- 1Department of Life Sciences, Graduate School of Arts and Sciences, University of Tokyo, Tokyo, Japan
- 2Japan Science and Technology Agency, CREST, Tokyo, Japan
- 3Department of Biological Science, Faculty of Science, and Research Institute of Electronics, Shizuoka University, Shizuoka, Japan
Cyanobacteria and chloroplasts perform oxygenic photosynthesis, and share a common origin. Galactolipids are present in the photosynthetic membranes of both cyanobacteria and chloroplasts, but the biosynthetic pathways of the galactolipids are significantly different in the two systems. In this minireview, we explain the history of the discovery of the cyanobacterial pathway, and present a probable scenario of the evolution of the two pathways.
Introduction
Cyanobacteria perform oxygenic photosynthesis like chloroplasts of land plants and algae. The initial reactions of photosynthesis such as photochemical reactions, electron transport reactions, and ATP synthesis are performed in the thylakoid membranes, namely, flattened sac-like membranes specialized for photosynthesis. Thylakoid membranes are built up with galactolipids and acidic lipids. There are two major classes of galactolipids, monogalactosyl diacylglycerol (MGDG) and digalactosyl diacylglycerol (DGDG), which are both major components of all thylakoid membranes, in other words, typical of photosynthetic organisms. MGDG was shown to be required for normal development of chloroplasts (Kobayashi et al., 2007, 2013). The universality of galactolipids in photosynthetic membranes has been understood in terms of endosymbiotic theory, namely, that chloroplasts originated from cyanobacteria (see for example, Sato, 2001, 2006; Petroutsos et al., 2014).
Biosynthesis of these galactolipids is, however, quite different in cyanobacteria and chloroplasts (Figure 1). In the chloroplasts of land plants, MGDG is synthesized by galactosylation of diacylglycerol (DAG), and DGDG is synthesized by the second galactosylation of MGDG (Shimojima and Ohta, 2011; Dörmann, 2013). In contrast, cyanobacteria have monoglucosyl diacylglycerol (GlcDG; Feige et al., 1980), which serves as a precursor to MGDG (Sato and Murata, 1982a). The conversion of GlcDG to MGDG was presumed to be epimerization, namely, the isomerization at the C-4 of the glucose moiety (Sato and Murata, 1982a). The glucosyltransferase activity was subsequently demonstrated (Sato and Murata, 1982b), while the enzymatic activity of the epimerase has never been detected in vitro. This was the reason why the epimerization hypothesis remained elusive until the identification of the responsible gene (Awai et al., 2014). Plant galactosyltransferases for the synthesis of MGDG and DGDG have been identified in the late 1990s and named MGD1 (Shimojima et al., 1997) and DGD1 (Dörmann et al., 1999), respectively, and homologs MGD2/3 and DGD2 were found later. In contrast, the enzymes involved in the synthesis of galactolipids in cyanobacteria have been uncharacterized until quite recently.
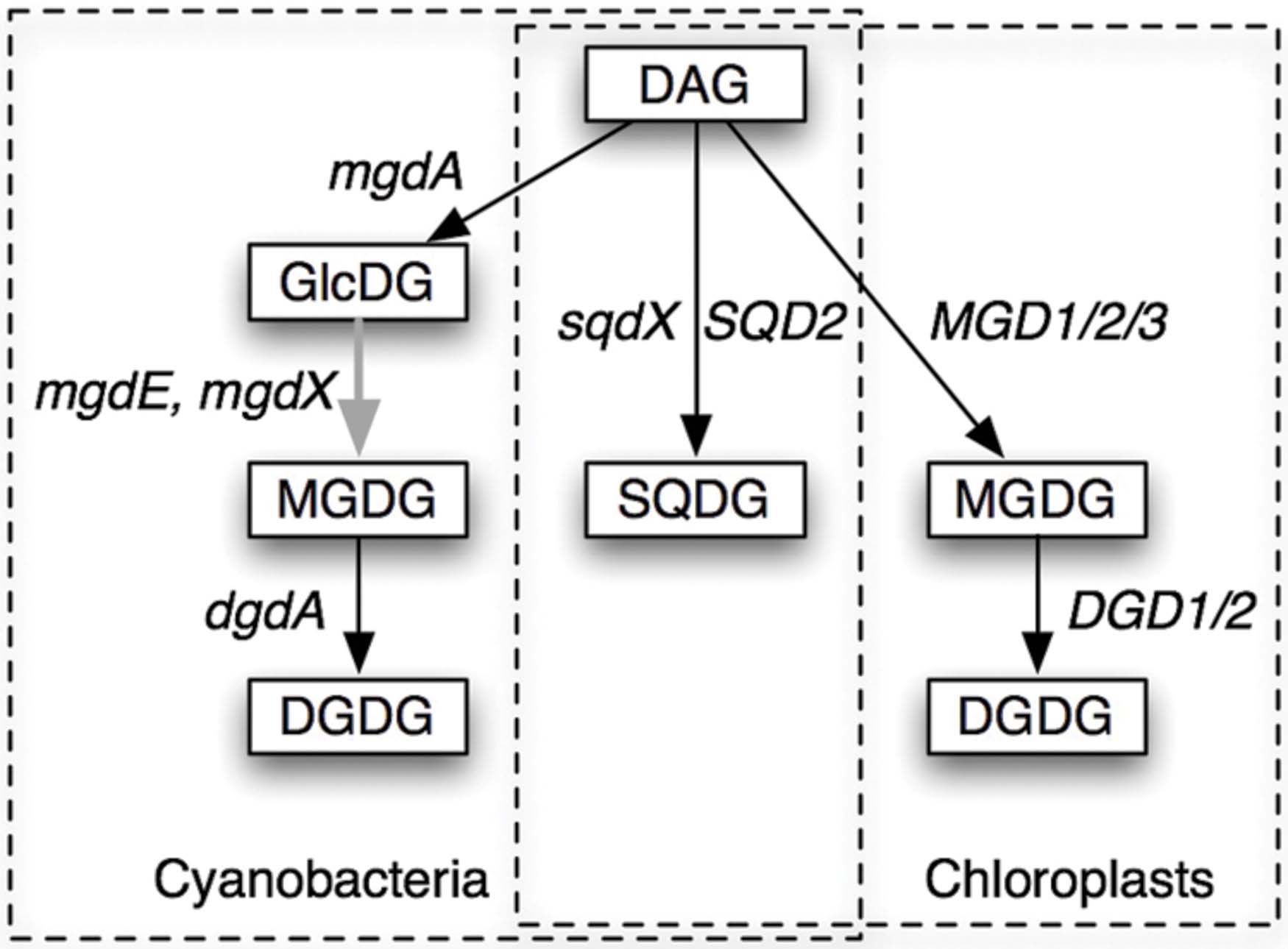
FIGURE 1. Pathways of synthesis of glycolipids in cyanobacteria and chloroplasts. The pathways and the genes involved in the processes are shown. DAG, diacylglycerol; GlcDG, monoglucosyl diacylglycerol; MGDG, monogalactosyl diacylglycerol; DGDG, digalactosyl diacylglycerol; SQDG, sulfoquinovosyl diacylglycerol. Gene names are explained in the text. mgdX is a hypothetical alternative gene encoding the epimerase. Solid arrows indicate glycosyltransferases (GT), while the gray arrow indicates an epimerase.
Identification of the Enzymes/Genes for the Synthesis of Galactolipids
Identification of Glycosyltransferases (GT)
Awai et al. (2006) identified the gene mgdA encoding the glucosyltransferase for the synthesis of GlcDG in Synechocystis sp. PCC 6803. The enzyme MgdA belongs to the glycosyltransferase (GT) family 2, and its domain structure was quite different from the plant galactosyltransferases MGD1/2/3, which belong to the GT 28, according to the CAZy database (Lombard et al., 2014). They also contain different InterPro domains (Mitchell et al., 2015). Homologs of MGD1 were found in green non-sulfur (GNS) bacteria (Yuzawa et al., 2012; Supplementary Figure S1; see Cluster 2103 of dataset Gclust2012_42 in comparative genomic database Gclust: Sato, 2009, available at http://gclust.c.u-tokyo.ac.jp/), but they are different from MgdA. Distant homologs of MgdA are found in α-proteobacteria (Supplementary Figure S2; see Gclust Cluster 2866).
No homolog of DGD1 had been detected in the sequenced cyanobacterial genomes or in the genome of Cyanidioschyzon merolae, which was the only red alga that was sequenced in the early 2000s. Cyanobacterial galactosyltransferase that catalyzes the synthesis of DGDG was identified by exploiting comparative genomics (Sato et al., 2005; Sato, 2009), which identified a GT that was conserved in cyanobacteria and C. merolae (Awai et al., 2007; Sakurai et al., 2007). The enzyme named DgdA has two InterPro domains, N-terminal Glycosyltransf_like_4 and C-terminal Glycosyltransf_1, whereas DGD1/2 has a single Glycosyltransf_1 domain. Both DgdA and DGD1//2 belong to the CAZy GT4 family. DGD1/2 has homologs in only plants and algae (Gluster 2454 in Gclust). It is interesting to note that DgdA is a close relative of SqdX/SQD2, an enzyme catalyzing the transfer of sulfoquinovose (Figure 1). This enzyme has a dual GT domain similar to that in DgdA. Phylogenetic analysis suggests that plant SQD2 originates from cyanobacterial SqdX, which is also found in various bacteria such as α-proteobacteria and GNS bacteria (Supplementary Figure S3). This points to a possibility that the gene dgdA originated from an sqdX-like gene before the emergence of cyanobacteria. Based on these considerations, it is likely that mgdA and dgdA originated from α-proteobacteria and GNS bacteria, respectively (Figure 2). Curiously, the SQDG synthesis pathway has been lost in Gloeobacter violaceus, which is the most deeply branching species in cyanobacteria.
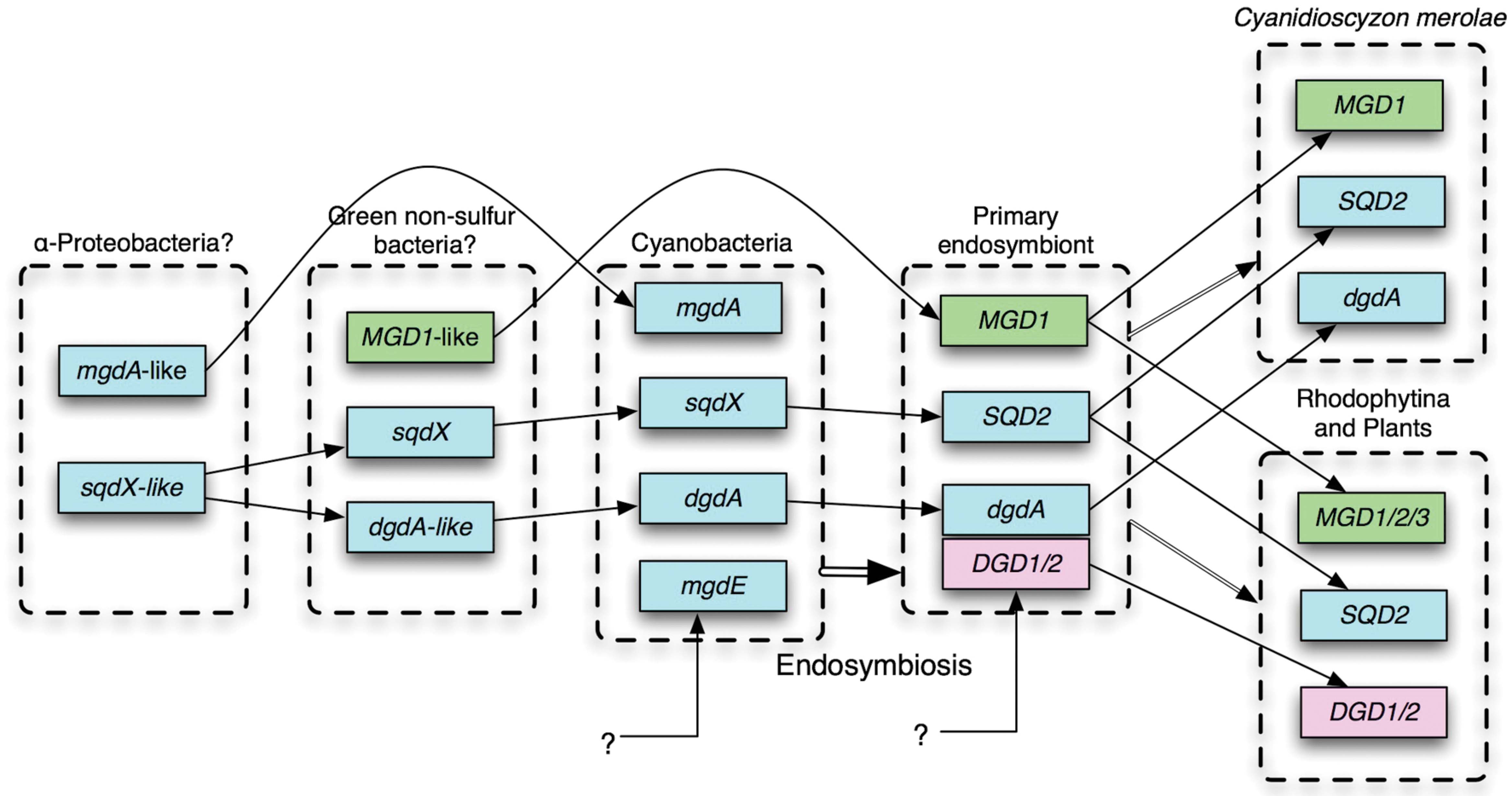
FIGURE 2. Probable evolutionary scenario of glycolipid biosynthesis enzymes. For details, see text. Cyan, genes identified in cyanobacteria; green, originated from green non-sulfur (GNS) bacteria; pink, unknown eukaryotic origin.
Identification of GlcDG Epimerase
The epimerase remained unidentified for a long time despite considerable efforts of many researchers. The use of Gclust, however, gave the clue again. Important additional information was given by the chromatophore genome of Paulinella chromatophora. The chromatophore of this microorganism looks like a chloroplast, but the sequencing of the chromatophore genome suggested that it originated from Prochlorococcus-like cyanobacteria (Nowack et al., 2008), but does not belong to the lineage of known chloroplasts, which are all monophyletic and originated from the deep root of cyanobacteria (Shih et al., 2013). The chromatophore genome encodes mgdA, dgdA, and sqdX. It was quite possible that an unknown GlcDG epimerase is also encoded in the genome. Comparative genomic analysis using Gclust indeed revealed that a putative membrane-bound oxidoreductase is conserved in most cyanobacteria and the chromatophore (Awai et al., 2014). Expression in Escherichia coli of the corresponding gene sll1376 in Synechocystis sp. PCC 6803 demonstrated that it encodes an enzyme converting GlcDG to MGDG. Disruption of the gene in Synechocystis resulted in the cells in which all MGDG and DGDG were replaced by GlcDG. The gene was named mgdE. The enzyme MgdE consisted of an N-terminal hydrophobic domain assigned as the ‘fatty acid hydroxylase domain’ and a C-terminal oxidoreductase domain known as the ‘Rossmann-fold’ (Rao and Rossmann, 1973), thus considered as having a reasonable structure as a membrane-bound epimerase.
Role of MgdE in Epimerization
The enzyme MgdE was conserved in many cyanobacteria, but curiously, not in all cyanobacteria. The gene mgdE was not detected in at least G. violaceus, Thermosynechococcus elongatus, and Acariochloris marina. In addition, the enzyme in various strains of Prochlorococcus marinus, marine Synechococcus species as well as P. chromatophora lacked the N-terminal domain. This raised again a fundamental question regarding the pathway of galactolipid synthesis in cyanobacteria, namely, if all cyanobacteria contain GlcDG as a precursor to MGDG. The detection of GlcDG has been especially difficult because it is a very minor component in many cyanobacteria. Even in a recently published review [Figure 1 and Supplementary Table S1 in Petroutsos et al. (2014)], GlcDG was described as undetected in many cyanobacteria such as Synechocystis sp. PCC 6803 and G. violaceus.
This situation made it important to re-analyze GlcDG in the cyanobacteria in which mgdE was not detected or truncated. Sato (2015b) isolated and identified GlcDG in G. violaceus, T. elongatus, A. marius as well as in P. marinus. In addition, the conversion of GlcDG to MGDG was demonstrated by radiolabeling experiments in G. violaceus and P. marinus. Comparison of the composition of molecular species of GlcDG and MGDG also suggested that GlcDG can be considered as a precursor to MGDG in all the cyanobacteria analyzed, on the assumption that the molecular species containing saturated fatty acids are synthesized first and then the acyl groups are desaturated on the intact glycolipids while keeping the overall structure of lipids (Sato et al., 1986). This raised a question as to the universality of the role of mgdE in the epimerization of GlcDG in cyanobacteria. Is there another GlcDG epimerase in some cyanobacteria? In that case, how was the pathway of galactolipid synthesis acquired in the cyanobacteria?
MgdE belongs to a large family of bifunctional sterol desaturases/short-chain dehydrogenases (Kramm et al., 2012), including various enzymes related to lipid metabolism such as FabG, 3-oxoacyl-ACP reductase, involved in fatty acid biosynthesis [Supplementary Figure S4 in Sato (2015b)]. In this respect, other members of this family could act as GlcDG epimerase (encoded by a hypothetical gene mgdX in Figure 1). This will be a new perspective of MGDG synthesis in cyanobacteria.
Evolution of Galactolipid Biosynthesis
Phylogenetic Distribution of the Cyanobacterial Pathway in Algae
As described above, the pathway of galactolipid biosynthesis is significantly different in cyanobacteria and chloroplasts of land plants. An immediate question arises as to what the situation is in algae. The chloroplasts of Archaeplastida (green plants, red algae, and glaucophytes) are monophyletic and are believed to originate from a single endosymbiotic event (Sato, 2001, 2006; Nowack et al., 2008; Shih et al., 2013). A survey of the Gclust database shows that the green algae, such as Chlamydomonas reinhardtii and Ostreococcus tauri, have a pathway consisting of MGD1 and DGD1, like land plants. The situation in red algae is complicated. Cyanidioschyzon merolae has a plastid-encoded dgdA, but has a copy of MGD1. Curiously, an mgdA homolog is also encoded in the C. merolae genome, although no GlcDG was detected by careful analysis (Sato and Moriyama, 2007). The dgdA gene (also known as ycf82) is also found in the plastid genomes of Cyanidiales algae, Galdieria sulphuraria and Cyanidium caldarium. This is not the case in another red alga Porphyridium purpureum, in which dgdA is not encoded in the plastid genome (Tajima et al., 2014), but a putative DGD1 is encoded in the nuclear genome (Bhattacharya et al., 2013). The same is true for other red algae in Rhodophytina (Awai, 2015), which are unicellular or macrophytic red algae belonging to non-Cyanidiales clades. The glaucophyte Cyanophora paradoxa encodes MGD1 and dgdA (Awai, 2015). The heterokonts such as diatoms are supposed to originate from a red algal secondary endosymbiosis, but none of them encodes dgdA.
Concluding Remarks: Probable Evolutionary Scenarios
Comparative genomics clearly shows that all eukaryotes have MGD1 whereas all cyanobacteria have MgdA. Although there could be at least two different entities of GlcDG epimerase, all cyanobacteria are likely to synthesize MGDG by epimerization of GlcDG. In this respect, the replacement of MgdA–MgdE system by green bacterial MGD1 accompanied the primary endosymbiosis (Figure 2), which also accompanied drastic changes in the transcriptional and genomic machineries (Sato, 2001): Namely, prokaryotic transcription factors and DNA-binding proteins are not conserved in the chloroplasts, while the sigma factors become encoded by the nucleus.
In contrast, the replacement of DgdA by DGD1 seemed to occur at least two times, namely, during the evolution of green algae on the one hand, and during the evolution of Rhodophytina in the red algae on the other hand (Sato, 2015a), because all red algae are monophyletic (Tajima et al., 2014). Another, more plausible possibility implies that the dgdA gene and the DGD1 gene coexisted in a hypothetical primary endosymbiont, and one of them was lost subsequently in different lineages (Figure 2). This is more likely because all DGD1 in both green plants and red algae are monophyletic (Supplementary Figure S3). A simple question should be asked: what is the functional difference between DgdA and DGD1, and why the two types of enzymes are present? To answer the question, we will have to know the enzymology and three-dimensional structure of the respective enzymes. In addition, it is clear that a more extensive survey of all algal phyla will be necessary to reveal the whole picture of the distribution of the two pathways, and hence the evolution of the DGDG biosynthesis.
Author Contributions
Both authors contributed to all parts of collaborative works, including experiments and writing.
Funding
This work was supported in part by grants-in-aid for the Core Research for Evolutional Science and Technology (CREST) and the Precursory Research for Embryonic Science and Technology (PRESTO) from the Japan Science and Technology Agency.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2016.00117
References
Awai, K. (2015). Evolution and distribution of galactolipid biosynthetic pathways in photosynthetic organisms. News Lett. Japan. Soc. Photosynth. Res. 25, 143–150.
Awai, K., Kakimoto, T., Awai, C., Kaneko, T., Nakamura, Y., Takamiya, K., et al. (2006). Comparative genomic analysis revealed a gene for monoglucosyldiacylglycerol synthase, an enzyme for photosynthetic membrane lipid synthesis in cyanobacteria. Plant Physiol. 141, 1120–1127. doi: 10.1104/pp.106.082859
Awai, K., Ohta, H., and Sato, N. (2014). Oxygenic photosynthesis without galactolipids. Proc. Natl. Acad. Sci. U.S.A. 111, 13571–13575. doi: 10.1073/pnas.1403708111
Awai, K., Watanabe, H., Benning, C., and Nishida, I. (2007). Digalactosyldiacylglycerol is required for better photosynthetic growth of Synechocystis sp. PCC6803 under phosphate limitation. Plant Cell Physiol. 48, 1517–1523. doi: 10.1093/pcp/pcm134
Bhattacharya, D., Price, D. C., Chan, C. X., Qiu, H., Rose, N., Ball, S., et al. (2013). Genome of the red alga Porphyridium purpureum. Nat. Commun. 4:1941. doi: 10.1038/ncomms2931
Dörmann, P. (2013). Galactolipids in Plant Membranes, Encyclopedia of Life Sciences. Hoboken, NJ: Wiley Online Library.
Dörmann, P., Balbo, I., and Benning, C. (1999). Arabidopsis galactolipid biosynthesis and lipid trafficking mediated by DGD1. Science 284, 2181–2184. doi: 10.1126/science.284.5423.2181
Feige, G., Heinz, E., Wrage, K., Cochems, N., and Ponzelar, E. (1980). “Discovery of a new glyceroglycolipid in blue-green algae and its role in galactolipid biosynthesis,” in Biogenesis and Function of Plant Lipids, eds P. Mazliak, P. Benveniste, C. Castes, and R. Douce (Amsterdam: Elsevier/North Holland Biomedical Press), 135–140.
Kobayashi, K., Kondo, M., Fukuda, H., Nishimura, M., and Ohta, H. (2007). Galactolipid synthesis in chloroplast inner envelope is essential for proper thylakoid biogenesis, photosynthesis, and embryogenesis. Proc. Natl Acad. Sci. U.S.A. 104, 17216–17221. doi: 10.1073/pnas.0704680104
Kobayashi, K., Narise, T., Sonoike, K., Hashimoto, H., Sato, N., Kondo, M., et al. (2013). Role of galactolipid biosynthesis in coordinated development of photosynthetic complexes and thylakoid membranes during chloroplast biogenesis in Arabidopsis. Plant J. 73, 250–261. doi: 10.1111/tpj.12028
Kramm, A., Kisiela, M., Schulz, R., and Maser, E. (2012). Short-chain dehydrogenases/reductases in cyanobacteria. FEBS J. 279, 1030–1043. doi: 10.1111/j.1742-4658.2012.08494.x
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M., and Henrissat, B. (2014). The Carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495. doi: 10.1093/nar/gkt1178
Mitchell, A., Chang, H. Y., Daugherty, L., Fraser, M., Hunter, S., Lopez, R., et al. (2015). The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res. 43, D213–D221. doi: 10.1093/nar/gku1243
Nowack, E. C., Melkonian, M., and Glockner, G. (2008). Chromatophore genome sequence of Paulinella sheds light on acquisition of photosynthesis by eukaryotes. Curr. Biol. 18, 410–418. doi: 10.1016/j.cub.2008.02.051
Petroutsos, D., Amiar, S., Abida, H., Dolch, L. J., Bastien, O., Rébeillé, F., et al. (2014). Evolution of galactoglycerolipid biosynthetic pathways–from cyanobacteria to primary plastids and from primary to secondary plastids. Prog. Lipid Res. 54, 68–85. doi: 10.1016/j.plipres.2014.02.001
Rao, S. T., and Rossmann, M. G. (1973). Comparison of super-secondary structures in proteins. J. Mol. Biol. 76, 241–256. doi: 10.1016/0022-2836(73)90388-4
Sakurai, I., Mizusawa, N., Wada, H., and Sato, N. (2007). Digalactosyldiacylglycerol is required for stabilization of the oxygen-evolving complex in photosystem II. Plant Physiol. 145, 1361–1370. doi: 10.1104/pp.107.106781
Sato, N. (2001). Was the evolution of plastid genetic machinery discontinuous? Trends Plant Sci. 6, 151–156. doi: 10.1016/S1360-1385(01)01888-X
Sato, N. (2006). “Origin and evolution of plastids: genomic view on the unification and diversity of plastids,” in The Structure and Function of Plastids, eds R. Robert, J. Wise, and H. Kenneth (Berlin: Springer), 75–102. doi: 10.1007/978-1-4020-4061-0_4
Sato, N. (2009). Gclust: trans-kingdom classification of proteins using automatic individual threshold setting. Bioinformatics 25, 599–605. doi: 10.1093/bioinformatics/btp047
Sato, N. (2015a). Glycolipid biosynthesis and oxygenic photosynthesis in cyanobacteria. Seikagaku 87, 209–211.
Sato, N. (2015b). Is monoglucosyl diacylglycerol a precursor to monogalactosyl diacylglycerol in all cyanobacteria? Plant Cell Physiol. 56, 1890–1899. doi: 10.1093/pcp/pcv116
Sato, N., Ishikawa, M., Fujiwara, M., and Sonoike, K. (2005). Mass identification of chloroplast proteins of endosymbiont origin by phylogenetic profiling based on organism-optimized homologous protein groups. Genome Inform. 16, 56–68.
Sato, N., and Moriyama, T. (2007). Genomic and biochemical analysis of lipid biosynthesis in the unicellular rhodophyte Cyanidioschyzon merolae: lack of plastidic desaturation pathway results in mixed pathway of galactolipid synthesis. Eukaryot. Cell 6, 1006–1017. doi: 10.1128/EC.00393-06
Sato, N., and Murata, N. (1982a). Lipid biosynthesis in the blue-green alga, Anabaena variabilis: I. Lipid classes. Biochim. Biophys. Acta 710, 271–278. doi: 10.1016/0005-2760(82)90109-6
Sato, N., and Murata, N. (1982b). Lipid biosynthesis in the blue-green alga (cyanobacterium), Anabaena variabilis III. UDP-glucose:diacylglycerol glucosyltransferase activity in vitro. Plant Cell Physiol. 23, 1115–1120.
Sato, N., Seyama, Y., and Murata, N. (1986). Lipid-linked desaturation of palmitic acid in monogalactosyl diacylglycerol in the blue-green alga (cyanobacterium) Anabaena variabilis. Plant Cell Physiol. 27, 819–835.
Shih, P. M., Wu, D., Latifi, A., Axen, S. D., Fewer, D. P., Talla, E., et al. (2013). Improving the coverage of the cyanobacterial phylum using diversity-driven genome sequencing. Proc. Natl. Acad. Sci. U.S.A. 110, 1053–1058. doi: 10.1073/pnas.1217107110
Shimojima, M., and Ohta, H. (2011). Critical regulation of galactolipid synthesis controls membrane differentiation and remodeling in distinct plant organs and following environmental changes. Prog. Lipid Res. 50, 258–266. doi: 10.1016/j.plipres.2011.03.001
Shimojima, M., Ohta, H., Iwamatsu, A., Masuda, T., Shioi, Y., and Takamiya, K. (1997). Cloning of the gene for monogalactosyldiacylglycerol synthase and its evolutionary origin. Proc. Natl. Acad. Sci. U.S.A. 94, 333–337. doi: 10.1073/pnas.94.1.333
Tajima, N., Sato, S., Maruyama, F., Kurokawa, K., Ohta, H., Tabata, S., et al. (2014). Analysis of the complete plastid genome of the unicellular red alga Prophyridium purpureum. J. Plant Res. 127, 389–397. doi: 10.1007/s10265-014-0627-1
Yuzawa, Y., Nishihara, H., Haraguchi, T., Masuda, S., Shimojima, M., Shimoyama, A., et al. (2012). Phylogeny of galactolipid synthase homologs together with their enzymatic analyses revealed a possible origin and divergence time for photosynthetic membrane biogenesis. DNA Res. 19, 91–102. doi: 10.1093/dnares/dsr044
Keywords: cyanobacteria, galactolipids, glucolipid, photosynthesis, endosymbiosis
Citation: Sato N and Awai K (2016) Diversity in Biosynthetic Pathways of Galactolipids in the Light of Endosymbiotic Origin of Chloroplasts. Front. Plant Sci. 7:117. doi: 10.3389/fpls.2016.00117
Received: 25 November 2015; Accepted: 22 January 2016;
Published: 05 February 2016.
Edited by:
Jeffrey Leblond, Middle Tennessee State University, USACopyright © 2016 Sato and Awai. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Naoki Sato, naokisat@bio.c.u-tokyo.ac.jp
 Naoki Sato
Naoki Sato Koichiro Awai2,3
Koichiro Awai2,3