- 1Pacific Northwest National Laboratory, Biological Sciences Division, Richland, WA, USA
- 2Department of Microbiology, School of Earth Sciences, The Ohio State University, Columbus, OH, USA
- 3Environmental and Molecular Sciences Laboratory, Pacific Northwest National Laboratory, Richland, WA, USA
Geologic carbon dioxide (CO2) sequestration drives physical and geochemical changes in deep subsurface environments that impact indigenous microbial activities. The combined effects of pressurized CO2 on a model sulfate-reducing microorganism, Desulfovibrio vulgaris, have been assessed using a suite of genomic and kinetic measurements. Novel high-pressure NMR time-series measurements using 13C-lactate were used to track D. vulgaris metabolism. We identified cessation of respiration at CO2 pressures of 10 bar, 25 bar, 50 bar, and 80 bar. Concurrent experiments using N2 as the pressurizing phase had no negative effect on microbial respiration, as inferred from reduction of sulfate to sulfide. Complementary pressurized batch incubations and fluorescence microscopy measurements supported NMR observations, and indicated that non-respiring cells were mostly viable at 50 bar CO2 for at least 4 h, and at 80 bar CO2 for 2 h. The fraction of dead cells increased rapidly after 4 h at 80 bar CO2. Transcriptomic (RNA-Seq) measurements on mRNA transcripts from CO2-incubated biomass indicated that cells up-regulated the production of certain amino acids (leucine, isoleucine) following CO2 exposure at elevated pressures, likely as part of a general stress response. Evidence for other poorly understood stress responses were also identified within RNA-Seq data, suggesting that while pressurized CO2 severely limits the growth and respiration of D. vulgaris cells, biomass retains intact cell membranes at pressures up to 80 bar CO2. Together, these data show that geologic sequestration of CO2 may have significant impacts on rates of sulfate reduction in many deep subsurface environments where this metabolism is a key respiratory process.
Introduction
The injection of CO2 gas into deep subsurface environments has emerged as one possible process to reduce anthropogenic greenhouse gas (GHG) emissions to the atmosphere (Benson and Cole, 2008). This technology involves the pumping of CO2 into suitable geologic environments such as depleted gas and oil fields, saline aquifers, and basalt formations (Holloway, 2005; McGrail et al., 2006). In many of these environments, elevated pressures and temperatures result in a CO2 phase change, from a gaseous to a supercritical state. Within deep subsurface environments, supercritical CO2 (scCO2) can undergo dissolution and disassociation reactions in formation fluids, resulting in pH decreases between 1.5 and 4 units (Mitchell et al., 2010). This pH change can subsequently cause mineral dissolution (and pH buffering) and re-precipitation reactions; the formation of new carbonate minerals may sequester the injected carbon over extended time periods (Krevor and Lackner, 2011; Qafoku et al., 2014).
A significant fraction of microbial biomass on Earth is present in the deep subsurface (Whitman et al., 1998; Kallmeyer et al., 2012; McMahon and Parnell, 2014), in many terrestrial and oceanic locations that could potentially be used for CO2 sequestration (Stevens and McKinley, 1995; Krumholz, 2000; Basso et al., 2009; Edwards et al., 2012). scCO2 has been shown to significantly reduce bacterial membrane potential and membrane integrity (Tamburini, 2013), and is commercially used as a microbial sterilizing agent in the food industry (Amanatidou et al., 1999). The injection of scCO2 into the subsurface could therefore potentially kill a significant fraction of biomass in the storage formation (White et al., 2006). However, the heterogeneity of the subsurface may provide microenvironments where less harsh conditions can be found; indeed, a field test site in Ketzin, Germany reported active populations of sulfate-reducing bacteria (SRB) and methanogenic Archaea following a CO2 injection (Morozova et al., 2010). Similarly, studies in an Australian sandstone aquifer revealed increases in Comamonadaceae and Sphingomonadaceae following CO2 injection (Mu et al., 2014). The activity of such microbial populations in subsurface regions receiving injected CO2 has both beneficial and deleterious implications for the efficiency of this process. Biofilm formation might act to block small fractures and pores, preventing upward migration of CO2 into overlying aquifers (Mitchell et al., 2009), while microbially-enhanced mineral precipitation could accelerate rates of CO2 incorporation into insoluble phases (Mitchell et al., 2010). Conversely, microbial growth around injection wells can cause localized clogging (Morozova et al., 2010), corrosion (Pitonzo et al., 2004), and reductions in injection rates, while the potential for certain microbial strains to metabolize CO2 and H2 to methane, a more potent GHG than CO2, remains poorly understood under relevant conditions.
Although in situ microbial activity has been reported following CO2 injection, little is known about how CO2 injection and possible upward migration of CO2 into overlying formations can affect microbial physiology, metabolism, and stress responses. Studies on a range of different microorganisms have reported reductions in growth rates and cell yields associated with increasing CO2 exposures, while a combination of increased CO2 at high pressures caused large reductions in viable cells (Coyne, 1933; Eklund, 1984; Wu et al., 2010; Schulz et al., 2012). Mitchell et al. (2008) reported that biomass present in a biofilm were more resistant to scCO2 than similar cells in a planktonic state, and hypothesized that differences in cell viability might be due to protective effects of extracellular polymeric substances (EPS) that accumulate within a biofilm. However, despite these studies there is currently little understanding of the stress response mechanisms that may be utilized by microbial populations that are suddenly exposed to CO2 at elevated pressures. Sulfate reducing bacteria (SRB) inhabit a wide range of diverse ecosystems, under conditions that are frequently considered to be “extreme” for microbial growth, including the deep subsurface in terrestrial and ocean environments (Bale et al., 1997; Sass and Cypionka, 2004; Basso et al., 2005; Klouche et al., 2009). Here we exposed a model subsurface SRB species, Desulfovibrio vulgaris, to CO2 under a range of pressures, using a novel combination of high-pressure nuclear magnetic resonance spectroscopy (HP-NMR) and RNA-Seq to track both physiological and transcriptional responses to CO2 stresses that may occur in deep subsurface injection sites.
Materials and Methods
Bacterial Growth
Desulfovibrio vulgaris str. Hildenborough (ATCC 29579) was obtained from the ATCC (Manassas, VA, USA), and grown in a defined freshwater media containing lactate (20 mM) and sulfate (20 mM) as electron donor and acceptor, respectively (Lovley et al., 1993) at pH 7.2. HEPES was added at 50 mM as a buffering agent. Cells were grown under strict anaerobic conditions at 30°C with a 100% N2 gas headspace. This strain was used as a representative for heterotrophic SRB in the environment.
CO2 Exposure Experiments
D. vulgaris was grown overnight for 20 h at 30°C to a density of OD600 ~ 0.5. Growth culture (50 mL) was harvested in an anaerobic glove box (M Braun, Garching, Germany) via centrifugation at 7000 × g for 10 min, and subsequently re-suspended in 65 ml fresh media to an OD600 cell density of ~0.2. For CO2 and N2 exposure experiments at atmospheric pressure, 160 mL serum bottles were sealed with thick rubber butyl stoppers. The headspace of these bottles was flushed with laboratory-grade CO2 or N2 for 15 min, before being incubated at 37°C. For exposure studies at elevated pressures, re-suspended biomass was added to a custom-built Parr high-pressure incubation vessel (Parr Instrument Company, Moline, IL, USA) within an anaerobic glove box (M Braun). The sealed pressure vessel was removed from the glove box and connected to a high-pressure syringe pump (Teledyne Isco, Lincoln, NE, USA), and pressurized to a desired pressure using CO2 or N2 gas. The pressure vessel was incubated at 37°C using a warming sleeve (Parr). Experiments were carried out under both N2 or CO2 at atmospheric (1 bar), 10 bar, 25 bar, 50 bar, and 80 bar pressures. pH was monitored during experiments via an in situ high pressure pH probe (Endress & Hauser, Greenwood, IN, USA). All experiments were performed in triplicate.
Sulfide and Cell Growth Measurements
During incubation experiments, samples were removed from the pressure vessel via a sampling dip tube, while maintaining pressurized conditions. Samples (1 mL) were removed at 0, 1, 2, 3, and 4 h following pressurization and analyzed for aqueous sulfide measurements using the Methylene Blue assay (Hach, Loveland, CO, USA). Sulfide concentrations were measured at 665 nm using a Shimadzu Biospec-1601 spectrophotometer (Shimadzu, Columbia, MD, USA) and absorbance intensities were compared against a standard curve generated from known sulfide concentrations. Cell densities were also measured using the Shimadzu Biospec-1601 spectrophotometer at 600 nm. Estimates of live cells were obtained using a Molecular Probes Live/Dead BacLight Bacterial Viability Kit, according to manufacturer's instructions (Life Technologies, Grand Island, NY, USA). Cell counts were performed using a Nikon 720 Light Microscope with an attached Nikon Super High Pressure mercury lamp, and a fluorescence cube. The results from eight randomized field of view counts were averaged and the Dead/Live ratio determined.
Transmission Electron Microscopy
To image CO2-impacted cells, a 5 μ L drop of planktonic D. vulgaris cell suspension was applied to a 100 mesh Cu grid covered with formvar support film sputtered with carbon (Electron Microscopy Sciences, Hatfield, PA, USA). The cells were allowed to adhere to the grids for 1 min before being blotted with a filter paper, and negatively stained with a 5 uL drop of NANO-W™ negative stain (Nanoprobes, Yaphank, NY, USA). After 30 s, the excess liquid was removed by wicking, and the sample was allowed to air dry. Whole mount samples were examined with a Tecnai T-12 transmission electron microscope (TEM) (FEI Co., Hillsboro, OR, USA) operating at 120 kV with a LaB6 filament. Images were collected digitally with a 2 × 2K Ultrascan 1000 charge-coupled device (Gatan, Pleasanton, CA, USA). Representative images of at least 50 cells per each sample were collected at 10,000× magnification and analyzed to distinguish the presence or absence of cell ultrastructural change.
EPS Extraction and Characterization
EPS isolation from D. vulgaris biomass was performed using a modified ethylenediaminetetraacetic acid (EDTA) treatment designed to extract EPS from biofilm communities (Eboigbodin and Biggs, 2008; Cao et al., 2011). Samples were collected at 0 and 4 h time points at 50 bar CO2 pressure. Biomass from 50 mL of culture was centrifuged at 5000 × g for 20 min at 4°C. The supernatant was discarded and the pellet was resuspended with 10 mL of sterilized 0.9% NaCl solution and centrifuged at 5000 × g for 20 min at 4°C. The supernatant was collected and defined as a “loosely associated EPS” (laEPS) fraction. The process of washing and laEPS collection was repeated three times and the remaining biomass was resuspended in 1% EDTA in 0.9% NaCl to obtain 5 mg biomass/mL of solution. This mixture was incubated at 4°C for 3 h. After incubation, the mixture was centrifuged at 5000 × g for 20 min at 4°C, and the supernatant collected as a “bound EPS” (bEPS) fraction. The resulting pellet was washed three times by resuspension in 10 mL 0.9% NaCl solution and the supernatant was added to the bEPS fraction. Both EPS fractions were clarified using a 0.22 μm filter and kept at 4°C until concentration. EPS fractions were concentrated and washed using Amicon Ultra Centrifugal Filters (Millipore, Billerica, MA, USA) with a molecular weight cut off of 3000 Da. The EPS fractions were reduced to ~3 mL, and were placed separately into Slide-A-Lyzer Dialysis Cassettes (Thermo Fisher Scientific Inc., Rockford, IL, USA) with a molecular weight cut off of 3500 Da. The dialysis cassettes were placed into a 4 L beaker of ultrapure water and were allowed to spin with a stir bar for 16 h at 4°C. After the overnight dialysis, samples were removed from the cassettes and concentrated to ~1 mL. These samples were flash frozen in liquid N2 and lyophilized overnight. For analysis, the lyophilized samples were resuspended in sterile ultrapure water and measured using ATR-FTIR (Attenuated Total Reflectance Fourier Transform Infrared Spectroscopy) using a Bruker Vector 22 FTIR spectrometer (Bruker Optics Inc., Billerica, MA, USA). Resulting data was baseline corrected and normalized to the 1045 cm−1 sugar peak using IGOR Pro version 6.3.1.2 (WaveMetrics, Inc., Lake Oswego, OR, USA).
RNA-Seq Library Preparation and Sequencing
For RNA-Seq measurements, biomass was incubated under CO2 pressurized conditions (1, 25, 50, and 80 bar) for 1 h at 37°C. Material was quickly sampled, centrifuged at 7000 × g for 10 min at 4°C, and the resulting cell pellet flash frozen in liquid N2. RNA was extracted from biomass using Invitrogen TRIzol® reagent, followed by genomic DNA removal and cleaning using the Qiagen RNase-Free DNase Set kit and Qiagen Mini RNeasy™ kit. An Agilent 2100 Bioanalyzer (Agilent Technologies, Inc., Santa Clara, CA, USA) was used to assess the integrity of the RNA samples. Only RNA samples having an RNA Integrity Number between 8 and 10 were selected for downstream analyses. The Applied Biosystems SOLiD™ Total RNA-Seq kit was used to generate the cDNA template library. The SOLiD™ EZ Bead system (Life Technologies, Grand Island, NY, USA) was used to perform emulsion clonal bead amplification to generate bead templates for SOLiD™ platform sequencing. Samples were sequenced on the 5500XL SOLiD™ platform. The 50-base short read sequences produced by the 5500XL SOLiD™ sequencer were mapped in color space using SOLiD™ LifeScope™ software version 2.5 using default parameters against the D. vulgaris str. Hildenborough reference genome. A second mapping step removed reads with low quality value scores (<20) to generate output files for downstream analyses. Normalized data as RPKM (Reads per Kilobase per Million mapped) values were used for downstream analyses. Genes were classified as being up- or down-regulated via One-Way ANOVA analysis to identify differences in means across the different conditions. Results were tested via the Tukey HSD test with a Benjamini-Hochberg multiple hypothesis correction to identify differences with p < 0.05.
NMR Methods
D. vulgaris biomass was prepared as for the batch kinetic experiments, with the following modification: washed cell suspensions were supplemented with 3 mM 13C-lactate (99% uniformed labeled 13C, Sigma-Aldrich, St. Louis, MO, USA) as an electron and carbon source. An 800 μ L aliquot of cell suspension was added to an appropriate high-pressure NMR tube or NMR rotor in an M Braun anaerobic glove bag (100% N2 atmosphere). High pressure tubes were either 5 mm heavy-walled precision pressure valve NMR tubes (Wilmad, Vineland, NJ, USA) for the 1H observed NMR experiments (up to 25 bar), or specialized custom-made 7.5 mm NMR rotors described previously (Hoyt et al., 2011; Turcu et al., 2013) for experiments up to 80 bar for the 13C NMR experiments. Experiments were carried similarly as above using laboratory grade N2, CO2, or Carbon-13C dioxide (99%-labeled, Sigma-Aldrich- Isotech, Miamisburg, OH) at atmospheric (1 bar), 5 bar, 10 bar, 15 bar, 25 bar, and 80 bar pressures. All experiments were carried out in duplicate. The sealed pressure vessels once removed from the glove box, were connected to a high-pressure syringe pump system (Teledyne Isco), and pressurized to a desired pressure using CO2 or N2 gas. The pressure tubes or rotors were incubated at 37°C ± 0.5°C using a Blue-M lab oven (Thermal Product Solutions, White Deer, PA) and equilibrated 15 min after gas was brought to the desired pressure. Rotors were mechanically manipulated in a loading chamber/safety cell as previously described (Turcu et al., 2013). After samples were pressurized to temperature and pressure, they were transferred using a safety cell to the NMR instrument and immediately equilibrated at 37°C in the NMR probe.
1H observed NMR data was acquired on a Varian Direct Drive (VNMRS) 600 MHz spectrometer (Agilent Technologies) equipped with a Dell PrecisionT3500 Linux workstation running VNMRJ4.0. The spectrometer system was outfitted with a Varian z-gradient triple resonance HCN probe. A time series of short (every 15 min) 2-D gChsqc spectra (1H detect and 13C-filtered) were collected with a sweep width of 12 ppm (1H) in the direct detect dimension with 2048 complex points and 80 ppm (13C) in the indirect dimension with 32 increments and acquisition time of 0.285 s per scan. 1H 1-dimensional 1D) NMR spectra were derived from the first increments of these 2-D experiments. The 1D 1H NMR spectra were the resultant averaged free-induction decays (FIDs) of 8 scans with a 1 s recycle delay between scans. Data were processed and analyzed using VNMRJ 4.0 software and 1H spectra with 2048 complex points were processed with zero filling to digitize 4K points and using a sine-bell squared apodization window prior to Fourier Transform (FT). Spectral peak positions of the lactate and acetate peaks were externally referenced relative to a 0.5 mM 2,2-dimethyl- 2-silapentane-5-sulfonate (DSS) (Chenomx, Edmonton, Alberta, Canada) standard. During D. vulgaris growth experiments in CO2 or N2 at various pressures, protons attached to the C2 methylene and C3 methyl of lactate show 1H resonances disappearing at ~4.08 and ~1.30 ppm, respectively, while the C2 methyl of acetate shows a 1H resonance emerging over time as lactate is oxidized to acetate.
All 13C NMR measurements were performed on an Agilent-Varian 300 MHz VNMRS spectrometer (H0 = 7.05 T) at 75.44 MHz Larmor frequency using a direct polarization Bloch-decay sequence with 31.2 kHz proton decoupling for 300 ms. All 13C experiments employed a pulse width of 2 μ s (45° flip angle) and used a relaxation delay of 5 s to acquire 120 transients. Spectra are referenced with respect to TMS via a secondary adamantane standard (37.85 ppm). The 13C spectral width was 50 kHz and 15,008 data points were acquired per transient. The 13C NMR spectra were zero filled to a Fourier number of 32,000 points and given the equivalent of 40 Hz exponential apodization prior to FT. 13C resonances for lactate were observed at 185, 74, and 28 ppm for the C1, C2, and C3 carbons, respectively, and for acetate at 184 and 30 ppm for the C1 and C2 carbons, respectively.
Results
In this study, metabolic and physiological indicators were monitored during exposure of an actively growing SRB model organism, D. vulgaris, to CO2 over a range of environmentally relevant pressures. Using a high-pressure batch incubation reactor, biomass was incubated under CO2 at atmospheric, 10 bar, 25 bar, 50 bar, and 80 bar pressures at 37°C, with samples recovered for kinetic, microscopic, and transcriptomic analyses. High-pressure NMR techniques were applied concurrently to measure microbial oxidation of lactate under similar pressure and temperature conditions.
Despite the use of HEPES-buffered media, incubation under CO2 (≥5 bar) resulted in pH decreases over the 4 h incubation period from 7.2 to values ranging from 5.5 to 6.5. Atmospheric pressure incubations at pH 6 (N2 headspace) indicated that the decrease in media pH had little effect on cell growth or sulfide production (data not shown). CO2 exposure had a dramatic effect on cell respiration however, as inferred from sulfide measurements, cell density measurements (Figures 1A,B), and time-course NMR data collection (Figure 2).
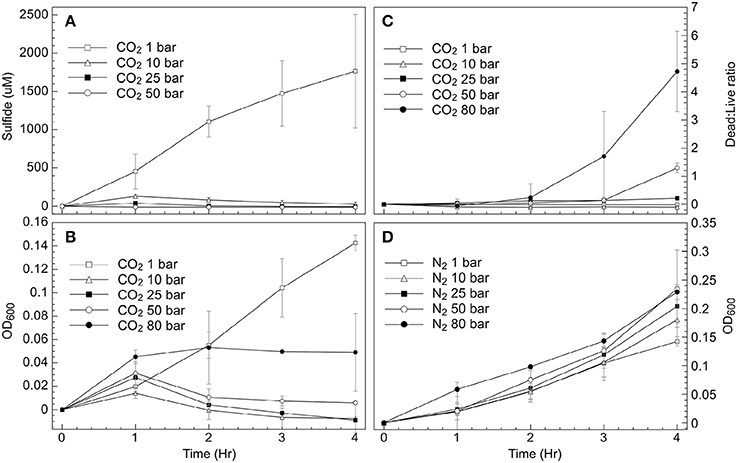
Figure 1. Exposure of D. vulgaris biomass to CO2 or N2 at a series of pressures. Values show changes from measurements made at T = 0 (A) Sulfide concentrations under CO2; (B) Optical density (OD600) under CO2 headspace; (C) Live:Dead cell ratios under CO2; (D) Optical density (OD600) under N2 headspace.
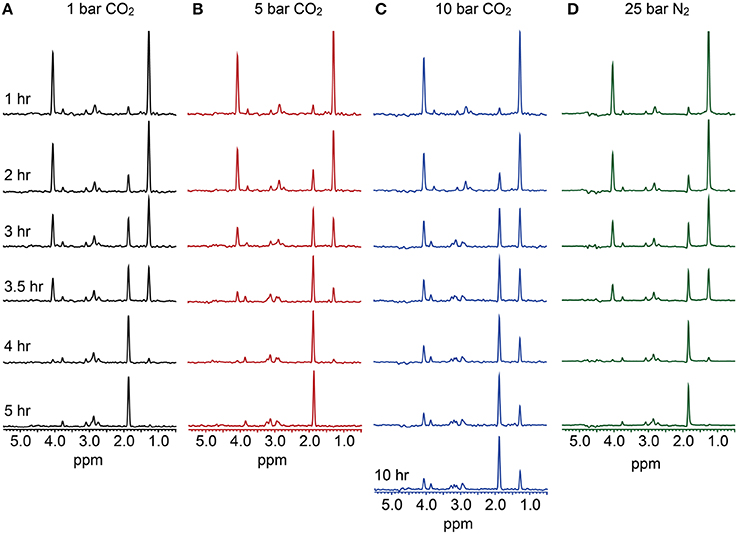
Figure 2. Carbon (CO2 as the pressurizing phase) and proton (N2 as the pressurizing phase) NMR data. Oxidation of 13C-3-lactate to acetate was monitored via the disappearance of lactate peaks at 4.04 and 1.26 ppm, which represent the lactate C-H and C-H3 bonds, respectively. Concurrent to this, the peak appearing at 1.84 ppm represents the acetate CH3 bond. Data indicates that at 1 bar CO2 (A), 5 bar CO2 (B), and 25 bar N2 (D), complete conversion of lactate to acetate occurs over a 4–5 h. period. At 10 bar CO2 (C) however, incomplete lactate oxidation is observed, supporting observations made in high-pressure CO2 batch experiments.
Sulfide generation was greatly reduced at CO2 pressures above atmosphere (1 bar), with any S2− increases only observed over the first hour of exposure to 10, 25, 50, or 80 bar CO2. By contrast, sulfide was consistently generated over the 4-h measurement period under CO2 at atmospheric pressure (Figure 1A). These trends were mirrored when measuring cell biomass increases via OD600 readings (Figure 1B), with consistent increases in cell density observed at atmospheric pressure under a CO2 headspace. At elevated CO2 pressures however (≥10 bar), small increases in cell density were generally only observed during the first hour (Figure 1B), with cessation of growth at later time periods. No additional cell growth was observed in these experiments over longer time periods (~48 h) (data not shown). Biomass samples were also recovered at each time point and stained for cell-viability measurements using live-dead cell staining techniques. These data indicated that little cell death occurred at 10 bar and 25 bar CO2 pressures during the 4 h incubation period (Figure 1C). Similar trends were initially observed at 50 bar and 80 bar CO2. Cell viability began to decrease in these samples at later time points (>3 h), with the greatest proportion of cells with compromised membranes measured after 4 h incubation at 80 bar CO2 (Figure 1C). To decouple the effects of elevated pressure and CO2 stress, parallel incubations were carried out using N2 gas as the pressurizing phase rather than CO2. These experiments were carried out at pH 6 to mirror the pH change associated with CO2 exposure. Results from these incubations demonstrated that pressure alone did not impact D. vulgaris activity. The growth rate in cultures exposed to 80 bar N2 (μ = 0.19 h−1) was even greater than the growth rate measured in cultures exposed to 1 bar CO2 (μ = 0.13 h−1) during the 4-h incubation period (Figures 1A,D).
13-Carbon-NMR was used to track the oxidation of 13C-3-lactate, and formation of 13C-2-acetate by D. vulgaris across a range of CO2 pressures at 37°C. At 10 bar CO2, incomplete oxidation of lactate was observed, indicating inhibition of microbial respiration and supporting observations made in batch reactions (Figure 2). Similar observations were made via NMR at pressures of 25, 50, and 80 bar CO2. However, the complete conversion of lactate to acetate was observed at both 1 and 5 bar CO2 over similar reaction times, suggesting that some exposure to increased CO2 pressures could be tolerated by D. vulgaris (Figure 2). In contrast, 1-H proton NMR measurements revealed complete lactate oxidation to acetate by D. vulgaris under 25 bar N2. These data support batch incubation results that indicated no negative effects of pressure alone on growth rate (Figure 2).
Biomass samples taken at 0 h and 4 h time points during 50 bar CO2 exposure were imaged using TEM. Initial TEM images showed healthy D. vulgaris cells at the beginning of the experiment (Figures 3A–C). After 4 h incubation under 50 bar CO2, there was visual evidence for compromised cells, as observed by increasingly shriveled cell membranes (Figures 3D–F). While there are sometimes artifacts from negative strain preparations, here the images may indicate the secretion of large quantities of EPS around cells following 50 bar CO2 incubations (Figures 3D–F). To investigate the EPS chemistry, we modified a biofilm EPS extraction protocol to isolate two “operational” EPS fractions, laEPS and bEPS, from the 0 and 4 h time points at 50 bar CO2 for chemical characterization. EPS yields were low from all samples. This was not unexpected since biomass in the high pressure batch reactor were generally planktonic. Interestingly, ATR-FTIR spectroscopy did indicate chemical changes in lipid chemistry of the bEPS (EDTA-extractable) fraction after 4 h exposure to CO2 (Figure 4). Predominant peaks representing CH2- stretches of lipids were observed at 2846 and 2915 wavenumber (cm−1) in the 0 h exposure. After the 4 h CO2 exposure, these peaks were replaced with CH3- lipid peaks at 2888 and 2944 wavenumber (cm−1). No chemical differences could be conclusively identified within other classes of biomolecules (proteins, polysaccharides, nucleic acids) in the EPS fractions using ATR-FTIR spectroscopy (not shown).
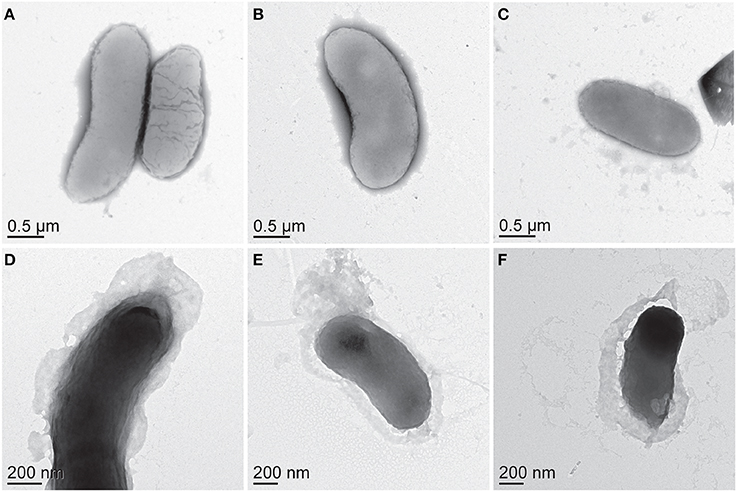
Figure 3. Negatively stained, whole mount TEM images of D. vulgaris biomass (A–C) before, and (D–F) after 4 h exposure to CO2 at 50 bar.
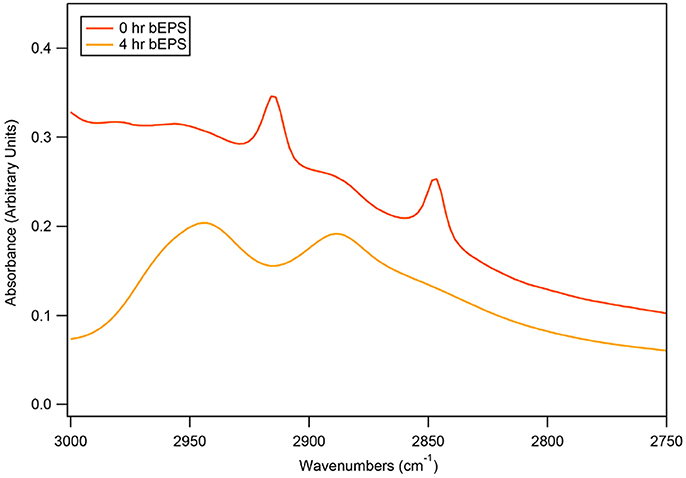
Figure 4. Comparisons of bEPS chemistry produced by D. vulgaris biomass after 0 or 4 h exposure to 50 bar of CO2. EPS was extracted and compared using ATR-FTIR spectroscopy. Ten-minute scans were obtained, baseline corrected, and normalized to 1045 cm−1 sugar peak.
Coupled to these physiological measurements, RNA was extracted from D. vulgaris biomass that had been exposed to CO2 at atmospheric pressure, 25 bar, 50 bar, and 80 bar. Duplicate sets of biomass were generated for each condition. Following conversion of the extracted RNA to cDNA, this material was sequenced using the SOLiD platform. Sequenced reads were mapped to the D. vulgaris genome to identify gene expression patterns characteristic of a stress response. Despite the measurement of only limited metabolic activity at elevated pressures, shifts in transcript abundance were observed across these conditions, indicating that cells were responding to CO2 stresses (Supplementary Material). Genes associated with chemotaxis (flagella subunits, methyl-accepting chemotaxis proteins) were clearly up-regulated across all CO2 exposures, suggesting that cells were attempting to find more suitable environmental conditions (Table 1). However, more genes associated with two-component environmental sensing systems were down-regulated across the same set of conditions (Table 1). The regulation and expression of certain amino acids appears to play a key role in the D. vulgaris stress response to pressurized CO2 conditions; relative to abundances at atmospheric pressure, transcripts for leucyl and isoleucyl-tRNA synthetases (DVU1196; DVU1927) were both up-regulated at 25 bar and 50 bar CO2 pressures, while expression of a tRNA modification GTPase (TrmE; DVU1079) exhibited a similar pattern. Further evidence for a significant amino-acid based response to these stresses was inferred from genes associated with leucine biosynthesis. In addition to the leucyl-tRNA synthetase, transcripts from a 2-isopropylmalate synthase (DVU2981), and both the large and small subunits of a 3-isopropylmalate dehydratase (DVU2982 and DVU2983, respectively) were similarly up-regulated. This pattern was maintained in the 50 bar and 80 bar CO2 exposures (Figure 5). Other general stress responses were observed during CO2 exposure at 25 bar. The phage shock protein (Psp) system responds to extracytoplasmic stresses, and is induced by dissipation of proton motive force (PMF) in the periplasm (Darwin, 2005). However, although PsPA is the most abundant protein upon induction of the PsP system, little is currently known as to its physiological function. PspA was up-regulated at both 25 bar and 50 bar CO2 exposures, possibly suggesting a loss of membrane integrity in some cells. In addition, a periplasmic Fe-hydrogenase that has been shown to play a key role in protecting against oxidative stress (Fournier et al., 2004) was up-regulated over the same conditions. Interestingly, the transcription of carbon starvation protein A (CstA) was up regulated under all pressurized conditions; given that lactate was provided in excess as a carbon and electron source, the cstA gene may play a role in a more generalized stress response. The role of hypothetical genes in many cellular functions was also emphasized via transcriptomic results. Across all three pressurized conditions approximately 25% of up- and down-regulated genes were hypothetical, hinting at unknown stress mechanisms and responses to CO2 exposure.
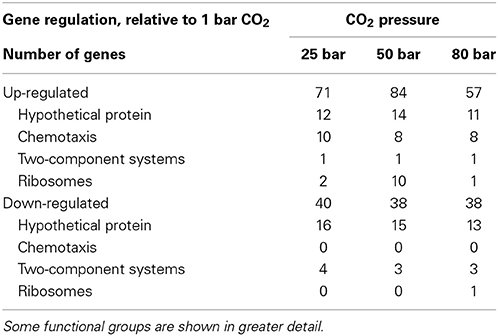
Table 1. Numbers of statistically significant up- and down-regulated genes across the CO2 exposure conditions.
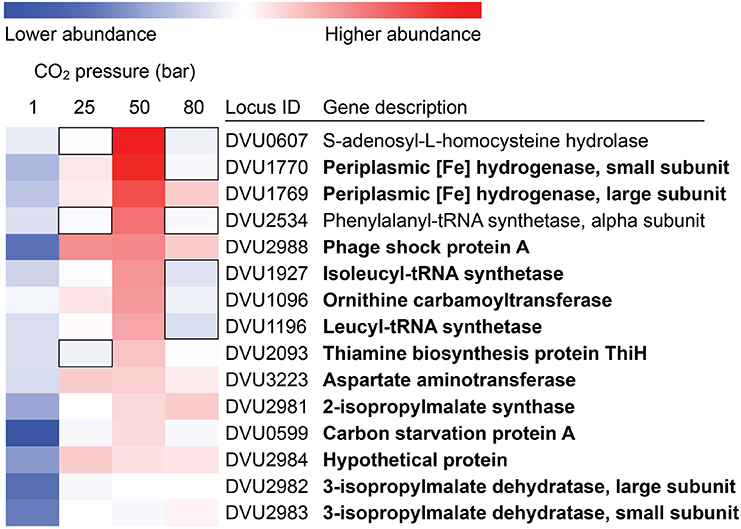
Figure 5. Expression profiles for up-regulated mRNA gene transcripts under CO2 pressurized conditions (25, 50, and 80 bar), relative to 1 bar CO2. Bolded gene descriptions show genes that were up-regulated (p < 0.05) under multiple conditions. Conversely, boxed areas show genes that had no statistically significant changes under a certain condition.
Discussion
In this study, we illustrate the negative effects of CO2 exposure on microbial activity in the model organism D. vulgaris. As CO2 injection into subsurface systems occurs, microorganisms are exposed both within the reservoir, and in shallower regions via CO2 leakage and migration through cap rock (Harvey et al., 2012). Our findings indicate that even at relatively low pressures (10 bar) some microbial activity is significantly impacted by exposure to CO2. Growth of D. vulgaris is significantly limited under all pressurized CO2 conditions, and respiration (inferred from the measurements of microbial oxidation of lactate, and accumulation of sulfide) is similarly impacted. The rapid growth of D. vulgaris under N2 pressurized conditions (up to 80 bar) suggests that pressure itself has no negative impact on cell health. Instead, toxic effects from CO2 exposure linked to compromised membrane integrity would appear to be responsible for decreases in D. vulgaris activity in certain experiments. Kinetic measurements during pressurized CO2 exposure experiments suggest that some microbial activity persisted for approximately the first hour, before respiration and cell growth ceased. Despite this cessation of growth under lower pressure conditions (10 bar and 25 bar CO2), epifluorescence microscopy observations reveal that cells remained viable. In addition, the continued production of mRNA transcripts at all CO2 pressures studied here further supports the observation that D. vulgaris cells remain viable during CO2 exposures. These results support observations from environmental samples recovered from a saline aquifer in Germany, where SRB biomass remained viable for up to 30 days, despite the cessation of active sulfate reduction following CO2 injection (Frerichs et al., 2013).
Electron microscopy images suggest that EPS may be produced in response to CO2 exposure, potentially to buffer against deleterious effects. Given that previous studies have at least partly attributed increased resistance to CO2 toxicity in biofilm forming bacteria to greater EPS production (Mitchell et al., 2008, 2009; Wu et al., 2010), D. vulgaris may utilize a similar approach. ATR-FTIR measurements on EPS extracted from biomass exposed to CO2 showed little overall chemical changes in most classes of biomolecules except the lipid region. Notably, bEPS chemical shifts from CH2 to CH3 stretches may indicate that membrane lipids were being fragmented as a result of the CO2 exposure. The relatively short exposure (4 h) of the D. vulgaris cells to 50 bar CO2 may not have been adequate to completely access chemical changes in EPS composition, or the protective effect of biofilm formation under CO2 exposure conditions. In contrast, measuring gene expression may provide a better representation of the physiological responses to short-term 50 bar CO2 exposure. Gene transcript data indicates that D. vulgaris utilizes additional stress responses, including the accumulation of certain amino acids such as leucine. The accumulation of specific amino acids (in addition to other solutes) is a well-recognized osmotic stress response in many bacteria (Csonka, 1989); indeed, the up-regulation of several genes involved in leucine biosynthesis has been reported in response to KCl stress (Mukhopadhyay et al., 2006). Under salt stress conditions, the role of these amino acids is generally to buffer the cytoplasm and prevent hyperosmotic shock; glutamate and proline are two of the more well studied examples (Csonka et al., 1994; Amin et al., 1995). A potential role for leucine/isoleucine in CO2 stress response is less clear, although these amino acids may be important components of other buffering or chaperone proteins, such as ferritin (Götze et al., 2014). Alternatively, amino acid production may be a more general stress response in this microorganism, stimulated via multiple factors. Likewise, phage shock proteins have been observed to increase in abundance under a range of stress conditions, including nitrate, heat, salt, and oxygen stresses (Chhabra et al., 2006; Mukhopadhyay et al., 2006, 2007; He et al., 2010). Previous studies with E. coli have identified increased bacterial motility in response to low pH (Soutourina et al., 2002). Although these experiments were all performed under similar pH conditions (~pH 6), CO2 stress may have induced greater expression of genes characteristic of motility in D. vulgaris. However, unlike nitrate stress, there was little evidence of CO2 stress resulting in up-regulation of genes associated with energy metabolism (He et al., 2010).
Using a model SRB system, we tested the combinations of stresses associated with deep subsurface CO2 injection (elevated CO2 concentrations, pH decreases, and high-pressure) to ascertain the impact of CO2 injection on the physiology and metabolism of indigenous deep-subsurface bacteria. We observed that high pressures (80 bar N2) had less inhibitory effects on cellular growth than CO2 exposures at relatively low pressures. Although some of the pressures used in this study are lower than those in deep subsurface systems, these results indicate that upward migration of CO2 from storage reservoirs would impact microbiology in shallower overlying materials (Harvey et al., 2012), with implications for carbon cycling and metal fate and transport. In addition, the potential for inhibition of sulfate reduction in the presence of pressurized CO2 may help to limit deleterious subsurface processes such as sulfide-induced corrosion and oil field souring in the subsurface. However, it is worth noting that active microbial processes have been observed in CO2 sequestration reservoirs (Morozova et al., 2010; Mu et al., 2014), likely catalyzed by microbial species that exhibit greater tolerance for scCO2. Given the results of this study, injected CO2 in subsurface systems is likely to act as a strong driving force for shifts in microbial community structure. The effects of these changes on ecosystem function are currently unclear. Therefore, having observed laboratory responses to CO2 stresses in a model microbial strain (D. vulgaris), future work will attempt to understand the mechanisms that indigenous microorganisms use for survival and growth in CO2-impacted deep subsurface environments, and the impact of microbial persistence on system biogeochemistry.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This research was supported by a Laboratory Directed Research Development (LDRD) grant to Michael J. Wilkins at Pacific Northwest National Laboratory (PNNL). EPS characterization studies were funded by a US DOE Office of Science Early Career Research Award (project no. 60385, Matthew J. Marshall). Portions of this work were performed under a Science Themed Proposal (proposal ID: 44725, Michael J. Wilkins) using EMSL, a national scientific user facility sponsored by DOE's Office of Biological and Environmental Research and located at PNNL. PNNL is operated by Battelle for the DOE under Contract DE-AC05-76RL01830. We thank Dr. Sara Mae Belchik for assistance in ATR-FTIR data analysis.
Supplementary Material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/journal/10.3389/fmicb.2014.00507/abstract
References
Amanatidou, A., Smid, E. J., and Gorris, L. G. M. (1999). Effect of elevated oxygen and carbon dioxide on the surface growth of vegetable-associated micro-organisms. J. Appl. Microbiol. 86, 429–438. doi: 10.1046/j.1365-2672.1999.00682.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Amin, U. S., Lash, T. D., and Wilkinson, B. J. (1995). Proline betaine is a highly effective osmoprotectant for Staphylococcus aureus. Arch. Microbiol. 163, 138–142. doi: 10.1007/BF00381788
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Bale, S. J., Goodman, K., Rochelle, P. A., Marchesi, J. R., Fry, J. C., Weightman, A. J., et al. (1997). Desulfovibrio profundus sp. nov., a novel barophilic sulfate-reducing bacterium from deep sediment layers in the Japan Sea. Int. J. Syst. Bacteriol. 47, 515–521.
Basso, O., Caumette, P., and Magot, M. (2005). Desultovibrio putealis sp. nov., a novel sulfate-reducing bacterium isolated from a deep subsurface aquifer. Int. J. Syst. Evol. Microbiol. 55, 101–104. doi: 10.1099/ijs.0.63303-0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Basso, O., Lascourreges, J. F., Le Borgne, F., Le Goff, C., and Magot, M. (2009). Characterization by culture and molecular analysis of the microbial diversity of a deep subsurface gas storage aquifer. Res. Microbiol. 160, 107–116. doi: 10.1016/j.resmic.2008.10.010
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Benson, S. M., and Cole, D. R. (2008). CO2 sequestration in deep sedimentary formations. Elements 4, 325–331. doi: 10.2113/gselements.4.5.325
Cao, B., Shi, L., Brown, R. N., Xiong, Y., Fredrickson, J. K., Romine, M. F., et al. (2011). Extracellular polymeric substances from Shewanella sp. HRCR-1 biofilms: characterization by infrared spectroscopy and proteomics. Environ. Microbiol. 13, 1018–1031. doi: 10.1111/j.1462-2920.2010.02407.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chhabra, S. R., He, Q., Huang, K. H., Gaucher, S. P., Alm, E. J., He, Z., et al. (2006). Global analysis of heat shock response in Desulfovibrio vulgaris Hildenborough. J. Bacteriol. 188, 1817–1828. doi: 10.1128/JB.188.5.1817-1828.2006
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Coyne, F. P. (1933). The effect of carbon dioxide on bacterial growth. Proc. R. Soc. Lond. B 113, 196–217. doi: 10.1098/rspb.1933.0041
Csonka, L. N. (1989). Physiological and genetic responses of bacteria to osmotic stress. Microbiol. Rev. 53, 121–147.
Csonka, L. N., Ikeda, T. P., Fletcher, S. A., and Kustu, S. (1994). The accumulation of glutamate is necessary for optimal growth of Salmonella typhimurium in media of high osmolality but not induction of the proU operon. J. Bacteriol. 176, 6324–6333.
Darwin, A. J. (2005). The phage-shock-protein response. Mol. Microbiol. 57, 621–628. doi: 10.1111/j.1365-2958.2005.04694.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Eboigbodin, K. E., and Biggs, C. A. (2008). Characterization of the extracellular polymeric substances produced by Escherichia coli using infrared spectroscopic, proteomic, and aggregation studies. Biomacromolecules 9, 686–695. doi: 10.1021/bm701043c
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Edwards, K. J., Becker, K., and Colwell, F. (2012). The deep, dark energy biosphere: intraterrestrial life on Earth. Annu. Rev. Earth Plant Sci. 40, 551–568. doi: 10.1146/annurev-earth-042711-105500
Eklund, T. (1984). The effect of carbon dioxide on bacterial growth and on uptake processes in bacterial membrane vesicles. Int. J. Food Microbiol. 1, 179–185. doi: 10.1016/0168-1605(84)90014-X
Fournier, M., Dermoun, Z., Durand, M.-C., and Dolla, A. (2004). A new function of the Desulfovibrio vulgaris Hildenborough [Fe] hydrogenase in the protection against oxidative stress. J. Biol. Chem. 279, 1787–1793. doi: 10.1074/jbc.M307965200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Frerichs, J., Rakoczy, J., Ostertag-Henning, C., and Krüger, M. (2013). Viability and adaptation potential of indigenous microorganisms from natural gas field fluids in high pressure incubations with supercritical CO2. Environ. Sci. Technol. 48, 1306–1314. doi: 10.1021/es4027985
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Götze, S., Matoo, O. B., Beniash, E., Saborowski, R., and Sokolova, I. M. (2014). Interactive effects of CO2 and trace metals on the proteasome activity and cellular stress response of marine bivalves Crassostrea virginica and Mercenaria mercenaria. Aquat. Toxicol. 149, 65–82. doi: 10.1016/j.aquatox.2014.01.027
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Harvey, O. R., Qafoku, N. P., Cantrell, K. J., Lee, G., Amonette, J. E., and Brown, C. F. (2012). Geochemical implications of gas leakage associated with geologic CO2 Storage—a qualitative review. Environ. Sci. Technol. 47, 23–36. doi: 10.1021/es3029457
He, Q., He, Z., Joyner, D. C., Joachimiak, M., Price, M. N., Yang, Z. K., et al. (2010). Impact of elevated nitrate on sulfate-reducing bacteria: a comparative Study of Desulfovibrio vulgaris. ISME J. 4, 1386–1397. doi: 10.1038/ismej.2010.59
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Holloway, S. (2005). Underground sequestration of carbon dioxide—a viable greenhouse gas mitigation option. Energy 30, 2318–2333. doi: 10.1016/j.energy.2003.10.023
Hoyt, D. W., Turcu, R. V. F., Sears, J. A., Rosso, K. M., Burton, S. D., Felmy, A. R., et al. (2011). High-pressure magic angle spinning nuclear magnetic resonance. J. Magn. Reson. 212, 378–385. doi: 10.1016/j.jmr.2011.07.019
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kallmeyer, J., Pockalny, R., Adhikari, R. R., Smith, D. C., and D'Hondt, S. (2012). Global distribution of microbial abundance and biomass in subseafloor sediment. Proc. Natl. Acad. Sci. U.S.A. 109, 16213–16216. doi: 10.1073/pnas.1203849109
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Klouche, N., Basso, O., Lascourreges, J. F., Cayol, J. L., Thomas, P., Fauque, G., et al. (2009). Desulfocurvus vexinensis gen. nov., sp nov., a sulfate-reducing bacterium isolated from a deep subsurface aquifer. Int. J. Syst. Evol. Microbiol. 59, 3100–3104. doi: 10.1099/ijs.0.010363-0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Krevor, S. C. M., and Lackner, K. S. (2011). Enhancing serpentine dissolution kinetics for mineral carbon dioxide sequestration. Int. J. Greenh. Gas Con. 5, 1073–1080. doi: 10.1016/j.ijggc.2011.01.006
Krumholz, L. (2000). Microbial communities in the deep subsurface. Hydrogeol. J. 8, 4–10. doi: 10.1007/s100400050003
Lovley, D., Giovannoni, S., White, D., Champine, J., Phillips, E., Gorby, Y., et al. (1993). Geobacter metallireducens gen. nov. sp. nov., a microorganism capable of coupling the complete oxidation of organic compounds to the reduction of iron and other metals. Arch. Microbiol. 159, 336–344. doi: 10.1007/BF00290916
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
McGrail, B. P., Schaef, H. T., Ho, A. M., Chien, Y.-J., Dooley, J. J., and Davidson, C. L. (2006). Potential for carbon dioxide sequestration in flood basalts. J. Geophys. Res. 111, B12201. doi: 10.1029/2005JB004169
McMahon, S., and Parnell, J. (2014). Weighing the deep continental biosphere. FEMS Microbiol. Ecol. 87, 113–120. doi: 10.1111/1574-6941.12196
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mitchell, A. C., Dideriksen, K., Spangler, L. H., Cunningham, A. B., and Gerlach, R. (2010). Microbially enhanced carbon capture and storage by mineral-trapping and solubility-trapping. Environ. Sci. Technol. 44, 5270–5276. doi: 10.1021/es903270w
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mitchell, A. C., Phillips, A. J., Hamilton, M. A., Gerlach, R., Hollis, W. K., Kaszuba, J. P., et al. (2008). Resilience of planktonic and biofilm cultures to supercritical CO2. J. Supercrit. Fluid. 47, 318–325. doi: 10.1016/j.supflu.2008.07.005
Mitchell, A. C., Phillips, A. J., Hiebert, R., Gerlach, R., Spangler, L. H., and Cunningham, A. B. (2009). Biofilm enhanced geologic sequestration of supercritical CO2. Int. J. Greenh. Gas Con. 3, 90–99. doi: 10.1016/j.ijggc.2008.05.002
Morozova, D., Wandrey, M., Alawi, M., Zimmer, M., Vieth, A., Zettlitzer, M., et al. (2010). Monitoring of the microbial community composition in saline aquifers during CO2 storage by fluorescence in situ hybridisation. Int. J. Greenh. Gas Con. 4, 981–989. doi: 10.1016/j.ijggc.2009.11.014
Mu, N., Boreham, C., Leong, H. X., Haese, R. R., and Moreau, J. W. (2014). Changes in the deep subsurface microbial biosphere resulting from a field-scale CO2 geosequestration experiment. Front. Microbiol. 5:209. doi: 10.3389/fmicb.2014.00209
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mukhopadhyay, A., He, Z., Alm, E. J., Arkin, A. P., Baidoo, E. E., Borglin, S. C., et al. (2006). Salt stress in Desulfovibrio vulgaris Hildenborough: an integrated genomics approach. J. Bacteriol. 188, 4068–4078. doi: 10.1128/JB.01921-05
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mukhopadhyay, A., Redding, A. M., Joachimiak, M. P., Arkin, A. P., Borglin, S. E., Dehal, P. S., et al. (2007). Cell-wide responses to low-oxygen exposure in Desulfovibrio vulgaris Hildenborough. J. Bacteriol. 189, 5996–6010. doi: 10.1128/JB.00368-07
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Pitonzo, B. J., Castro, P., Amy, P. S., Southam, G., Jones, D. A., and Ringelberg, D. (2004). Microbiologically influenced corrosion capability of bacteria isolated from Yucca Mountain. Corrosion 60, 64–74. doi: 10.5006/1.3299233
Qafoku, O., Hu, J., Hess, N. J., Hu, M. Y., Ilton, E. S., Feng, J., et al. (2014). Formation of submicron magnesite during reaction of natural forsterite in H2O-saturated supercritical CO2. Geochim. Cosmochim. Acta 134, 197–209. doi: 10.1016/j.gca.2013.09.024
Sass, H., and Cypionka, H. (2004). Isolation of sulfate-reducing bacteria from the terrestrial deep subsurface and description of Desulfovibrio cavernae sp. nov. Syst. Appl. Microbiol. 27, 541–548. doi: 10.1078/0723202041748181
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Schulz, A., Vogt, C., and Richnow, H.-H. (2012). Effects of high CO2 concentrations on ecophysiologically different microorganisms. Environ. Pollut. 169, 27–34. doi: 10.1016/j.envpol.2012.05.010
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Soutourina, O. A., Krin, E., Laurent-Winter, C., Hommais, F., Danchin, A., and Bertin, P. N. (2002). Regulation of bacterial motility in response to low pH in Escherichia coli: the role of H-NS protein. Microbiology 148, 1543–1551.
Stevens, T. O., and McKinley, J. P. (1995). Lithoautotrophic microbial ecosystems in deep basalt aquifers. Science 270, 450–455. doi: 10.1126/science.270.5235.450
Tamburini, S. (2013). Molecular and Cellular Effects of Supercritical Carbon Dioxide on Some Important Food-Borne Pathogens. Ph.D, University of Trento.
Turcu, R. V. F., Hoyt, D. W., Rosso, K. M., Sears, J. A., Loring, J. S., Felmy, A. R., et al. (2013). Rotor design for high pressure magic angle spinning nuclear magnetic resonance. J. Magn. Reson. 226, 64–69. doi: 10.1016/j.jmr.2012.08.009
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
White, A., Burns, D., and Christensen, T. W. (2006). Effective terminal sterilization using supercritical carbon dioxide. J. Biotechnol. 123, 504–515. doi: 10.1016/j.jbiotec.2005.12.033
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Whitman, W. B., Coleman, D. C., and Wiebe, W. J. (1998). Prokaryotes: the unseen majority. Proc. Natl. Acad. Sci. U.S.A. 95, 6578–6583. doi: 10.1073/pnas.95.12.6578
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Wu, B., Shao, H., Wang, Z., Hu, Y., Tang, Y. J., and Jun, Y.-S. (2010). Viability and Metal Reduction of Shewanella oneidensis MR-1 under CO2 stress: implications for ecological effects of CO2 leakage from geologic CO2 sequestration. Environ. Sci. Technol. 44, 9213–9218. doi: 10.1021/es102299j
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Keywords: Desulfovibrio vulgaris, geologic CO2 sequestration, RNA-Seq, NMR spectroscopy, microbial stress
Citation: Wilkins MJ, Hoyt DW, Marshall MJ, Alderson PA, Plymale AE, Markillie LM, Tucker AE, Walter ED, Linggi BE, Dohnalkova AC and Taylor RC (2014) CO2 exposure at pressure impacts metabolism and stress responses in the model sulfate-reducing bacterium Desulfovibrio vulgaris strain Hildenborough. Front. Microbiol. 5:507. doi: 10.3389/fmicb.2014.00507
Received: 27 June 2014; Accepted: 09 September 2014;
Published online: 25 September 2014.
Edited by:
Dawn Cardace, University of Rhode Island, USAReviewed by:
John Stolz, Duquesne University, USAWilliam D. Orsi, Woods Hole Oceanographic Institution, USA
Copyright © 2014 Wilkins, Hoyt, Marshall, Alderson, Plymale, Markillie, Tucker, Walter, Linggi, Dohnalkova and Taylor. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Michael J. Wilkins, Department of Microbiology, School of Earth Sciences, The Ohio State University, 315 Mendenhall Laboratory, South Oval Mall, Columbus, OH 43210, USA e-mail: wilkins.231@osu.edu
†Present address: Bryan E. Linggi, Janssen R&D, LLC, San Diego, USA
 Michael J. Wilkins
Michael J. Wilkins David W. Hoyt3
David W. Hoyt3 Matthew J. Marshall
Matthew J. Marshall Alice C. Dohnalkova
Alice C. Dohnalkova