- 1The Pirbright Institute, Woking, UK
- 2Institute for Virology, Faculty of Medicine, University Medical Center, University of Freiburg, Freiburg, Germany
- 3Department of Microbiology and Immunology, University of Rochester Medical Center, Rochester, NY, USA
Interferon (IFN) responses, mediated by a myriad of IFN-stimulated genes (ISGs), are the most profound innate immune responses against viruses. Cumulatively, these IFN effectors establish a multilayered antiviral state to safeguard the host against invading viral pathogens. Considerable genetic and functional characterizations of mammalian IFNs and their effectors have been made, and our understanding on the avian IFNs has started to expand. Similar to mammalian counterparts, three types of IFNs have been genetically characterized in most avian species with available annotated genomes. Intriguingly, chickens are capable of mounting potent innate immune responses upon various stimuli in the absence of essential components of IFN pathways including retinoic acid-inducible gene I, IFN regulatory factor 3 (IRF3), and possibility IRF9. Understanding these unique properties of the chicken IFN system would propose valuable targets for the development of potential therapeutics for a broader range of viruses of both veterinary and zoonotic importance. This review outlines recent developments in the roles of avian IFNs and ISGs against viruses and highlights important areas of research toward our understanding of the antiviral functions of IFN effectors against viral infections in birds.
Introduction
For efficient replication and spread, viruses have to breach a potent and multilayered immune system in the host. Occasionally, either due to defects in host immune responses [e.g., complement system, interferons (IFNs), and adaptive immunity] or due to successful immune-antagonism, viruses overcome these antiviral mechanisms and replicate extensively in the host. This results in the engagement of diverse cascades of cellular signaling pathways (1). One of the most potent and essential events in this host–pathogen battle is the activation of the IFN pathways (1–3).
Three classes of nucleic acid receptors are associated with the activation of the IFN pathways. The first category of intracellular pattern recognition receptors (PRRs) is the family of retinoic acid-inducible gene I (RIG-I)-like helicases (RLH), which includes RIG-I, melanoma differentiation-associated gene 5 (MDA5), and laboratory of genetics and physiology 2 (LGP2) (3). A second class of PRRs is the family of toll-like receptors (TLR) including TLR3, TLR7, and TLR9, which senses extracellular, phagosomal, or endosomal pathogen-associated molecular patterns (1). The third category of PRRs is the family of DNA sensors, which include absent in melanoma 2 (AIM2) and cyclic GMP-AMP synthetase (cGAS) (4). Upon activation, these PRRs recruit downstream signaling molecules and result, directly or indirectly, in the activation of IFN regulatory factors 3 (IRF3) and 7 (IRF7), as well as activating protein 1 (AP-1) and nuclear factor kappa B (NF-κB) transcription factors (1–4). These are minimally essential events to initiate transcription of type I IFN genes and establishment of an antiviral state by expressing hundreds of IFN-stimulated genes (ISGs) (1, 5) in infected cells.
Extensive structural and functional models have been proposed on the plasticity and dynamics of nucleic acid sensing by intracellular PRRs and on the mechanisms of IFN-induced antiviral states in mammals (1–4, 6). For a detailed description of IFN induction and other innate immune responses in mammals against viruses of diverse genetic backgrounds, we refer to other in-depth reviews (1, 2, 4, 5, 7, 8).
In this article, we offer a review of the IFN pathways and transcriptional activation of ISGs in different avian species. First, we provide an overview of the chicken IFN pathways and highlight areas that differ from mammalian IFN induction and signaling. Then, we convey a comparative genetic and genomic analysis of characterized components of IFN systems among different avian species. We conclude with a description of currently studied antiviral effectors, their implications for avian diseases, and future perspectives.
The Chicken IFN Pathways: Sensing of Viral Nucleic Acids
RLH-Mediated IFN Induction
The principles of mammalian IFN pathways (exemplified by humans) are in general transferable to chickens. However, there are considerable evolutionary divergences in some of the key elements of the chicken IFN responses to avian viruses if compared to their mammalian counterparts. In mammals, RIG-I primarily senses 5′-triphosphorylated blunt-ended or double-stranded RNA (dsRNA) produced during RNA virus infections. On the other hand, MDA5 can be activated by long dsRNA, whereas LGP2, which differs from RIG-I and MDA5 in lacking caspase activation and recruitment domain domain, can positively regulate MDA5 and negatively regulate RIG-I signaling (9, 10). One of the most striking features of chickens and other members of the order Galliformes (e.g., turkeys) is the absence of RIG-I (11). Despite the absence of this key PRR, chickens respond to highly pathogenic avian influenza virus (HPAIVs) and mount potent type I IFN responses, probably due to cooperative actions of MDA5 and LGP2 (10, 12, 13) (Figure 1). Additionally, unlike the mammalian MDA5, which senses only long dsRNA, it appears that chicken MDA5 can also sense short dsRNA implying that chicken MDA5 may compensate, to some extent, the function of RIG-I in chickens (13). Recently, Uchikawa et al. have resolved the structures of dsRNA-bound chicken LGP2 and MDA5 and revealed the plasticity of nucleic acid sensing by these RLH (10). It was shown that chicken LGP2 carries two properties of RLH: an MDA5-like helicase domain and a RIG-I-like C-terminal domain. Chicken LGP2, similar to human RIG-I, is an “end binder,” whereas chicken MDA5 is a “stem binder” of dsRNA (Figure 1). Based on structural (10) and functional studies (12, 13), it is likely that chicken LGP2-mediated enhancement of MDA5 sensing of dsRNA is dependent on RNA binding. However, it remains to be demonstrated if the mechanisms of LGP2-mediated enhancement of MDA5 signaling are similar to its mammalian counterparts or if the absence of RIG-I in chickens can contribute in the dynamics of cooperative nucleic acid sensing in chickens.
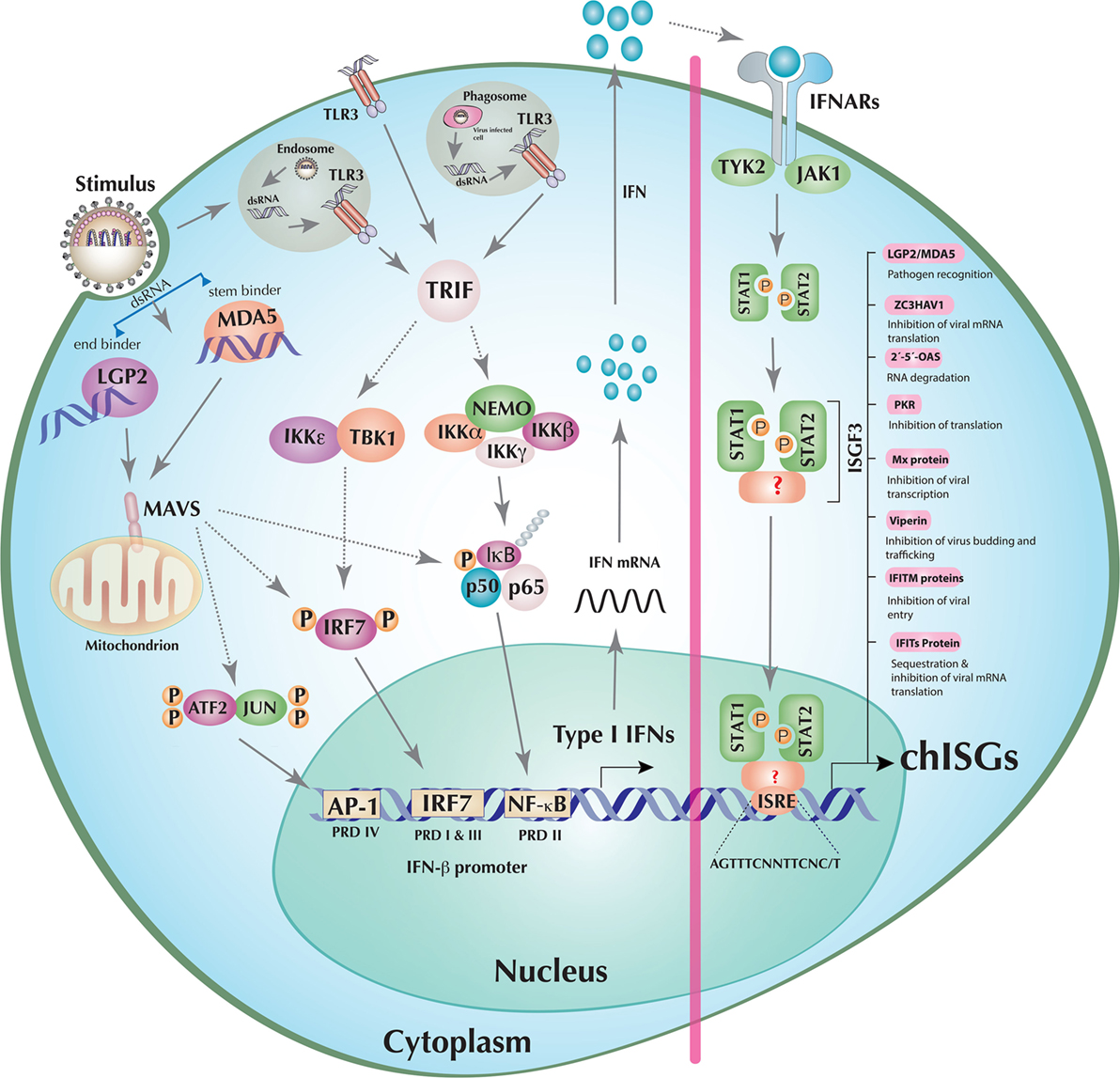
Figure 1. Induction of interferons (IFNs) and establishment of an antiviral state in a model chicken cell. The double-stranded RNA (dsRNA), detected by either chicken retinoic acid-inducible gene I (RIG-I)-like helicase (RLH) [melanoma differentiation-associated gene 5 (MDA5) or laboratory of genetics and physiology 2 (LGP2) individually or in cooperation] or toll-like receptor (TLR)3 (endosomal, phagosomal, or transmembrane) initiates downstream signaling mediated through mitochondrial antiviral-signaling protein (MAVS) or TRIF, respectively. These adaptor molecules then activate the transcription factors IFN regulatory factor (IRF)7, nuclear factor kappa B (NF-κB), and activating protein 1 (AP-1) (ATF2/JUN) by orchestrating the assembly of multi-protein complexes. Once activated, IRF7, NF-κB, and AP-1 translocate to the nucleus where they stimulate the transcription of, among others, type I IFNs (e.g., IFN-β). The transcribed, translated, and secreted type I IFNs initiate the JAK–STAT pathway by both autocrine (depicted in the figure) and paracrine signaling through cognate type I IFN receptor recognition. Activated JAK–STAT leads to the phosphorylation of STAT1 and STAT2 molecules, which (together with factors that are currently unknown in chicken) results in the formation of the IFN-stimulated gene factor 3 (ISGF3) transcription factor complex. This multifunctional transcription factor then scans and recognizes unique IFN-stimulated response element (ISRE) sequences to initiate the transcription of hundreds of chicken IFN-stimulated genes (chISGs), which subsequently establish the antiviral state against the invading viruses. Few examples of IFN-stimulated genes (ISGs) along with a summarized description of their functions are enlisted in the right panel of the figure. Abbreviations used in the figure and are not described in the main text are as follows: IκB kinase (IKK) epsilon (IKKε), alpha (IKKα), beta (IKKβ), and gamma (IKKγ); NF-κB essential modulator (NEMO); TANK-binding kinase 1 (TBK1); inhibitors of NF-κB (IκB), NF-κB subunits p50 and p65; activating transcription factor 2 (ATF2); tyrosine kinase 2 (TYK2); Janus kinase 1 (JAK1); signal transducer and activator of transcription 1 (STAT1), and STAT2. “P” represents the phosphorylation state of the protein, and dotted lines indicate the involvement of multiple intermediary steps.
It has been hypothesized that the lack of RIG-I makes chickens highly susceptible to RNA viruses, and therefore chickens continue to play a central role in the emergence of zoonotic influenza viruses (14). However, more research is still required to support this generally accepted concept. Although MDA5 and LGP2 seem to be sufficient to induce a potent activation of the type I IFN pathway, ectopic expression of duck RIG-I in chicken cells potentiated the downstream signaling pathway, including increased induction of several ISGs such as myxovirus-resistance protein (Mx), protein kinase R (PKR), IFN-induced protein with tetratricopeptide repeats 5 (IFIT5), or 2′-5′-oligoadenylate synthetase (2′-5′-OAS) (14, 15). These studies indicate that chickens have acquired mechanisms to compensate the deficiency of the RIG-I signaling molecule; however, it is not possible to assess the outcome of nucleic acid sensing in chickens as it would have been in the presence of endogenous RIG-I. Nevertheless, chickens are one of the most successfully domesticated animal species and are immunologically competent in mounting an effective antiviral type I IFN state against diverse stimuli.
TLR-Mediated IFN Induction
Toll-like receptors are type I transmembrane proteins and have a highly conserved architecture in a variety of species, including insects, fish, amphibians, birds, and mammals (16). Comparative biological approaches revealed that chicken TLRs carry unique properties regarding ligand specificity, formation of TLR receptor complexes, and activation of signaling pathways (17). At least 10 different TLR members (TLR1–10) have been identified in humans (16). Chickens have been shown to have two TLR2 isoforms (chTLR2 types 1 and 2), two TLR1/6/10 orthologs, and single genes for TLR3, TLR4, TLR5, and TLR7. Interestingly, chickens do not possess the viral DNA sensor TLR9. However, TLR-mediated DNA sensing is mediated by a functional ortholog TLR21, which is absent in humans (16). Additionally, it has been proposed that chicken TLR8 is non-functional and that the chicken genome encodes for an additional TLR gene, TLR15, which requires protease-cleavage for activation (18). Beside genomic variations, functional differences exist in the mechanism of TLR-mediated signal induction in chickens. In contrast to humans, lipopolysaccharides failed to stimulate the TLR4–TRAM–TRIF pathway in chicken cells (19). Among all mammalian TLRs, TLR3, TLR7/8, and TLR9 are known to sense viral dsRNA, ssRNA, and DNA molecules, respectively. Since in chicken TLR8 is non-functional and TLR9 is absent, only TLR3 and TLR7 are involved in the recognition of RNA viruses. All TLR family members, with the notable exception of TLR3, signal via myeloid differentiation primary response 88 (Myd88). TLR3 recruits TIR-domain-containing adapter-inducing IFN-β (TRIF) through transmembrane, phagosomal, or endosomal compartments (Figure 1). Both modes of TLR-dependent signal induction culminate in the activation of the transcription factors required for the transcription of type I IFNs.
DNA Sensors-Mediated IFN Induction
In addition to TLR9-mediated DNA sensing in mammals, cytosolic DNA, which can be either non-self DNA or results from gross nuclear/mitochondrial damage, can elicit type I IFN responses in mammals (9). Currently two major cytosolic sensors of DNA have been characterized: the PYHIN family member AIM2 and cGAS. Additionally, several proteins have been recognized as DNA receptors, including Z DNA binding protein 1 (ZBP1/DAI), the helicase DDX41, and IFI16, another member of the PYHIN/HIN-200 family (20, 21). Downstream of these DNA sensors, the stimulator of IFN genes (STING) acts as an adapter and stimulates type I IFN production through the activation of IRF3 and NF-κB transcription factors (9). Although DNA sensing in chickens has not yet been explored in greater detail, genetic analysis indicate that the AIM2 gene has been lost independently in several animals, including bats and chickens (22). Even in the latest Ensembl release of the chicken genome, ZBP1 and IFI16 were not identified, suggesting fundamental differences in DNA sensing mechanisms in chickens. However, it has been shown recently that chicken STING can actively sense DNA and in cooperation with the mitochondrial antiviral-signaling protein induces type I IFN responses independent of RIG-I, interfering with the replication of RNA viruses (23). Interestingly, STING-mediated type I IFN induction was synergistically supported by RLHs in chickens (23). This warrants future investigations to understand the molecular mechanisms underlining DNA sensing in chickens.
Transcriptional Activation of IFNs
Signals initiated by the sensing of viral nucleic acids by RLHs, TLRs, or DNA sensors lead to the activation of at least three transcription factors (AP-1, IRF3, and NF-κB) in the mammalian type I IFN enhanceosome (1). There is scarcity in our current understanding of the mechanism and structure of the chicken IFN enhanceosome. Comparative genomics analysis indicates that chickens are IRF3 deficient (detailed below). Currently, it is not known if the presence of functional IRF7 in chickens compensates for the IRF3 deficiency. Components of AP-I and NF-κB transcription factors are encoded in the chicken genome, and it is likely that these signaling cascades are functionally similar to mammals. Thus, a direct functional comparison may be plausible. While inactive, NF-κB, IRF3/IRF7 (in mammals and IRF7 in chicken), and AP-1 remain in the cytoplasm; however, upon stimulation (e.g., nucleic acids) these transcription factors get activated and subsequently translocated to the nucleus of viral-infected cells by unique mechanisms (1). The activation signals result in phosphorylation of IRF7. Conformational changes caused by this post-translational modification result in IRF7 dimerization and exposure of the nuclear localization signal (NLS) (1). This NLS mediates the nuclear translocation of IRF7 (1, 24). The inhibitor of NF-κB (IκB) retains NF-κB molecules in the cytoplasm. However, upon activation by phosphorylation, IκB undergoes ubiquitination and proteasomal degradation. Degradation of IκB exposes the NLS of NF-κB, which leads to its nuclear translocation (7). Phosphorylation of c-jun and activating transcription factor 2, two heterodimeric components of AP-1, also causes nuclear translocation (1). In the nucleus, these three transcription factors assemble in a cooperative manner to build a type I IFN enhanceosome, which binds to its respective positive regulatory domains (PRDs). IRF7, NF-κB, and AP-1 bind to PRD I/III, PRD II, and PRD IV, respectively, where they induce the transcription of type I IFNs and pro-inflammatory cytokines (TNF, IL-6, IL-1β, etc.) (25) (Figure 1). These type I IFNs lead to transcriptional activation of several hundreds ISGs to mount an antiviral state in the host (detailed below).
Comparative Genomics and Evolution by Gene Loss
Even in the updated version of chicken Ensembl (Ensembl release 85—July 2016, accessed on September 11, 2016), it appears that chickens lack IRF3 and IRF9 (depicted in Figure 1), which are essential components of the type I IFN system in mammals (1). Lately, there have been substantial improvements in the genetic analysis and functional characterization of the avian type I IFN pathway, particularly in chicken. However, the annotation of the chicken genome is not yet completed, leaving open questions on the presence or absence of the mammalian homologs in avian species. Improved annotation of chicken and other avian genomes is required to unambiguously declare the presence or absence of a particular gene in the future. This fact can be exemplified by a recent analysis of IRF3/7 in the chicken genome. The first identified and characterized member in the chicken IRF family (named cIRF3) was classified as IRF3 based on its sequence and overall functional conservation with corresponding IRF3 in other species (26). Availability of updated annotation of chicken genome in the Ensembl has filled the gaps in the chicken chromosome 5, which encodes for the IRF3/7 genes and allowed to reevaluate the IRF locus in the chicken. Based on the analysis of gene loci in different species including human, mouse, dog, and fish (Figure 2A) and previous reports (25, 27), it is convincing that the formerly reported cIRF3 is actually IRF7. Furthermore, genetic clustering and sequence divergence analysis indicate that the chicken IRF7 clusters closely with IRF7 of human, mouse, and cattle compared to the IRF3 of corresponding species (Figure 2B). Therefore, it is suggested to use the term chicken IRF7 instead of cIRF3 to avoid any misunderstanding in the functional nomenclature between these two transcription factors.
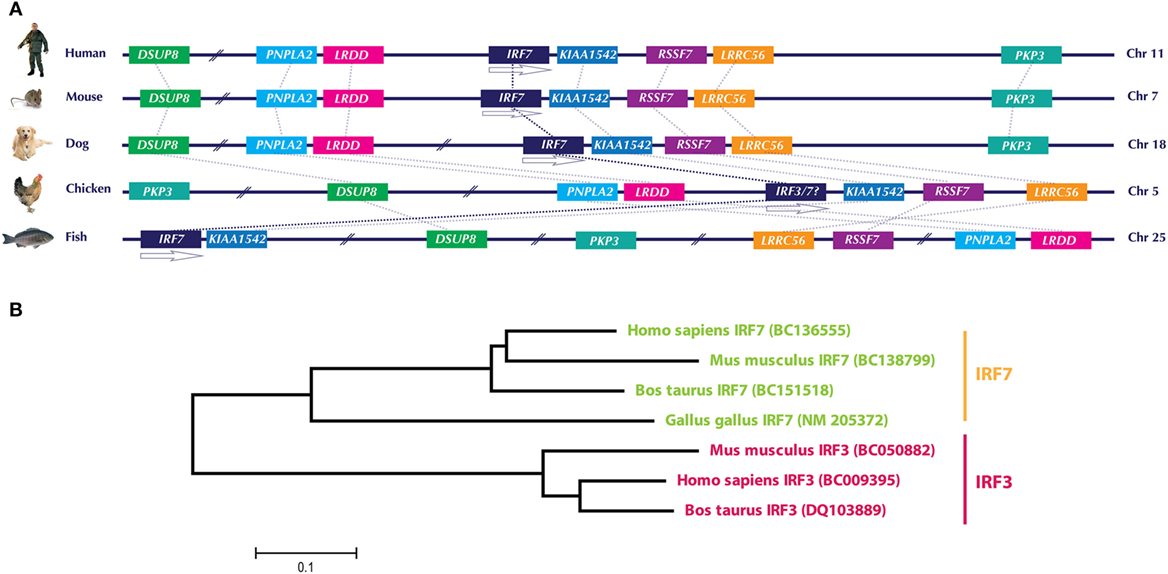
Figure 2. (A) Genomic architecture along with relative loci around the IRF7 gene in human, mouse, dog, chicken, and fish. The IRF7 genes in the compared species are flanked upstream with LRDD gene and downstream with KIAA1542 and RSSF7 genes. Direct comparison of previously identified chicken IRF3 with these species indicates that this gene is in fact IRF7. (B) Phylogenetic analysis of IRF3 and IRF7 genes in different species. Based on the clustering patterns and sequence homologies, the gene previously identified as “chicken IRF3” clustered closer to IRF7 of other mammals compared to mammalian IRF3. It is therefore proposed to rename “chicken IRF3” to “chicken IRF7.” Gene abbreviations used in the figure are dual specificity phosphatase 8 (DSUP8), patatin-like phospholipase domain containing 2 (PNPLA2), leucine-rich repeats and death domain containing (LRDD), interferon regulatory factor 7 (IRF7), CTD-binding SR-like protein rA9 (KIAA1542); Ras association (RalGDS/AF-6) domain family 7 (RSSF7); leucine-rich repeat containing 56 (LRRC56); plakophilin 3 (PKP3).
Similar to IRF3/IRF7, the currently annotated chicken IRF9 sequence is both genetically (Figure 3A) and phylogenomically (Figure 3B) similar to IRF10 in dog and fish. Comparison of the gene orientation and architecture between species in which IRF10 is detected (dog and fish) and species in which IRF10 is lacking (human and mice) provides direct evidence that these crucial elements of the IFN pathways are currently incorrectly annotated. In addition, our analysis on global IRF family members confirms that the chicken genome lacks any significant sequence identity to the mammalian IRF9 orthologs. It remains to be explored how chickens still manage to efficiently trigger the production of ISGs without the need of IRF9 to constitute a functional type I IFN-stimulated gene factor 3 (ISGF3) complex. However, it is plausible that factor(s) other than IRF9 are involved in the formation of an active ISGF3 complex in chickens. Since type II IFNs-mediate induction of ISGs is IRF9-independent, it may be possible that under virus infection the gamma-activated sequence (GAS) promoter may overwhelm the overall induction of ISGs compared to type I and III IFN-induced expression of ISGs.
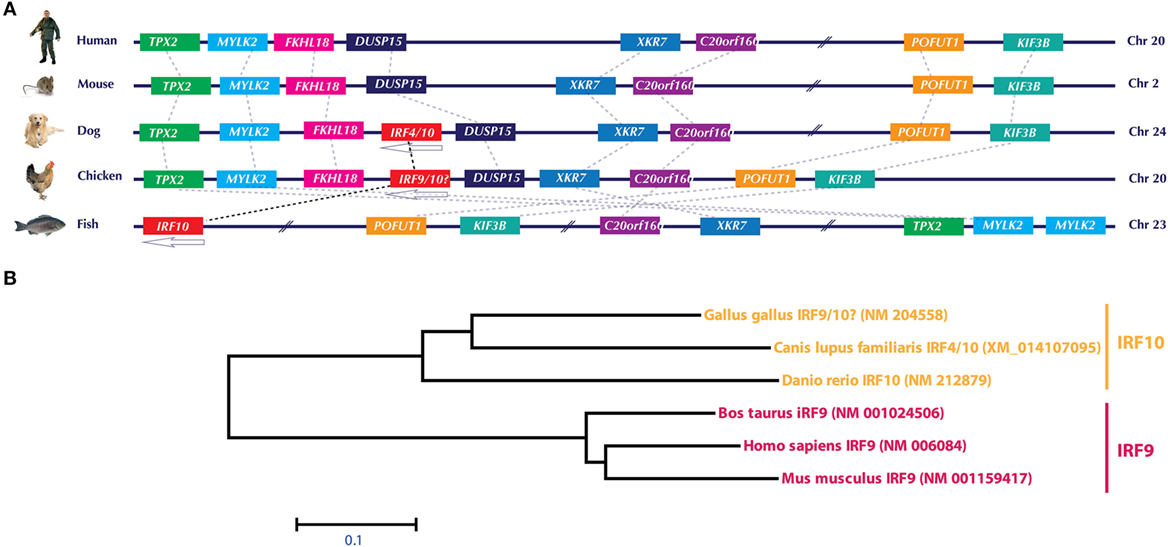
Figure 3. (A) Genomic architecture of interferon regulatory factor (IRF)10 loci and phylogenetic analysis of IRF9 and IRF10 in human, mouse, dog, chicken, and fish. Upstream and downstream genes architecture in the IRF10 of chicken, dog, and fish indicate that this locus is similar to the corresponding locus in human and mice, which lack IRF10. Based on this and phylogenetic analysis (B), it is evident that the currently annotated chicken IRF9 is in fact an ortholog of IRF10. Gene abbreviations used in the figure are microtubule-associated protein homolog (Xenopus laevis) (TPX2); myosin, light polypeptide kinase 2, skeletal muscle (MYLK2); forkhead-like 18 (Drosophila) (FLKHL18); dual specificity phosphatase-like 15 (DUSP15); X Kell blood group precursor-related family member 7 homolog (XKR7); chromosome 20 open reading frame 160 (C20orf160); protein O-fucosyltransferase 1 (POFUT1); kinesin family member 3B (KIF3B).
From these examples, it is clear that our understanding of the avian genome is still insufficient to accurately annotate the newly identified genes. Efforts have recently become intense through the avian consortium to not only characterize the genetics of endangered and newly sequenced bird species but also to improve the annotation of the existing genome drafts of avian species, especially chicken and duck. As a result of this, a bunch of genome sequences from more than 40 avian species was published recently (28), providing a valuable source for gene mapping. These resources would certainly advance our understanding in exploring genes, which are conserved across avian species, and to confirm existing genes. A special database (AvianBase) has been established to facilitate comparative genomics and immunogenetics in avian species (29).
Beside the fact that genes are incorrectly annotated and important genomic loci are not characterized in the avian species, it is likely that birds have evolutionary lost some genes during their domestication and subsequent division into required phenotypes (egg-laying versus meat-producing) (30). It requires extensive genetic and genomic investigations to confirm gene loss in the evolutionary process of avian species and to identify a minimum number of genes that can be readily lost from avian genomes without compromising the survivability. Although several models can be proposed, loss of genes due to “gene function bias” appears to be operative in chicken and other avian species (31). Gene function bias refers to the gene loss that is preferentially evident in a specific functional category; gene loss in gene ontology category of “immune responses” is highly probable in mammals compared to other vertebrates (32). A similar scenario can be applied to the gene loss in innate immune signaling pathways compared to other gene ontology categories in avian species mainly due to dispensable functional constraints. In this context, different type I IFN-induced proteins with tetratricopeptide repeats (IFITs), including IFIT1, IFIT2, IFIT3, and IFIT5 (33), have been described to play essential roles in nucleic acid sensing, antiviral responses, and protein translation in humans. All these functions of IFIT proteins are redundant, and thus the protein family is likely under selection constraints in chicken where only one IFIT protein (IFIT5) has been identified compared to four in human and mice (34). In summary, understanding the mechanisms and impacts of gene loss would reveal crucial evolutionary aspects of animal domestication and may highlight unexplored ways that could be exploited both for antiviral therapy and disease control.
Evolution and Nomenclature of Avian IFNs
Phylodynamic analysis of homology-based coding sequences of all three types of IFNs (I, II, and III) indicates that these evolutionary IFN classes are only distantly related and lack apparent sequence homology among each other (Figure 4A). However, type I and II IFNs appear to be more closely related to each other compared to type III IFN, despite the fact that type I and III share functional and signaling homologies.
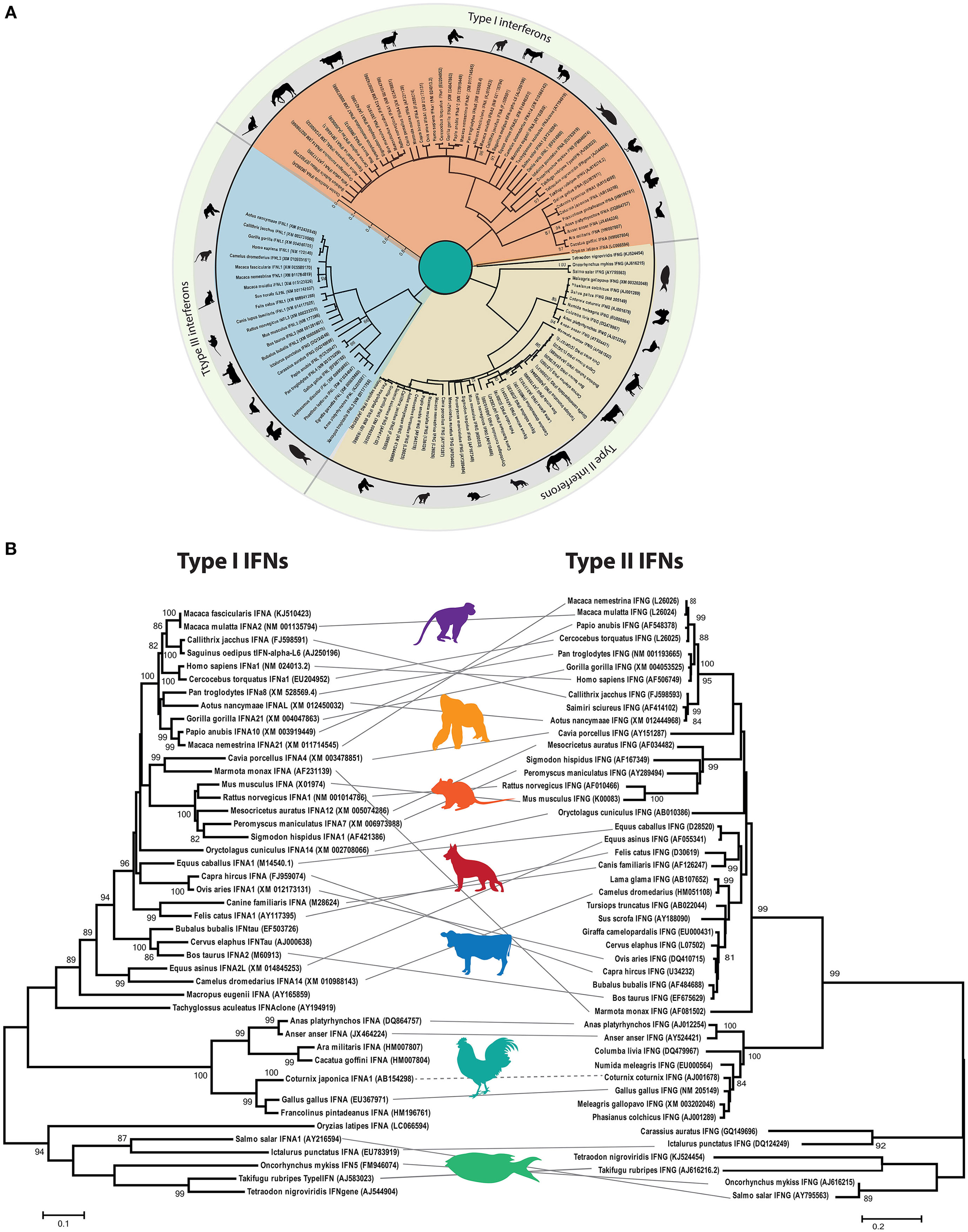
Figure 4. Phylogenetic analysis of interferon (IFN) genes of mammals (including rodents, primates, and domestic animals), avian, and fish species. (A) The open reading frames of type I, II, and III IFN genes were manually extracted from public databases and were aligned in BioEdit. The phylogenetic tree was constructed by MEGA6 software using the Kimura-2 model with 1,000 bootstrap replicates. The three types of IFNs clustered distantly and were labeled according to their clustering patterns. Approximate branching position was marked with a representative animal in the class. Only bootstrap values higher than 50 are shown. (B) Parallel comparison of type I and type II IFNs. Type III IFNs have been identified in limited numbers of species, and thus direct comparison was avoided. Clustering pattern of type I IFNs were linked to the type II IFN gene of the corresponding species for comparison purposes, and a representative animal image was shown to illustrate the clustering pattern.
Although chicken IFNs have functional homologies with their mammalian counterparts, gene duplication of each IFN subtype varies markedly among different animal species. In all birds investigated so far, type II and type III IFNs exist as a single gene each (35), whereas in mammals two to four copies of type III IFNs have been identified (35). Compared to fish, where generally only a single type I IFN homolog is detected, 3 to 10 type I IFN copies have been identified on the sex-determining Z chromosome of avian species (36–39). The maintenance of type II IFNs in avian and mammalian speciation indicates their constant function and evolutionary pressures. In both chicken and mammalian genomes, the functional transcript of the single type II IFN gene is encoded by four exons, and the gene architecture resembles that of IL-10-like cytokines.
Direct and parallel comparison of clustering patterns of type I and type II IFNs indicates the divergence of IFN-alpha (IFN-α) and IFN-gamma (IFN-γ) across mammals, rodents, primates, fish, and avian species (Figure 4B). It is evident that chicken IFNs and IFN genes of other vertebrates included in this evolutionary tree cluster distinctly from the fish IFNs. However, both fish and chicken type I and type II IFNs formed separate clades with markedly high resolution (bootstrap value of >90%). These clustering patterns may support the evolutionary and structural architecture of at least type I IFNs in different vertebrates, where fish encodes for five exons compared to single exon in birds and mammals. This is postulated to be due to the retrotransposition events in which four exons were lost between divergence of tetrapods and radiation of amniote lineages (37).
Consistent with vertebrate evolution, there are insufficient relationships between type I and type II IFNs in avian and mammalian species (Figures 4A,B). Thus, it is concluded that mammalian and avian type I IFNs evolved independently by gene duplication of a progenitor after segregation of mammals and birds (38, 39). Therefore, the avian type I IFNs are no true orthologs of their mammalian counterparts, and the nomenclature used for mammalian type I IFNs is strictly not appropriate for avian species. This is further supported by the level of genetic and functional differences between mammalian and avian type I IFNs (detailed below).
Avian IFNs
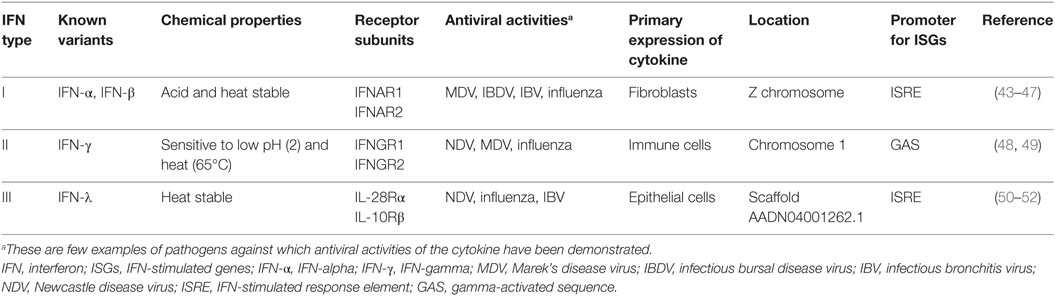
Based on their receptor specificity, sequence homology, and nature of ISG induction, IFNs are divided into those that bind IFNαR1 and IFNαR2 (type I IFNs), those that interact with receptors complexes of IFNγR1 and IFNγR2 (type II IFNs), and those that interact with heterodimeric receptor complex of IL-28Rα and IL-10Rβ (type III IFNs or IL-28/29). Our understanding of the avian IFN pathways is gradually increasing, and recently several significant contributions have been made to characterize existing genes (40–42) and previously identified IFNs, especially in chicken. In the following sections, our current understanding on chicken IFNs and comparative genomics in other avian species will be discussed. Several known features of chicken IFNs are summarized in Table 1.
Avian Type I IFNs
In contrast to the numerous members of type I IFNs in mammals (IFN-α, IFN-β, IFN-ε, IFN-κ, IFN-ω, IFN-δ, and IFN-τ), so far only two serologically distinct, intron-less, acid and heat stable type I IFNs (IFN-α and IFN-β sharing nt homology of 57%) have been identified in avian species on the short arm of Z (sex) chromosome (53). Unlike IFN-β, which is encoded only by a single gene copy, chicken IFN-α exists as a family of several genes (Table 1). Although there is low overall amino acid identity between avian and mammalian IFN-α protein sequences (24%), a core region in the chicken IFN-α carries four of six conserved cysteine residues, an α-helix and a high sequence identity (80%) compared to mammalian IFN-α protein (Figure 5A).
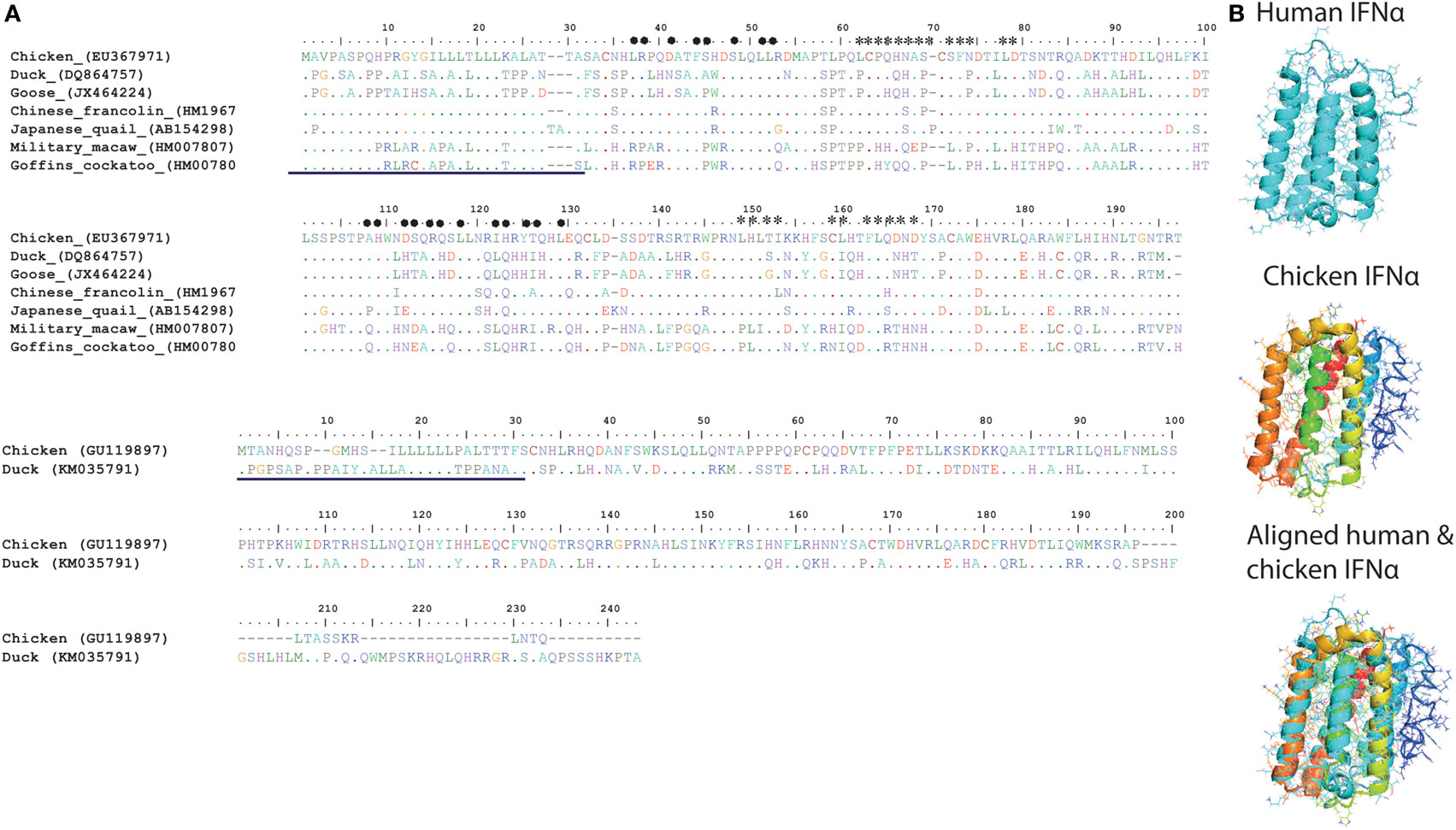
Figure 5. (A) Structural and amino acid sequence homologies between type I interferons (IFNs) in different avian species. Alignment and sequence homology of avian IFN-alpha (IFN-α) [(A), top panel] and avian IFN-beta (IFN-β) [(A), bottom panel] amino acid sequences. Putative sites for IFN-α binding to IFNAR1 are marked with heart symbol, whereas the sites that are important for binding to IFNAR2 are marked with star sign. In comparison to mammals, sites required for interaction of IFNs with IFNARs are more variable among avian species (54). The previously reported signal peptides are underlined in both IFN-α (top) and IFN-β (bottom) sequences (54). (B) A modeled cartoon structure of human and chicken IFN-α. IFN-α protein structures were predicted using I-TASSER online tool and were annotated and aligned in MacPyMOL. Similar to human IFN-α (PBD ID: 1ITF), chicken IFN-α carries five helices and is structurally similar to human IFN-α. Direct structure comparison between human (mammalian) and chicken (avian) IFN-α proteins indicate that the chicken IFN-α protein carries five alpha-helices, which are considered crucial for the functionality of type I IFNs in mammals.
Moreover, recombinant goose IFN-α has been shown to carry partial cross-species antiviral properties (55, 56). These results indicate that type I IFNs have attained certain levels of functional flexibilities (56). Nevertheless, all type I IFNs are known to be involved in inducing an antiviral state, inhibiting cell proliferation, modulating cell fate, and mediating cell differentiation and migration (57). To accomplish the primary function of IFNs, it is essential for these cytokines to bind to their respective receptors. Receptors for type I IFNs (IFNAR1 and IFNAR2) have been sequenced in chicken (58); however, little information is available about their functional domains and their crucial roles in type I IFN signaling.
Among avian type I IFNs, most of the research has been conducted on chickens, and IFN-α has been identified and more extensively characterized than IFN-β in different avian species (59) (Figures 5A,B, respectively). Chicken IFN-α and IFN-β genes were first identified from a cDNA library of aged chicken embryo cells, and subsequent analysis indicated the functionally and evolutionarily conserved properties compared to mammalian type I IFNs (60). Several recent studies have mapped the expression dynamics of chicken type I IFNs triggered by different stimuli (60–63). Collectively, type I IFNs (especially IFN-α) are potent antiviral agents and can ameliorate viral infections including Marek’s disease virus (MDV), infectious bursal disease virus (IBDV), infectious bronchitis virus (IBV), and HPAIV in different avian species (43–47). These antiviral properties of type I IFNs are identified not only in vitro but also in ovo and in vivo (43, 47).
Antiviral properties of type I IFNs are essentially mediated by the induction of ISGs. Both chicken IFN-α and IFN-β bind to the same IFN receptors (IFNAR1 and IFNAR2). However, it has been recently found that IFN-α and IFN-β differentially regulate ISGs in chickens (62). The antiviral state induced by chicken IFN-α was observed to be significantly more potent than that induced by chicken IFN-β, although both share genetic and structural similarities (64). These differential effects can be explained by differential binding affinity of IFN-α and IFN-β for the IFNAR1 and IFNAR2 (44). This hypothesis is further supported by a recent ontological study on the development of the chicken type I IFN system in which a markedly stronger upregulation of IFNAR1 as compared to IFNAR2 was observed during embryonic development in chicken lung and spleen cells (65). Since IFN-α and IFN-β differentially regulate the transcriptional activation of ISGs, it is imperative to consider that 5′ upstream regions of the chicken IFN-α genes lack NF-κB-binding sites and carry several binding sites for IRF members in their promoters regions (64). Moreover, observed differences in the ISGs induced by chicken IFN-α and IFN-β could be due to intrinsic functional components of the cell lines under investigation. For instance, type I IFNs induces TLR3 upregulation in the chicken fibroblasts cell line DF-1, whereas this induction was not observed in the chicken macrophages cell line HD11 (66). It cannot be excluded that constitutively primed cells may respond better to IFN-α compared to IFN-β, as has been observed in human lymphocytes that produce IFN-α, without the need to produce IFN-β, by viral infections (67). In conclusion, differential regulation of type I IFN-induced ISG signaling can be multifactorial and represents an interesting area for future investigations on the avian innate immunity.
Type I IFNs, especially IFN-α, have been characterized and assessed for their antiviral activities against IFN-sensitive viruses in various additional avian species. The duck type I IFNs were first detected in duck embryo fibroblasts (DEFs) after infection with high doses of reovirus serotype 3 (strain Dearing). Exogenous expression of this IFN blocked the release of avian RNA tumor virus particles in B77 virus-transformed DEFs (68) and showed antiviral effects for chronic hepatitis B virus infections (69). Recently, IFN-α has been identified and expressed in cells from the red-crowned crane (70), and an initial bioassay indicated its antiviral activities against vesicular stomatitis virus (VSV) in heterologous chicken fibroblasts. IFN-α has also been cloned from geese and turkeys, and initial functional insights including the antiviral actions have been determined (55, 71).
The results obtained so far on avian type I IFNs indicate that these cytokines are functionally, structurally (Figure 5B), and evolutionary related to mammalian IFNs and may have originated from common ancestor genes. However, extensive studies are required to identify other homologs of type I IFNs in all avian species, their mechanisms of action, how they exert individual and cumulative antiviral effects, and their potential for cross-species reactivity.
Avian Type II IFNs
Interferon-gamma is the only member of type II IFN in birds and mammals and serves as a bridge between innate and adaptive immunity. IFN-γ plays a crucial role in regulating the maturation and differentiation process of several immune cells and activates T helper 1-type immune responses (3). Due to these unique properties, significant research has been conducted to map the antiviral potential and mechanistic effects of IFN-γ in chicken, and considerable information is also available for other avian species. Direct gene comparison and evolutionary analysis of avian IFN-γ genes clearly demonstrate the significant identity both at the genome architecture and at the core functional transmembrane domain levels (Figures 6A,B). Receptors for type II IFNs have been identified and genetically characterized in chicken (72, 73). It is interesting to observe that unlike IFN-γ receptor β-chain (IFNGR2), the IFN-γ receptor α-chain (IFNGR1) of chicken has a 110 amino acid domain of a fibronectin type III (59). The LPKS and YDKPH motifs in the intracellular domain, required for the interaction with Janus kinase 1 (JAK1) and signal transducer and activator of transcription 1 (STAT1), were found to be conserved between avian and mammalian IFNGR1 (59). From two studies conducted by the same group, it was found that chicken IFNGR1 was highly expressed in spleen, thymus, peripheral blood lymphocytes (PBLs), cecal tonsil lung, and liver, whereas chicken IFNGR2 was highly expressed in spleen, thymus, PBLs, cecal tonsil, and muscle (72, 73). Beside these fundamental investigations, our current understanding is limited to the nature and genetics of IFNGRs in avian species, which warrants extensive future research to underpin the mechanisms of the IFN-γ-induced antiviral states.

Figure 6. Structural and amino acid sequence homologies between type II and type III interferons (IFNs) in different avian species. (A) Protein sequence alignment of avian type II IFN (IFN-γ). (B) Predicted structure of chicken IFN-γ. (C) Protein sequence alignment of avian type III IFN (IFN-λ). (D) Predicted structure of chicken IFN-λ. Sequence alignments show that type II and type III IFNs are significantly conserved among avian species and may indicate interspecies cross-reactivity. Previously identified or predicted signal peptides are underlined in both IFN-γ (A) and IFN-λ (C) sequence alignments (54). These structures were predicted using I-TASSER online tool and were annotated using MacPyMOL. Structurally, these IFNs are well aligned with that of human IFNs (data not shown).
Chicken IFN-γ was first amplified from a cDNA expression library generated from a T cell line (CC8.1h) in 1995 (74). Chicken IFN-γ is encoded by a single gene located on the chromosome 1 and shares >30% amino acid homology with mammalian IFN-γ genes (74). Genetic and functional studies indicated its actions to be conserved as compared to mammalian IFN-γ-proteins (74). Unlike type I IFNs, IFN-γ is sensitive to low pH (2) and heat (65°C) (74). Several studies demonstrate that small interfering RNA mediated gene silencing of the IFN-γ to ascertain its antiviral effects (75, 76). Likewise, recent studies have clearly defined the antiviral role of IFN-γ and its adjuvant properties against viruses of diverse genetic nature including Newcastle disease virus (NDV), MDV, and influenza viruses (77–79). Similar to its mammalian counterparts, chicken IFN-γ also induces MHC class I and class II molecules and mediates the production of nitric oxide, which is an important inhibitory mechanism for viruses (80). These studies have collectively highlighted the potential and emerging roles of chicken IFN-γ in vaccine-conferred antiviral immunity.
After the initial identification of duck IFN-γ from a cDNA library generated from primary duck hepatocytes and demonstration that duck IFN-γ inhibits duck hepatitis B virus in a dose-dependent manner (81, 82), it has been found that duck IFN-γ shares both structural and functional identities with chicken IFN-γ (83). In contrast to chicken and duck IFN-γ, goose IFN-γ exerts only a weak antiviral state, which may indicate distinct biological activities between these two species (55). It is interesting to observe that the cross-species reactivity of type II IFN has been shown to be considerably higher compared to any other IFN types (48). For instance, recombinant pigeon and turkey IFN-γ was found to be functionally active in chicken cells (48, 49). In conclusion, despite of structural and functional similarities between type II IFNs in different avian species, drivers of differential antiviral activities and molecular mechanisms of diverse immunological responses induced by type II IFNs are yet to be determined in different avian species.
Avian Type III IFN
While at least four IFN-λ genes (IFN-λ1, IFN-λ2, IFN-λ3, and IFN-λ4) were identified in humans (84), only one functionally conserved IFN-λ copy was identified in chicken (50, 51). Chicken IFN-λ shows high sequence identity with human IFN-λ3. The antiviral activities of type III IFNs are dependent on the heterodimeric IFN-λ receptor, which is composed of the IFN-λ-specific IL-28Rα (IFNLR1) chain and the IL-10Rβ (IFNLR2) chain in mammals. Expression of chicken IL-28Rα was also shown to be indispensable for the antiviral activity of chicken IFN-λ (52). Similar to mammals, expression of chicken IL-28Rα appeared to be highest on epithelial cells and in epithelium-rich organs (52). Moreover, avian type III IFN might be functionally conserved compared to those of mammalian species, likely playing a predominant role in the antiviral defense of epithelial barriers (85). This view is further supported by antiviral activity of chicken IFN-λ against several respiratory pathogens, including NDV, IBV, and influenza viruses in vivo, in ovo, and in epithelial cells and in tissue culture systems (52). In contrast, chicken IFN-λ showed only low to moderate antiviral effects on non-epithelial cells, such as primary chicken embryo fibroblasts (CEFs), DF-1 chicken fibroblasts, or the chicken macrophage cell line HD11 (50, 52). This is in line with the assumption that the expression of chicken IL-28Rα is low or absent in most non-epithelial cell types (52).
In contrast to chicken IFN-γ, which induces high levels of nitric oxide in immune cells, IFN-λ as well as IFN-β induces significantly lower levels of nitric oxide in different non-epithelial cell types (50). Recently, it has been shown that chicken IFN-λ inhibits influenza virus replication in CEFs; however, it requires higher doses for achieving effective antiviral activities and to induce ISGs as compared to chicken IFN-γ and IFN-β (63).
In addition to chicken IFN-λ, Yao and colleagues have recently cloned IFN-λ from Pekin ducks and have found that duck IFN-λ is genetically and structurally highly conserved to other avian and mammalian IFN-λ genes (86). Recombinant duck IFN-λ was capable of inducing ISGs (2′-5′-OAS and Mx) in primary duck hepatocytes. Only very little information is available on IFN-λ homologs in other avian species (Figures 6C,D).
A Cross Talk Between Type I, II, and III IFNs
Following production, IFNs initiate the induction of ISGs by binding to their respective IFN receptors in autocrine and paracrine manners (1). Despite the fact that all types of IFNs play distinct and dedicated roles, a significant functional and regulatory overlap among all types of IFNs has been identified. Type I IFNs (IFN-α/β in the case of chickens) are produced from fibroblasts, whereas the antiviral actions of type III IFNs are mainly restricted to epithelial cells (52). These cell-specific roles are probably linked to the expression of cognate receptors in these organs for their importance in specific system.
It has been shown in mammals that type I and III IFNs initiate the same signaling pathway through phosphorylation of STAT1 and STAT2 heterodimers possibly by tyrosine kinase 2 (TYK2) and JAK1 kinases (1) (Figure 1). However, type II IFNs trigger ISGs’ induction via the activation of STAT1 homodimers by JAK1 and JAK2 kinases (1). Several protein phosphatases and the suppressors of cytokine signaling (SOCS), such as SOCS1 and SOCS3, were found to be involved in negative regulation of STATs phosphorylation (87). Although there are discrete downstream JAK–STAT signaling pathways for different type of IFNs, it has been shown that antibody-based neutralization of type I IFNs, or their receptors, attenuate the type II IFN responses. This may be linked to possible common receptor components or to the priming effect of type I IFNs on the expression of common transcription factors (e.g., STAT1) (1), which could cross-link the signaling between the three types of IFNs. Most components of JAK–STAT signaling pathway have been identified in chickens (Figure 1) and ducks, indicating possible functional homologies between mammals and avians.
To initiate the transcriptional activation of ISGs and other cytokines, type I and III IFNs mediate the recruitment and phosphorylation of IRF9 and STAT1/STAT2 heterodimer, to constitute a functional ISGF3 (1, 88). Type II IFNs initiate the formation of a STAT1–STAT1 homodimer to assemble GAF, without the need of IRF9. Upon nuclear translocation, ISGF3 and GAF bind to IFN-stimulated response elements (ISREs) (88) or GAS element, respectively (1). These events consequently lead to the transcriptional activation of hundreds of ISGs (Figure 1). In mammals, IRF9 is required for ISRE promoter activation (1). However, as indicated before, this transcription factor has not yet been identified in chickens, raising the question of alternative mechanisms of types I and III IFN-mediated ISG induction.
Regardless of the nature of their induction, ISGs play fundamental roles in a wide range of cellular activities, including transcriptional and translational regulation of immune responses (89, 90). The collective actions of these ISGs counteract viral replication and provide an antagonistic environment to limit virus propagation and spread (detailed below).
Avian Antiviral Effectors
Binding of type I, II, and III IFNs to their respective receptors leads to the initiation of signaling cascades that culminate in the induction of distinct set of >300 ISGs (at least in human, mouse, and rats) (1, 5). These ISGs create an antiviral state and safeguard the host with multilayered, often synergistic, and cumulative actions (91). ISGs act on several stages of the viral replication cycle, ranging from virus entry to virus release (91). Some of these ISGs are PRRs that potentiate virus detection and thus modulate IFN induction through an amplification loop resulting in enhanced IFN production and hence more efficient virus inhibition (1, 91). Some ISGs have direct antiviral roles by acting at the level of host protein translation, post-transcriptional, and post-translational modifications. Significant advancements have been made in screening and mapping the antiviral roles of many ISGs against a broad range of viral pathogens (5). However, similar investigations have just been started in avian species. High throughput host gene expression profiling strategies, such as next-generation sequencing and microarray transcriptome analysis, have provided a snapshot of the ISGs that might have essential roles against avian viruses (92, 93). While the majority of these identified ISGs are still uncharacterized, a comparative knowledge of chicken/avian ISGs with their mammalian counterparts indicates that some of these ISGs are genetically and functionally conserved and are likely crucial for the control of viral infections. Of the hundreds of ISGs identified in mammals, only few have been genetically and functionally characterized in chicken. These include IFN-inducible transmembrane protein (IFITM)3 (94), which can inhibit virus entry; Mx (95), which can block early stages of virus replication; viperin (40), which can inhibit virus release; ZAP (41), which can weaken viral mRNA translation; 2′-5′-oligoadenylate synthetases (2′-5′-OAS/RNaseL) (54), which can cleave viral RNA transcripts; and PKR (62), which can sense TLR-mediated immune responses (Figure 1). To our knowledge, these are the only ISGs that have been, to date, functionally characterized in chickens. A brief description of individually known avian ISGs is provided below.
CCCH-Type Zinc Finger Antiviral Protein (ZC3HAV1)
The antiviral action of ZC3HAV1 (ZAP) in mammals is mediated by its specific binding to the ZAP-responsive element encoded within viral mRNA (96). This binding recruits the host cellular degradation machinery to disable the viral mRNA translation specifically without any damage to host mRNA (96). Recently, chicken ZAP has been genetically characterized, and it appeared that the antiviral role of ZAP is probably evolutionarily conserved among vertebrates (41). In contrast to the presence of a long and a short ZAP isoforms in mammals, only one isoform (tentatively suggested to be the long isoform) has been found in chickens (41). The shorter isoform in mammals has recently been recognized as a positive regulator of the RIG-I pathway (97). While it remains to be finally clarified that chickens lack a shorter ZAP isoform, it may have been coevolutionary lost along with the RIG-I ortholog in chicken (41). The chicken ZAP gene can be prominently induced by polyinosinic:polycytidylic acid (poly I:C, a synthetic dsRNA analog) and type I IFN treatment in avian cells, suggesting that ZAP is an ISG (41). Moreover, the potential relevance of chicken ZAP in viral pathobiology is likely due to its upregulation in influenza H5N1 and IBDV-infected chickens (41). However, future studies are required to investigate whether all avian species have this protein and whether its functions are similar to those of its mammalian counterparts.
IIFITM Members
Several members of the IFITM family including IFITM1, IFITM2, IFITM3, and IFITM5 have been identified in humans (34). They are differentially expressed upon stimulation by type I and type II IFNs, either in the majority of body tissues (IFITM1, IFITM2, and IFITM3) or exclusively in osteoblasts (IFITM5) (34). Recently, functions of these ISGs have been studied extensively against viruses of medical, zoonotic, and veterinary importance (34). IFITM proteins inhibit viral infection by blocking cytoplasmic entry (98). Mechanistically, IFITM proteins suppress viral membrane fusion due to reduced membrane fluidity and thus forming curvature in the outer leaflets of cell membranes (99); or by disturbing the intracellular cholesterol homeostasis by preventing association of vesicle-membrane-protein-associated protein A with oxysterol-binding protein (100). Recently, three chicken IFITM proteins (IFITM1, IFITM2, and IFITM3) have been genetically characterized, and IFITM2 and IFITM3 have been functionally characterized (94). Despite of low sequence homology, human and chicken IFITM2 and IFITM3 are functionally conversed and are potent inhibitors of influenza and lyssaviruses (94). However, it remains to be determined whether the antiviral mechanisms of chicken and mammalian IFITM members are similar. Recently, it has been demonstrated that the duck IFITM3 confers antiviral activities against influenza viruses and that this action is independent of the N-terminal region of IFITM3 (101). Interestingly, several structural divergences were observed in the duck IFITMs probably owing to host–viral coevolution. Different publically available databases clearly indicate the presence of IFITM member proteins in several other avian species with variable levels of sequence and possible functional similarities. This leaves an opportunity to identify and characterize these important effector proteins of the innate immune system and to map their functions in avians.
Myxovirus-Resistance Proteins
Myxovirus-resistance proteins are GTPases that are key antiviral effector proteins of the type I and type III IFN pathways. In mammals, two major forms of Mx protein exist, namely MxA- and MxB-like Mx proteins (102–105). Mammalian MxA-like proteins, such as human MxA or mouse Mx1, are known to be potent inhibitors of influenza and a broad range of other viruses (102–104, 106). In contrast, the human MxB has only recently been shown to inhibit retrovirus infections (107). To date, only one lineage of Mx genes is known in birds (108, 109). Avian Mx proteins appear to be structurally similar to its mammalian counterparts, containing a GTP-binding and a leucine zipper motif, but they possess a unique N-terminal part that lacks significant homology with mammalian Mx proteins (109–111). Chicken Mx is distributed mainly in the cytoplasm (110, 112), while duck Mx has been shown to be located in cytoplasm and nucleus (111). To date, the GTPase activity for chicken Mx has not been demonstrated (113), and conflicting results have been reported on the antiviral activity of avian Mx proteins. In its first description, chicken Mx was reported to lack antiviral functions against a broad range of RNA viruses including influenza A viruses, Thogotovirus, VSV, and Sendai virus (110). A subsequent study identified a high degree of genetic diversity in the chicken Mx gene (114). Functional assays demonstrated that chicken Mx alleles carrying an asparagine at amino acid position 631 (Mx-Asn631) possess antiviral activity against VSV and HPAIV H5N1 in transfected mouse cells, whereas alleles carrying a serine at this position (Mx-Ser631) lacked antiviral activity (114, 115). While some studies confirmed the antiviral effects of Mx-Asn631 against VSV and NDV in cell culture (116–118), others failed to demonstrate Mx-mediated resistance of both Mx variants against influenza, NDV, and Thogotovirus using comparable approaches (112, 113, 119, 120). Artificial translocation of chicken Mx to the nucleus did not enhance its antiviral activity (112). In vivo studies either did not demonstrate an effect of the polymorphism at position 631 on the clinical course of an experimental HPAIV H7N1 infection (121) or reported an association of Mx–Asn631 with slightly reduced mortality and morbidity following HPAIV H5N2 infections of chickens (120). Overexpression of duck Mx in murine cells did not result in enhanced antiviral activity against VSV and HPAIV H7N1 (111).
In summary, the functional characteristics of avian Mx proteins, their role in innate antiviral immunity, and the effect of genetic polymorphisms are still poorly understood and require further investigations. It is possible that, similar to human MxB, avian Mx proteins possess unequivocal antiviral activities against viruses substantially differing from the few RNA virus families, which have been tested so far.
Protein Kinase R
Protein kinase R is a serine/threonine protein kinase and consists of two domains that are functionally independent; the dsRNA-binding N-terminus and the catalytic C-terminus domains (122). PKR was first identified during investigations on the translation inhibition of viral and cellular mRNAs in vaccinia virus (VV)-infected mammalian cells (123). In an inactive form, PKR localizes in the nucleus and upon activation, mediated through viral dsRNA recognition, oxidative stress, growth factors, cytokines, and cellular proteins such as PKR-associated activator, or following the stimulation of TLRs, phosphorylates the eukaryotic initiation factor 2. This action impairs the guanine nucleotide exchange reaction and thus inhibits translation of mRNA in infected cells (124, 125). Although different viruses, including influenza virus, herpes simplex virus type I, and hepatitis C virus, encode for counteracting factors to inhibit PKR actions, this kinase can still surpass and can exert antiviral activities.
It has been demonstrated that chicken PKR carries all features characteristic for RNA-binding proteins and kinase families (126). Similar to the chicken Mx gene, chicken PKR is also polymorphic and confers antiviral effects against VSV (126). However, in an in vivo study, transcriptionally upregulated PKR failed to protect chickens from highly pathogenic H5N1 infection (127).
Similar to Mx and several cytokines, it is likely that specific SNPs may define the function of PKR in a specific and understudied avian population. Although PKR is one of the first identified PRRs, our understanding of its function is still incomplete even in mammals. In this regards, a novel role of PKR in specifically maintaining the integrity of newly synthesized IFN mRNAs has been recently described (128), further highlighting the need for future research (124).
2′-5′-Oligoadenylate Synthetase
In an attempt to understand the molecular mechanism of PKR-induced inhibition of protein synthesis during VV replication, another enzyme called 2′-5′-OAS was identified in mouse (129). Interestingly, 2′-5′-OAS mRNA has been detected in erythrocytes and immature red blood cells in several avian species (chicken, goose, and pigeon) (130, 131). The same group also identified the existence of two alleles of the 2′-5′-OAS gene in chickens (132). They found that 2′-5′-OAS A/B allele encodes for 58 and 54 kDa synthetases, whereas chickens carrying 2′-5′-OAS A/A alleles produce only a single 58 kDa protein (133). Expression of each of these two chicken 2′-5′-OAS alleles has been revealed to be age-dependent (133). The stability and persistence of 2′-5′-OAS are determined by the ubiquitin-like domain in the carboxyl-terminus of the 2′-5′-OAS (134). Interestingly, basal 2′-5′-OAS expression was systemically detected in chicken embryos independent of stimuli (130). However, a significant induction of 2′-5′-OAS was observed in IFN-treated chicken embryo cells (135).
More recently, the antiviral activity of chicken 2′-5′-OAS against West Nile virus was demonstrated in a replicon assay in mammalian cells (136). Notably, this assay provides the ability to investigate the effect of allele-specific antiviral actions of 2′-5′-OAS against avian viruses with diverse genetic backgrounds.
Viperin
Viperin is one of the most important IFN effectors in mammals and confers antiviral activity by inhibiting the trafficking of soluble viral proteins in the cytoplasmic compartments. Limited availability of the viral components may restrict viral spread (137, 138). Moreover, several studies have also found that mammalian viperin impairs virus replication and restricted viral budding (139). The recently characterized chicken viperin exhibits mammalian-like domains, including a variable N-terminal variable region spanning 77 amino acids, a central radical SAM domain, and a C-terminal conserved region (40). While chicken viperin was significantly induced by influenza viruses and IBDV as well as by different innate immune receptor ligands both in vitro and in vivo (40), its antiviral potential requires future investigations. Since chicken viperin carries leucine zipper and radical SAM motifs, which are known to be essential for viperin-induced antiviral activities in mammals, it is conceivable that chicken viperin has functional conservation with the mammalian counterpart.
Conclusion and Future of Avian Innate Immunity Research
The currently available information on the immunogenetics of avian IFNs is a basis for future research aimed to understand the molecular mechanisms of IFN induction, associated factors, and to identify uncharacterized IFNs in different avian species, which differ significantly in their IFNs pathways and harbor viruses of both veterinary and medical importance. Because of existing functional and genetic differences, it might be needed to revise the nomenclature of avian IFNs to truly represent their origins and actions. Although IFNs were discovered by Isaacs and Lindenmann in chicken cells (140), knowledge on the dynamics and plasticity of chicken IFNs and their antiviral activities is markedly scarce compared to their mammalian counterparts. An important and evolutionary crucial area of research is to understand the potent innate immune responses in chicken in the apparent absence of essential components of IFN pathways, such as RIG-I, especially in chicken and turkey. Recent availability of genomics data on different avian species has significantly advanced comparative immunogenetics studies. However, extensive efforts are required to improve the current genome annotation of widely used poultry species (chicken, duck, and turkey) and to effectively characterize existing gaps in functionally important genomic loci. Investigations on functional implications of avian ISGs have been started; however, next-generation strategies would be required to map the antiviral or possible proviral roles of these IFN effectors. Most actions of ISGs have been studied using single isoforms of the ISGs exploiting either ectopic expression or silencing methods. Approaches such as CRISPR/Cas9 knockout/knockin will be required for future investigations on effective mapping avian ISG and their functions. One of the aspects that might require future efforts is to identify the overlapping antiviral roles of ISGs and the molecular combinatorial networking in these antiviral, or proviral, properties. Since silencing of individual ISGs leads to observable differences in virus pathobiology, these appear to be valuable targets for the development of potential therapeutics for a broader range of viruses, and for vaccine production. In this regard, human IFNs have been successfully applied for the treatment of virus-induced human diseases; however, the clinical potential of chicken or other avian IFNs has not yet been exploited. These applications may hold options for future economical antiviral therapy not only in commercial poultry but also in companion birds.
Author Contributions
Conception and writing of the manuscript: MM; restructuring and improvement of the contents: DR, LM-S, and DS; and designing the figures: MM and DS.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This study was supported by the Biotechnology and Biological Sciences Research Council (grant numbers BB/M008681/1 and BBS/E/I/00001852) and The British Council (grant number 172710323). We are thankful to Peter Stäheli, Daniel Schnepf, and Mirjam Schilling (all from Freiburg, Germany) for their valuable comments and suggestions.
Funding
Funding agencies have no role in design of the manuscript, nature of publication, and type of the manuscript.
References
1. Randall RE, Goodbourn S. Interferons and viruses: an interplay between induction, signalling, antiviral responses and virus countermeasures. J Gen Virol (2008) 89(Pt 1):1–47. doi:10.1099/vir.0.83391-0
2. Baron S, Dianzani F. The interferons: a biological system with therapeutic potential in viral infections. Antiviral Res (1994) 24(2–3):97–110. doi:10.1016/0166-3542(94)90058-2
3. Fensterl V, Sen GC. Interferons and viral infections. Biofactors (2009) 35(1):14–20. doi:10.1002/biof.6
4. Rathinam VA, Fitzgerald KA. Innate immune sensing of DNA viruses. Virology (2011) 411(2):153–62. doi:10.1016/j.virol.2011.02.003
5. Schoggins JW, Wilson SJ, Panis M, Murphy MY, Jones CT, Bieniasz P, et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature (2011) 472(7344):481–5. doi:10.1038/nature09907
6. Diner BA, Lum KK, Cristea IM. The emerging role of nuclear viral DNA sensors. J Biol Chem (2015) 290(44):26412–21. doi:10.1074/jbc.R115.652289
7. Hayden MS, Ghosh S. Signaling to NF-kappaB. Genes Dev (2004) 18(18):2195–224. doi:10.1101/gad.1228704
8. Munir M. TRIM proteins: another class of viral victims. Sci Signal (2010) 3(118):jc2. doi:10.1126/scisignal.3118jc2
9. Schlee M, Hartmann G. Discriminating self from non-self in nucleic acid sensing. Nat Rev Immunol (2016) 16(9):566–80. doi:10.1038/nri.2016.78
10. Uchikawa E, Lethier M, Malet H, Brunel J, Gerlier D, Cusack S. Structural analysis of dsRNA binding to anti-viral pattern recognition receptors LGP2 and MDA5. Mol Cell (2016) 62(4):586–602. doi:10.1016/j.molcel.2016.04.021
11. Karpala AJ, Stewart C, McKay J, Lowenthal JW, Bean AG. Characterization of chicken Mda5 activity: regulation of IFN-beta in the absence of RIG-I functionality. J Immunol (2011) 186(9):5397–405. doi:10.4049/jimmunol.1003712
12. Liniger M, Summerfield A, Zimmer G, McCullough KC, Ruggli N. Chicken cells sense influenza A virus infection through MDA5 and CARDIF signaling involving LGP2. J Virol (2012) 86(2):705–17. doi:10.1128/JVI.00742-11
13. Hayashi T, Watanabe C, Suzuki Y, Tanikawa T, Uchida Y, Saito T. Chicken MDA5 senses short double-stranded RNA with implications for antiviral response against avian influenza viruses in chicken. J Innate Immun (2014) 6(1):58–71. doi:10.1159/000351583
14. Barber MRW, Aldridge JR Jr., Webster RG, Magor KE. Association of RIG-I with innate immunity of ducks to influenza. Proc Natl Acad Sci U S A (2010) 107(13):5913–8. doi:10.1073/pnas.1001755107
15. Shao Q, Xu W, Yan L, Liu J, Rui L, Xiao X, et al. Function of duck RIG-I in induction of antiviral response against IBDV and avian influenza virus on chicken cells. Virus Res (2014) 191:184–91. doi:10.1016/j.virusres.2014.07.028
16. Roach JC, Glusman G, Rowen L, Kaur A, Purcell MK, Smith KD, et al. The evolution of vertebrate toll-like receptors. Proc Natl Acad Sci U S A (2005) 102(27):9577–82. doi:10.1073/pnas.0502272102
17. Keestra AM, de Zoete MR, Bouwman LI, Vaezirad MM, van Putten JPM. Unique features of chicken toll-like receptors. Dev Comp Immunol (2013) 41(3):316–23. doi:10.1016/j.dci.2013.04.009
18. de Zoete MR, Bouwman LI, Keestra AM, van Putten JPM. Cleavage and activation of a toll-like receptor by microbial proteases. Proc Natl Acad Sci U S A (2011) 108(12):4968–73. doi:10.1073/pnas.1018135108
19. Keestra AM, van Putten JPM. Unique properties of the chicken TLR4/MD-2 complex: selective lipopolysaccharide activation of the MyD88-dependent pathway. J Immunol (2008) 181(6):4354–62. doi:10.4049/jimmunol.181.6.4354
20. Cavlar T, Ablasser A, Hornung V. Induction of type I IFNs by intracellular DNA-sensing pathways. Immunol Cell Biol (2012) 90(5):474–82. doi:10.1038/icb.2012.11
21. Unterholzner L, Keating SE, Baran M, Horan KA, Jensen SB, Sharma S, et al. IFI16 is an innate immune sensor for intracellular DNA. Nat Immunol (2010) 11(11):997–1004. doi:10.1038/ni.1932
22. Cridland JA, Curley EZ, Wykes MN, Schroder K, Sweet MJ, Roberts TL, et al. The mammalian PYHIN gene family: phylogeny, evolution and expression. BMC Evol Biol (2012) 12:140. doi:10.1186/1471-2148-12-140
23. Cheng YQ, Sun YJ, Wang HA, Yan YX, Ding C, Sun JH. Chicken STING mediates activation of the IFN gene independently of the RIG-I gene. J Immunol (2015) 195(8):3922–36. doi:10.4049/jimmunol.1500638
24. Wang Z, Ning Z, Sun M, Gao S, Kang Y, Xie P, et al. Interferon regulatory factor 7- (IRF7-) mediated immune response affects Newcastle disease virus replication in chicken embryo fibroblasts. Acta Vet Hung (2014) 62(4):500–11. doi:10.1556/AVet.2014.023
25. Kim TH, Zhou H. Functional analysis of chicken IRF7 in response to dsRNA analog poly(I:C) by integrating overexpression and knockdown. PLoS One (2015) 10(7):e0133450. doi:10.1371/journal.pone.0133450
26. May DL, Grant CE, Deeley RG. Cloning and promoter analysis of the chicken interferon regulatory factor-3 gene. DNA Cell Biol (2000) 19(9):555–66. doi:10.1089/104454900439782
27. Barjesteh N, Abdelaziz KT, Sharif S. The role of IRF7 and NF-κB pathways in the induction of antiviral responses in chicken tracheal epithelial cells following exposure to TLR3 and 4 ligands. Poster in Veterinary and Comparative Immunology (2016). Available from: https://immunology2016.zerista.com/event/member/256436
28. Zhang G, Jarvis ED, Gilbert MT. Avian genomes. A flock of genomes. Introduction. Science (2014) 346(6215):1308–9. doi:10.1126/science.346.6215.1308
29. Eory L, Gilbert MT, Li C, Li B, Archibald A, Aken BL, et al. Avianbase: a community resource for bird genomics. Genome Biol (2015) 16:21. doi:10.1186/s13059-015-0588-2
30. Rubin CJ, Zody MC, Eriksson J, Meadows JR, Sherwood E, Webster MT, et al. Whole-genome resequencing reveals loci under selection during chicken domestication. Nature (2010) 464(7288):587–91. doi:10.1038/nature08832
31. Albalat R, Canestro C. Evolution by gene loss. Nat Rev Genet (2016) 17(7):379–91. doi:10.1038/nrg.2016.39
32. Demuth JP, De Bie T, Stajich JE, Cristianini N, Hahn MW. The evolution of mammalian gene families. PLoS One (2006) 1:e85. doi:10.1371/journal.pone.0000085
33. Zhou X, Michal JJ, Zhang L, Ding B, Lunney JK, Liu B, et al. Interferon induced IFIT family genes in host antiviral defense. Int J Biol Sci (2013) 9(2):200–8. doi:10.7150/ijbs.5613
34. Diamond MS, Farzan M. The broad-spectrum antiviral functions of IFIT and IFITM proteins. Nat Rev Immunol (2013) 13(1):46–57. doi:10.1038/nri3344
35. Kaiser P, Wain HM, Rothwell L. Structure of the chicken interferon-gamma gene, and comparison to mammalian homologues. Gene (1998) 207(1):25–32. doi:10.1016/S0378-1119(97)00600-8
36. Staeheli P, Puehler F, Schneider K, Gobel TW, Kaspers B. Cytokines of birds: conserved functions – a largely different look. J Interferon Cytokine Res (2001) 21(12):993–1010. doi:10.1089/107999001317205123
37. Lutfalla G, Roest Crollius H, Stange-Thomann N, Jaillon O, Mogensen K, Monneron D. Comparative genomic analysis reveals independent expansion of a lineage-specific gene family in vertebrates: the class II cytokine receptors and their ligands in mammals and fish. BMC Genomics (2003) 4(1):29. doi:10.1186/1471-2164-4-29
38. Roberts RM, Liu L, Guo Q, Leaman D, Bixby J. The evolution of the type I interferons. J Interferon Cytokine Res (1998) 18(10):805–16. doi:10.1089/jir.1998.18.805
39. Hughes AL, Roberts RM. Independent origin of IFN-alpha and IFN-beta in birds and mammals. J Interferon Cytokine Res (2000) 20(8):737–9. doi:10.1089/10799900050116444
40. Goossens KE, Karpala AJ, Rohringer A, Ward A, Bean AG. Characterisation of chicken viperin. Mol Immunol (2015) 63(2):373–80. doi:10.1016/j.molimm.2014.09.011
41. Goossens KE, Karpala AJ, Ward A, Bean AG. Characterisation of chicken ZAP. Dev Comp Immunol (2014) 46(2):373–81. doi:10.1016/j.dci.2014.05.011
42. Sarmento L, Afonso CL, Estevez C, Wasilenko J, Pantin-Jackwood M. Differential host gene expression in cells infected with highly pathogenic H5N1 avian influenza viruses. Vet Immunol Immunopathol (2008) 125(3–4):291–302. doi:10.1016/j.vetimm.2008.05.021
43. Xia C, Liu J, Wu ZG, Lin CY, Wang M. The interferon-alpha genes from three chicken lines and its effects on H9N2 influenza viruses. Anim Biotechnol (2004) 15(1):77–88. doi:10.1081/ABIO-120037900
44. Pei J, Sekellick MJ, Marcus PI, Choi IS, Collisson EW. Chicken interferon type I inhibits infectious bronchitis virus replication and associated respiratory illness. J Interferon Cytokine Res (2001) 21(12):1071–7. doi:10.1089/107999001317205204
45. Mo CW, Cao YC, Lim BL. The in vivo and in vitro effects of chicken interferon alpha on infectious bursal disease virus and Newcastle disease virus infection. Avian Dis (2001) 45(2):389–99. doi:10.2307/1592978
46. Levy AM, Heller ED, Leitner G, Davidson I. Effect of native chicken interferon on MDV replication. Acta Virol (1999) 43(2–3):121–7.
47. Penski N, Hartle S, Rubbenstroth D, Krohmann C, Ruggli N, Schusser B, et al. Highly pathogenic avian influenza viruses do not inhibit interferon synthesis in infected chickens but can override the interferon-induced antiviral state. J Virol (2011) 85(15):7730–41. doi:10.1128/JVI.00063-11
48. Fringuelli E, Urbanelli L, Tharuni O, Proietti PC, Bietta A, Davidson I, et al. Cloning and expression of pigeon IFN-gamma gene. Res Vet Sci (2010) 89(3):367–72. doi:10.1016/j.rvsc.2010.03.021
49. Farnell MB, El Halawani M, You S, McElroy AP, Hargis BM, Caldwell DJ. In vivo biologic effects of recombinant-turkey interferon-gamma in neonatal leghorn chicks: protection against Salmonella enteritidis organ invasion. Avian Dis (2001) 45(2):473–8. doi:10.2307/1592991
50. Karpala AJ, Morris KR, Broadway MM, McWaters PG, O’Neil TE, Goossens KE, et al. Molecular cloning, expression, and characterization of chicken IFN-lambda. J Interferon Cytokine Res (2008) 28(6):341–50. doi:10.1089/jir.2007.0117
51. Kaiser P, Poh TY, Rothwell L, Avery S, Balu S, Pathania US, et al. A genomic analysis of chicken cytokines and chemokines. J Interferon Cytokine Res (2005) 25(8):467–84. doi:10.1089/jir.2005.25.467
52. Reuter A, Soubies S, Hartle S, Schusser B, Kaspers B, Staeheli P, et al. Antiviral activity of lambda interferon in chickens. J Virol (2014) 88(5):2835–43. doi:10.1128/JVI.02764-13
53. Sick C, Schultz U, Staeheli P. A family of genes coding for two serologically distinct chicken interferons. J Biol Chem (1996) 271(13):7635–9. doi:10.1074/jbc.271.13.7635
54. Goossens KE, Ward AC, Lowenthal JW, Bean AG. Chicken interferons, their receptors and interferon-stimulated genes. Dev Comp Immunol (2013) 41(3):370–6. doi:10.1016/j.dci.2013.05.020
55. Li HT, Ma B, Mi JW, Jin HY, Xu LN, Wang JW. Cloning, in vitro expression and bioactivity of goose interferon-alpha. Cytokine (2006) 34(3–4):177–83. doi:10.1016/j.cyto.2006.04.013
56. Jiang H, Yang H, Kapczynski DR. Chicken interferon alpha pretreatment reduces virus replication of pandemic H1N1 and H5N9 avian influenza viruses in lung cell cultures from different avian species. Virol J (2011) 8:447. doi:10.1186/1743-422X-8-447
57. Hertzog PJ, Williams BR. Fine tuning type I interferon responses. Cytokine Growth Factor Rev (2013) 24(3):217–25. doi:10.1016/j.cytogfr.2013.04.002
58. Guo SY, Li C, Dai XM, Zhao C, Chang WS. Distribution of the type I interferon in different organs of chicken digestive system. Cent Eur J Immunol (2014) 39(4):455–60. doi:10.5114/ceji.2014.47728
59. Zhou H, Chen S, Wang M, Cheng A. Interferons and their receptors in birds: a comparison of gene structure, phylogenetic analysis, and cross modulation. Int J Mol Sci (2014) 15(11):21045–68. doi:10.3390/ijms151121045
60. Sekellick MJ, Ferrandino AF, Hopkins DA, Marcus PI. Chicken interferon gene: cloning, expression, and analysis. J Interferon Res (1994) 14(2):71–9. doi:10.1089/jir.1994.14.71
61. Meng S, Yang L, Xu C, Qin Z, Xu H, Wang Y, et al. Recombinant chicken interferon-alpha inhibits H9N2 avian influenza virus replication in vivo by oral administration. J Interferon Cytokine Res (2011) 31(7):533–8. doi:10.1089/jir.2010.0123
62. Qu H, Yang L, Meng S, Xu L, Bi Y, Jia X, et al. The differential antiviral activities of chicken interferon alpha (ChIFN-alpha) and ChIFN-beta are related to distinct interferon-stimulated gene expression. PLoS One (2013) 8(3):e59307. doi:10.1371/journal.pone.0059307
63. Masuda Y, Matsuda A, Usui T, Sugai T, Asano A, Yamano Y. Biological effects of chicken type III interferon on expression of interferon-stimulated genes in chickens: comparison with type I and type II interferons. J Vet Med Sci (2012) 74(11):1381–6. doi:10.1292/jvms.11-0517
64. Sick C, Schultz U, Munster U, Meier J, Kaspers B, Staeheli P. Promoter structures and differential responses to viral and nonviral inducers of chicken type I interferon genes. J Biol Chem (1998) 273(16):9749–54. doi:10.1074/jbc.273.16.9749
65. Karpala AJ, Bagnaud-Baule A, Goossens KE, Lowenthal JW, Bean AG. Ontogeny of the interferon system in chickens. J Reprod Immunol (2012) 94(2):169–74. doi:10.1016/j.jri.2012.02.008
66. Karpala AJ, Lowenthal JW, Bean AG. Activation of the TLR3 pathway regulates IFNbeta production in chickens. Dev Comp Immunol (2008) 32(4):435–44. doi:10.1016/j.dci.2007.08.004
67. Hata N, Sato M, Takaoka A, Asagiri M, Tanaka N, Taniguchi T. Constitutive IFN-alpha/beta signal for efficient IFN-alpha/beta gene induction by virus. Biochem Biophys Res Commun (2001) 285(2):518–25. doi:10.1006/bbrc.2001.5159
68. Ziegler RE, Joklik WK. Effect of interferon on multiplication of avian sarcoma virus B77 in duck embryo fibroblasts. J Interferon Res (1981) 1(4):521–38. doi:10.1089/jir.1981.1.521
69. Schultz U, Kock J, Schlicht HJ, Staeheli P. Recombinant duck interferon: a new reagent for studying the mode of interferon action against hepatitis B virus. Virology (1995) 212(2):641–9. doi:10.1006/viro.1995.1522
70. Tian L, Zhao P, Ma B, Guo G, Sun Y, Xing M. Cloning, expression and antiviral bioactivity of red-crowned crane interferon-alpha. Gene (2014) 544(1):49–55. doi:10.1016/j.gene.2014.04.036
71. Suresh M, Karaca K, Foster D, Sharma JM. Molecular and functional characterization of turkey interferon. J Virol (1995) 69(12):8159–63.
72. Han CL, Zhang W, Dong HT, Han X, Wang M. A novel gene of beta chain of the IFN-gamma receptor of Huiyang chicken: cloning, distribution, and CD assay. J Interferon Cytokine Res (2006) 26(7):441–8. doi:10.1089/jir.2006.26.441
73. Han X, Chen T, Wang M. Molecular cloning and characterization of chicken interferon-gamma receptor alpha-chain. J Interferon Cytokine Res (2008) 28(7):445–54. doi:10.1089/jir.2007.0135
74. Digby MR, Lowenthal JW. Cloning and expression of the chicken interferon-gamma gene. J Interferon Cytokine Res (1995) 15(11):939–45. doi:10.1089/jir.1995.15.939
75. Haq K, Wootton SK, Barjesteh N, St Paul M, Golovan S, Bendall AJ, et al. Small interfering RNA-mediated knockdown of chicken interferon-gamma expression. J Interferon Cytokine Res (2013) 33(6):319–27. doi:10.1089/jir.2012.0141
76. Yuk SS, Lee DH, Park JK, Tseren-Ochir EO, Kwon JH, Noh JY, et al. Pre-immune state induced by chicken interferon gamma inhibits the replication of H1N1 human and H9N2 avian influenza viruses in chicken embryo fibroblasts. Virol J (2016) 13:71. doi:10.1186/s12985-016-0527-1
77. Haq K, Wootton SK, Barjesteh N, Golovan S, Bendall A, Sharif S. Effects of interferon-gamma knockdown on vaccine-induced immunity against Marek’s disease in chickens. Can J Vet Res (2015) 79(1):1–7.
78. Sawant PM, Verma PC, Subudhi PK, Chaturvedi U, Singh M, Kumar R, et al. Immunomodulation of bivalent Newcastle disease DNA vaccine induced immune response by co-delivery of chicken IFN-gamma and IL-4 genes. Vet Immunol Immunopathol (2011) 144(1–2):36–44. doi:10.1016/j.vetimm.2011.07.006
79. Susta L, Cornax I, Diel DG, Garcia SC, Miller PJ, Liu X, et al. Expression of interferon gamma by a highly virulent strain of Newcastle disease virus decreases its pathogenicity in chickens. Microb Pathog (2013) 61-62:73–83. doi:10.1016/j.micpath.2013.05.009
80. Mallick AI, Parvizi P, Read LR, Nagy E, Behboudi S, Sharif S. Enhancement of immunogenicity of a virosome-based avian influenza vaccine in chickens by incorporating CpG-ODN. Vaccine (2011) 29(8):1657–65. doi:10.1016/j.vaccine.2010.12.046
81. Lavine JE, Ganem D. Inhibition of duck hepatitis B virus replication by interferon-gamma. J Med Virol (1993) 40(1):59–64. doi:10.1002/jmv.1890400112
82. Schultz U, Chisari FV. Recombinant duck interferon gamma inhibits duck hepatitis B virus replication in primary hepatocytes. J Virol (1999) 73(4):3162–8.
83. Huang A, Scougall CA, Lowenthal JW, Jilbert AR, Kotlarski I. Structural and functional homology between duck and chicken interferon-gamma. Dev Comp Immunol (2001) 25(1):55–68. doi:10.1016/S0145-305X(00)00041-0
84. O’Brien TR, Prokunina-Olsson L, Donnelly RP. IFN-lambda4: the paradoxical new member of the interferon lambda family. J Interferon Cytokine Res (2014) 34(11):829–38. doi:10.1089/jir.2013.0136
85. Zhang Z, Zou T, Hu X, Jin H. Type III interferon gene expression in response to influenza virus infection in chicken and duck embryonic fibroblasts. Mol Immunol (2015) 68(2 Pt C):657–62. doi:10.1016/j.molimm.2015.10.013
86. Yao Q, Fischer KP, Arnesen K, Tyrrell DL, Gutfreund KS. Molecular cloning, expression and characterization of Pekin duck interferon-lambda. Gene (2014) 548(1):29–38. doi:10.1016/j.gene.2014.06.066
87. Yasukawa H, Misawa H, Sakamoto H, Masuhara M, Sasaki A, Wakioka T, et al. The JAK-binding protein JAB inhibits Janus tyrosine kinase activity through binding in the activation loop. EMBO J (1999) 18(5):1309–20. doi:10.1093/emboj/18.5.1309
88. Kessler DS, Veals SA, Fu XY, Levy DE. Interferon-alpha regulates nuclear translocation and DNA-binding affinity of ISGF3, a multimeric transcriptional activator. Genes Dev (1990) 4(10):1753–65. doi:10.1101/gad.4.10.1753
89. de Veer MJ, Holko M, Frevel M, Walker E, Der S, Paranjape JM, et al. Functional classification of interferon-stimulated genes identified using microarrays. J Leukoc Biol (2001) 69(6):912–20.
90. To KK, Lau CC, Woo PC, Lau SK, Chan JF, Chan KH, et al. Human H7N9 virus induces a more pronounced pro-inflammatory cytokine but an attenuated interferon response in human bronchial epithelial cells when compared with an epidemiologically-linked chicken H7N9 virus. Virol J (2016) 13:42. doi:10.1186/s12985-016-0498-2
91. Schneider WM, Chevillotte MD, Rice CM. Interferon-stimulated genes: a complex web of host defenses. Annu Rev Immunol (2014) 32:513–45. doi:10.1146/annurev-immunol-032713-120231
92. Giotis ES, Robey RC, Skinner NG, Tomlinson CD, Goodbourn S, Skinner MA. Chicken interferome: avian interferon-stimulated genes identified by microarray and RNA-seq of primary chick embryo fibroblasts treated with a chicken type I interferon (IFN-alpha). Vet Res (2016) 47(1):75. doi:10.1186/s13567-016-0363-8
93. Kang Y, Feng M, Zhao X, Dai X, Xiang B, Gao P, et al. Newcastle disease virus infection in chicken embryonic fibroblasts but not duck embryonic fibroblasts is associated with elevated host innate immune response. Virol J (2016) 13:41. doi:10.1186/s12985-016-0499-1
94. Smith SE, Gibson MS, Wash RS, Ferrara F, Wright E, Temperton N, et al. Chicken interferon-inducible transmembrane protein 3 restricts influenza viruses and lyssaviruses in vitro. J Virol (2013) 87(23):12957–66. doi:10.1128/JVI.01443-13
95. Zhang Y, Fu D, Chen H, Zhang Z, Shi Q, Elsayed AK, et al. Partial antiviral activities detection of chicken Mx jointing with neuraminidase gene (NA) against Newcastle disease virus. PLoS One (2013) 8(8):e71688. doi:10.1371/journal.pone.0071688
96. Gao G, Guo X, Goff SP. Inhibition of retroviral RNA production by ZAP, a CCCH-type zinc finger protein. Science (2002) 297(5587):1703–6. doi:10.1126/science.1074276
97. Hayakawa S, Shiratori S, Yamato H, Kameyama T, Kitatsuji C, Kashigi F, et al. ZAPS is a potent stimulator of signaling mediated by the RNA helicase RIG-I during antiviral responses. Nat Immunol (2011) 12(1):37–44. doi:10.1038/ni.1963
98. Yount JS, Moltedo B, Yang YY, Charron G, Moran TM, Lopez CB, et al. Palmitoylome profiling reveals S-palmitoylation-dependent antiviral activity of IFITM3. Nat Chem Biol (2010) 6(8):610–4. doi:10.1038/nchembio.405
99. Li K, Markosyan RM, Zheng YM, Golfetto O, Bungart B, Li M, et al. IFITM proteins restrict viral membrane hemifusion. PLoS Pathog (2013) 9(1):e1003124. doi:10.1371/journal.ppat.1003124
100. Amini-Bavil-Olyaee S, Choi YJ, Lee JH, Shi M, Huang IC, Farzan M, et al. The antiviral effector IFITM3 disrupts intracellular cholesterol homeostasis to block viral entry. Cell Host Microbe (2013) 13(4):452–64. doi:10.1016/j.chom.2013.03.006
101. Blyth GA, Chan WF, Webster RG, Magor KE. Duck interferon-inducible transmembrane protein 3 mediates restriction of influenza viruses. J Virol (2015) 90(1):103–16. doi:10.1128/JVI.01593-15
102. Haller O, Gao S, von der Malsburg A, Daumke O, Kochs G. Dynamin-like MxA GTPase: structural insights into oligomerization and implications for antiviral activity. J Biol Chem (2010) 285(37):28419–24. doi:10.1074/jbc.R110.145839
103. Haller O, Kochs G. Human MxA protein: an interferon-induced dynamin-like GTPase with broad antiviral activity. J Interferon Cytokine Res (2011) 31(1):79–87. doi:10.1089/jir.2010.0076
104. Haller O, Staeheli P, Schwemmle M, Kochs G. Mx GTPases: dynamin-like antiviral machines of innate immunity. Trends Microbiol (2015) 23(3):154–63. doi:10.1016/j.tim.2014.12.003
105. Verhelst J, Hulpiau P, Saelens X. Mx proteins: antiviral gatekeepers that restrain the uninvited (vol 77, pg 551, 2013). Microbiol Mol Biol Rev (2014) 78(1):198–198. doi:10.1128/MMBR.00011-14
106. Goujon C, Moncorge O, Bauby H, Doyle T, Ward CC, Schaller T, et al. Human MX2 is an interferon-induced post-entry inhibitor of HIV-1 infection. Nature (2013) 502(7472):559–62. doi:10.1038/nature12542
107. Liu Z, Pan Q, Ding S, Qian J, Xu F, Zhou J, et al. The interferon-inducible MxB protein inhibits HIV-1 infection. Cell Host Microbe (2013) 14(4):398–410. doi:10.1016/j.chom.2013.08.015
108. Verhelst J, Hulpiau P, Saelens X. Mx proteins: antiviral gatekeepers that restrain the uninvited. Microbiol Mol Biol Rev (2013) 77(4):551–66. doi:10.1128/MMBR.00024-13
109. Zeng M, Chen S, Wang MS, Jia RY, Zhu DK, Liu MF, et al. Molecular identification and comparative transcriptional analysis of myxovirus resistance GTPase (Mx) gene in goose (Anser cygnoide) after H9N2 AIV infection. Comp Immunol Microbiol Infect Dis (2016) 47:32–40. doi:10.1016/j.cimid.2016.05.005
110. Bernasconi D, Schultz U, Staeheli P. The interferon-induced Mx protein of chickens lacks antiviral activity. J Interferon Cytokine Res (1995) 15(1):47–53. doi:10.1089/jir.1995.15.47
111. Bazzigher L, Schwarz A, Staeheli P. No enhanced influenza virus resistance of murine and avian cells expressing cloned duck Mx protein. Virology (1993) 195(1):100–12. doi:10.1006/viro.1993.1350
112. Benfield CT, Lyall JW, Tiley LS. The cytoplasmic location of chicken Mx is not the determining factor for its lack of antiviral activity. PLoS One (2010) 5(8):e12151. doi:10.1371/journal.pone.0012151
113. Schusser B, Reuter A, von der Malsburg A, Penski N, Weigend S, Kaspers B, et al. Mx is dispensable for interferon-mediated resistance of chicken cells against influenza A virus. J Virol (2011) 85(16):8307–15. doi:10.1128/JVI.00535-11
114. Ko JH, Jin HK, Asano A, Takada A, Ninomiya A, Kida H, et al. Polymorphisms and the differential antiviral activity of the chicken Mx gene. Genome Res (2002) 12(4):595–601. doi:10.1101/gr.210702
115. Ko JH, Takada A, Mitsuhashi T, Agui T, Watanabe T. Native antiviral specificity of chicken Mx protein depends on amino acid variation at position 631. Anim Genet (2004) 35(2):119–22. doi:10.1111/j.1365-2052.2004.01096.x
116. Sasaki K, Yoneda A, Ninomiya A, Kawahara M, Watanabe T. Both antiviral activity and intracellular localization of chicken Mx protein depend on a polymorphism at amino acid position 631. Biochem Biophys Res Commun (2013) 430(1):161–6. doi:10.1016/j.bbrc.2012.11.053
117. Li B, Fu D, Zhang Y, Xu Q, Ni L, Chang G, et al. Partial antiviral activities of the Asn631 chicken Mx against Newcastle disease virus and vesicular stomatitis virus. Mol Biol Rep (2012) 39(8):8415–24. doi:10.1007/s11033-012-1694-9
118. Yin YL, Huang RL, Li TJ, Ruan Z, Xie MY, Deng ZY, et al. Amino acid metabolism in the portal-drained viscera of young pigs: effects of dietary supplementation with chitosan and pea hull. Amino Acids (2010) 39(5):1581–7. doi:10.1007/s00726-010-0577-4
119. Benfield CT, Lyall JW, Kochs G, Tiley LS. Asparagine 631 variants of the chicken Mx protein do not inhibit influenza virus replication in primary chicken embryo fibroblasts or in vitro surrogate assays. J Virol (2008) 82(15):7533–9. doi:10.1128/JVI.00185-08
120. Ewald SJ, Kapczynski DR, Livant EJ, Suarez DL, Ralph J, McLeod S, et al. Association of Mx1 Asn631 variant alleles with reductions in morbidity, early mortality, viral shedding, and cytokine responses in chickens infected with a highly pathogenic avian influenza virus. Immunogenetics (2011) 63(6):363–75. doi:10.1007/s00251-010-0509-1
121. Sironi L, Williams JL, Moreno-Martin AM, Ramelli P, Stella A, Jianlin H, et al. Susceptibility of different chicken lines to H7N1 highly pathogenic avian influenza virus and the role of Mx gene polymorphism coding amino acid position 631. Virology (2008) 380(1):152–6. doi:10.1016/j.virol.2008.07.022
122. Feng GS, Chong K, Kumar A, Williams BR. Identification of double-stranded RNA-binding domains in the interferon-induced double-stranded RNA-activated p68 kinase. Proc Natl Acad Sci U S A (1992) 89(12):5447–51. doi:10.1073/pnas.89.12.5447
123. Metz DH, Esteban M. Interferon inhibits viral protein synthesis in L cells infected with vaccinia virus. Nature (1972) 238(5364):385–8. doi:10.1038/238385a0
124. Munir M, Berg M. The multiple faces of proteinkinase R in antiviral defense. Virulence (2013) 4(1):85–9. doi:10.4161/viru.23134
125. Clemens MJ, Elia A. The double-stranded RNA-dependent protein kinase PKR: structure and function. J Interferon Cytokine Res (1997) 17(9):503–24. doi:10.1089/jir.1997.17.503
126. Ko JH, Asano A, Kon Y, Watanabe T, Agui T. Characterization of the chicken PKR: polymorphism of the gene and antiviral activity against vesicular stomatitis virus. Jpn J Vet Res (2004) 51(3–4):123–33.
127. Daviet S, Van Borm S, Habyarimana A, Ahanda ML, Morin V, Oudin A, et al. Induction of Mx and PKR failed to protect chickens from H5N1 infection. Viral Immunol (2009) 22(6):467–72. doi:10.1089/vim.2009.0053
128. Schulz O, Pichlmair A, Rehwinkel J, Rogers NC, Scheuner D, Kato H, et al. Protein kinase R contributes to immunity against specific viruses by regulating interferon mRNA integrity. Cell Host Microbe (2010) 7(5):354–61. doi:10.1016/j.chom.2010.04.007
129. Hovanessian AG, Kerr IM. The (2’-5’) oligoadenylate (pppA2’-5’A2’-5’A) synthetase and protein kinase(s) from interferon-treated cells. Eur J Biochem (1979) 93(3):515–26. doi:10.1111/j.1432-1033.1979.tb12850.x
130. Sokawa J, Shimizu N, Sokawa Y. Presence of (2’-5’)oligoadenylate synthetase in avian erythrocytes. J Biochem (1984) 96(1):215–22.
131. Sokawa J, Sokawa Y. (2’-5’) oligoadenylate synthetase in chicken embryo erythrocytes and immature red blood cells. J Biochem (1986) 99(1):119–24.
132. Yamamoto A, Iwata A, Koh Y, Kawai S, Murayama S, Hamada K, et al. Two types of chicken 2’,5’-oligoadenylate synthetase mRNA derived from alleles at a single locus. Biochim Biophys Acta (1998) 1395(2):181–91. doi:10.1016/S0167-4781(97)00148-6
133. Tatsumi R, Hamada K, Sekiya S, Wakamatsu M, Namikawa T, Mizutani M, et al. 2’,5’-oligoadenylate synthetase gene in chicken: gene structure, distribution of alleles and their expression. Biochim Biophys Acta (2000) 1494(3):263–8. doi:10.1016/S0167-4781(00)00174-3
134. Tatsumi R, Sekiya S, Nakanishi R, Mizutani M, Kojima S, Sokawa Y. Function of ubiquitin-like domain of chicken 2’-5’-oligoadenylate synthetase in conformational stability. J Interferon Cytokine Res (2003) 23(11):667–76. doi:10.1089/107999003322558809
135. West DK, Ball LA. Induction and maintenance of 2’,5’-oligoadenylate synthetase in interferon-treated chicken embryo cells. Mol Cell Biol (1982) 2(11):1436–43. doi:10.1128/MCB.2.11.1436
136. Tag-El-Din-Hassan HT, Sasaki N, Moritoh K, Torigoe D, Maeda A, Agui T. The chicken 2’-5’ oligoadenylate synthetase A inhibits the replication of West Nile virus. Jpn J Vet Res (2012) 60(2–3):95–103.
137. Hinson ER, Cresswell P. The N-terminal amphipathic alpha-helix of viperin mediates localization to the cytosolic face of the endoplasmic reticulum and inhibits protein secretion. J Biol Chem (2009) 284(7):4705–12. doi:10.1074/jbc.M807261200
138. Wang X, Hinson ER, Cresswell P. The interferon-inducible protein viperin inhibits influenza virus release by perturbing lipid rafts. Cell Host Microbe (2007) 2(2):96–105. doi:10.1016/j.chom.2007.06.009
139. Tan KS, Olfat F, Phoon MC, Hsu JP, Howe JL, Seet JE, et al. In vivo and in vitro studies on the antiviral activities of viperin against influenza H1N1 virus infection. J Gen Virol (2012) 93(Pt 6):1269–77. doi:10.1099/vir.0.040824-0
Keywords: interferons, innate immunity, antivirals, viruses, avian, interferon-stimulated genes
Citation: Santhakumar D, Rubbenstroth D, Martinez-Sobrido L and Munir M (2017) Avian Interferons and Their Antiviral Effectors. Front. Immunol. 8:49. doi: 10.3389/fimmu.2017.00049
Received: 18 October 2016; Accepted: 12 January 2017;
Published: 31 January 2017
Edited by:
Claudia U. Duerr, McGill University, CanadaReviewed by:
Jiuzhou Song, University of Maryland, USAHua-Ji Qiu, Chinese Academy of Agricultural Sciences, China
Copyright: © 2017 Santhakumar, Rubbenstroth, Martinez-Sobrido and Munir. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Muhammad Munir, muhammad.munir@pirbright.ac.uk, drmunir.muhammad@gmail.com
 Diwakar Santhakumar
Diwakar Santhakumar Dennis Rubbenstroth2
Dennis Rubbenstroth2 Luis Martinez-Sobrido
Luis Martinez-Sobrido Muhammad Munir
Muhammad Munir