., Kemp, D. W., Kemp, K. M., Lipp, E. K., et al. (2016). Shifting white pox aetiologies affecting Acropora palmata in the Florida Keys, 1994–2014. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 371:20150205. doi: 10.1098/rstb.2015.0205
Tetzlaff, S., Allender, M., Ravesi, M., Smith, J., and Kingsbury, B. (2015). First report of snake fungal disease from Michigan, USA involving Massasaugas, Sistrurus catenatus (Rafinesque 1818). Herpetol. Notes 8, 31–33.
Verant, M. L., Carol, M. U., Speakman, J. R., Cryan, P. M., Lorch, J. M., and Blehert, D. S. (2014). White-nose syndrome initiates a cascade of physiologic disturbances in the hibernating bat host. BMC Physiol. 14:10. doi: 10.1186/s12899-014-0010-4
Weil, E., and Rogers, C. S. (2011). “Coral reef diseases in the Atlantic-Caribbean,” in Coral Reefs: An Ecosystem in Transition, eds Z. Dubinsky and N. Stambler (New York, NY: Springer), 465–491. doi: 10.1007/978-94-007-0114-4_27
Whilde, J., Martindale, M. Q., and Duffy, D. J. (2017). Precision wildlife medicine: applications of the human-centred precision medicine revolution to species conservation. Glob. Chang. Biol. 23, 1792–1805. doi: 10.1111/gcb.13548
Williams, E. S., Miller, M. W., Kreeger, T. J., Kahn, R. H., and Thorne, E. T. (2002). Chronic wasting disease of deer and elk: a review with recommendations for management. J. Wildl. Manage. 66, 551–563. doi: 10.2307/3803123
Willis, C. K. R., Menzies, A. K., Boyles, J. G., and Wojciechowski, M. S. (2011). Evaporative water loss is a plausible explanation for mortality of bats from white-nose syndrome. Integr. Comp. Biol. 51, 364–373. doi: 10.1093/icb/icr076
Woodhams, D. C., Geiger, C. C., Reinert, L. K., Rollins-Smith, L. A., Lam, B., Harris, R. N., et al. (2012). Treatment of amphibians infected with chytrid fungus: learning from failed trials with itraconazole, antimicrobial peptides, bacteria, and heat therapy. Dis. Aquat. Org. 98, 11–25. doi: 10.3354/dao02429
Keywords: bat conservation, disease treatment modeling, fungal pathogen, wildlife disease, white nose syndrome, wildlife disease management
Citation: Maslo B, Gignoux-Wolfsohn SA and Fefferman NH (2017) Success of Wildlife Disease Treatment Depends on Host Immune Response. Front. Ecol. Evol. 5:28. doi: 10.3389/fevo.2017.00028
Received: 03 February 2017; Accepted: 23 March 2017;
Published: 11 April 2017.
Edited by:
Ben L. Phillips, University of Melbourne, AustraliaReviewed by:
Sergio Andrés Estay, Austral University of Chile, ChileEugene B. Postnikov, Kursk State University, Russia
Copyright © 2017 Maslo, Gignoux-Wolfsohn and Fefferman. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Brooke Maslo, brooke.maslo@rutgers.edu
 Brooke Maslo
Brooke Maslo Sarah A. Gignoux-Wolfsohn
Sarah A. Gignoux-Wolfsohn Nina H. Fefferman
Nina H. Fefferman